Clear Sky Science · en
MiGPC: a comprehensive catalog of enzybiotics from environmental metagenomes
Why tiny enzymes matter for our future
As antibiotic resistance rises and plastics and pollutants spread through our air, water, and soil, scientists are searching for new ways to control harmful microbes without harming the rest of nature. This study introduces MiGPC, a vast catalog of natural germ fighting enzymes drawn from many different environments, from oceans and hot springs to animal guts and plastic litter. It shows how rich Earth’s hidden microbial world is in potential new antimicrobials and offers a roadmap for turning that diversity into practical tools for medicine, food safety, and environmental cleanup. 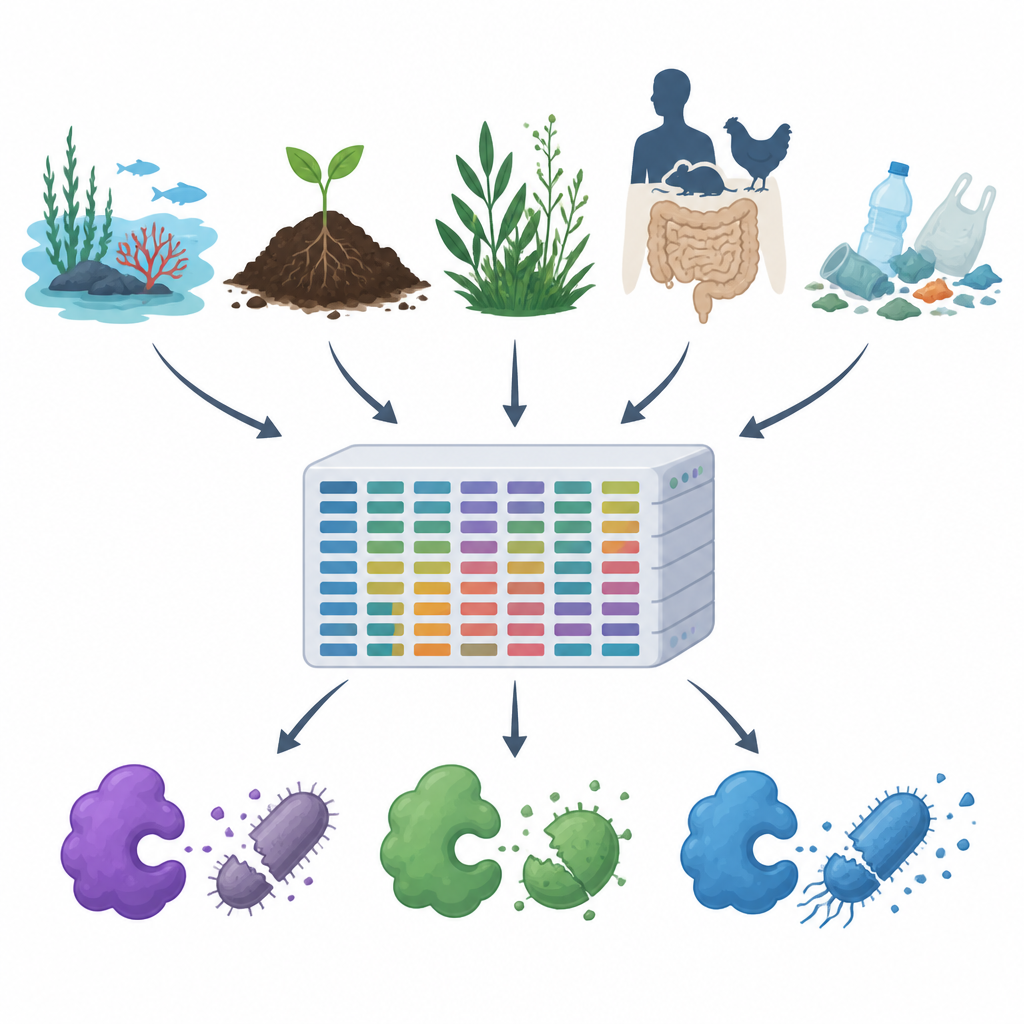
From microbes to natural germ killers
Traditional antibiotics often act like blunt instruments, wiping out helpful bacteria along with harmful ones and encouraging resistance over time. By contrast, many microbes use specialized enzymes to attack rivals with great precision, poking holes in cell walls or generating toxic molecules right where they are needed. These enzyme based antimicrobials, called enzybiotics, can burst bacterial cells, break down protective slime layers, or disrupt survival pathways. The authors set out to find how widely such enzymes occur in nature and to gather their genetic blueprints into a single, organized resource that anyone can explore.
Building a global enzyme catalog
The team first combed through public protein databases and scientific papers to collect 19 well studied antimicrobial enzymes, including cell wall cutters, protein digesting enzymes, and oxidizing enzymes that generate harmful reactive molecules. They then used these proven examples as bait to scan large DNA datasets from 15 very different environments, such as seawater, brine pools, river estuaries, hot springs, alpine plants, animal and human guts, and plastic contaminated soils and waters. Using a high powered computational pipeline, they cleaned and assembled about 1.2 terabytes of raw DNA reads into longer fragments, predicted genes and proteins, removed duplicates, and grouped related sequences, ending up with roughly 100 million unique genes and proteins.
Finding hidden functions in environmental DNA
To understand what these genes might do, the researchers compared them against major functional databases. Even with modern tools, about 62 percent of the genes could not be assigned a known role, hinting at a deep well of unexplored biology. Among recognizable functions, enzymes that work on complex sugars were especially common. Two families glycoside hydrolases and glycosyl transferases stood out, many of which can weaken bacterial cell walls or biofilms. A targeted search with a specialized tool uncovered nearly 20,000 likely antimicrobial enzymes. Most belonged to two main groups: hydrolases, which cut molecules apart, and oxidoreductases, which drive reactions involving electrons and can generate reactive oxygen species that damage microbes. 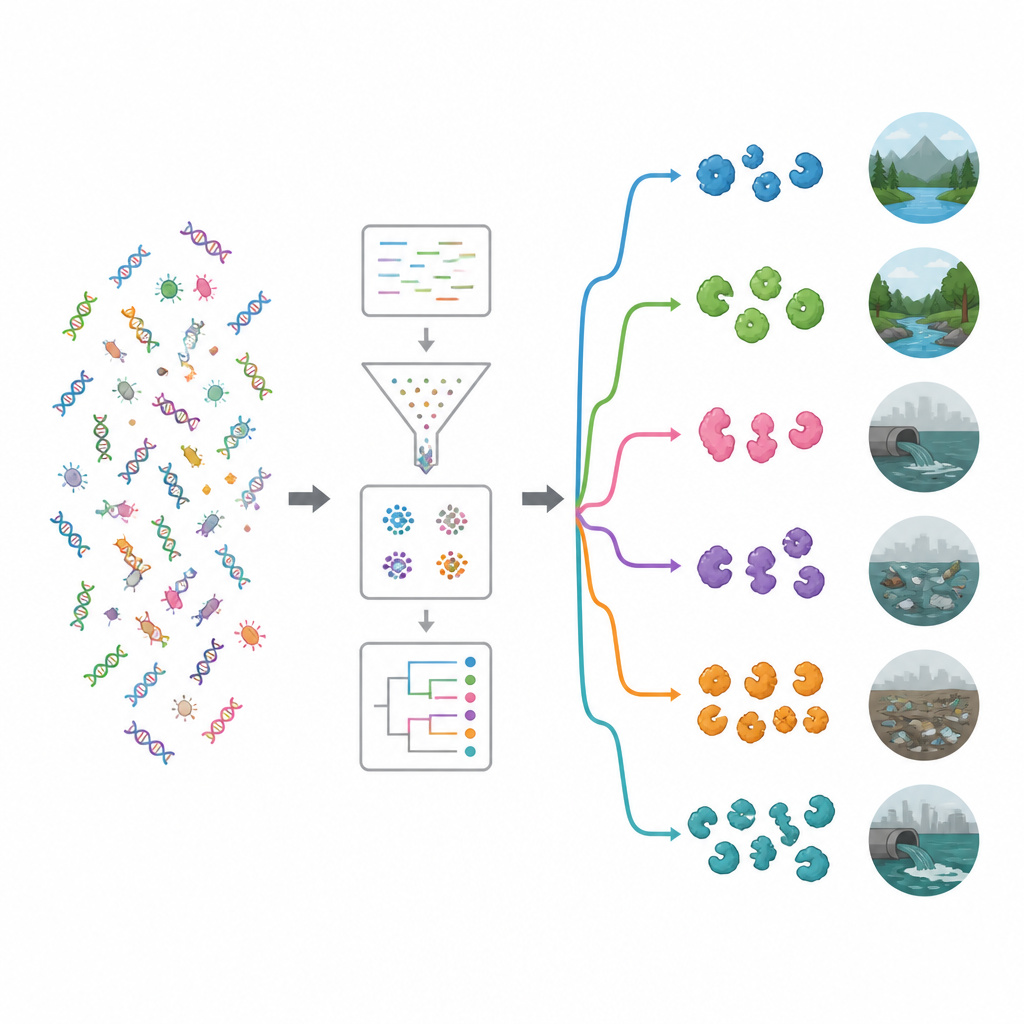
Who makes these enzymes and where they thrive
By matching enzyme bearing genes to their microbial hosts, the study showed that many came from bacterial groups already known to produce natural antimicrobials, particularly Pseudomonadota and Bacillota, as well as actinomycetes like Streptomyces. The team then examined which bacteria were abundant in each environment and used clustering and machine learning to sort the sites into ecological groups. One cluster contained relatively pristine settings such as rivers, groundwater, vegetation, brine, and hot springs. The other cluster held more human influenced habitats, including plastic polluted sites, wastewater, reservoirs, oceans near farms, and fecal samples. Polluted environments tended to harbor a richer and more diverse set of enzybiotic genes, suggesting that intense competition and chemical stress favor microbes that invest heavily in germ killing tools.
What this means for health and the environment
MiGPC is more than a giant list of genes; it is a map that links specific enzymes, microbes, and environments. For non specialists, the takeaway is that Earth’s stressed and untouched habitats alike are packed with natural molecules that can fight dangerous bacteria, including those that no longer respond well to standard antibiotics. Many of these enzymes are still functionally mysterious, but this catalog makes it far easier for researchers to find, test, and engineer them. Over time, MiGPC could guide the development of new enzyme based treatments, safer food preservatives, and tools to manage harmful microbes in farms and polluted sites, helping society respond to antimicrobial resistance without relying solely on traditional drugs.
Citation: Afshar Jahanshahi, D., Ariaeenejad, A., Hasannejad, A. et al. MiGPC: a comprehensive catalog of enzybiotics from environmental metagenomes. Sci Rep 16, 15153 (2026). https://doi.org/10.1038/s41598-026-44250-9
Keywords: enzybiotics, antimicrobial enzymes, metagenomics, microbial ecology, antibiotic resistance