Clear Sky Science · en
Detection and classification of chromosomes with sister chromatid cohesion defects using object detection models
Why tiny twists in DNA matter
Every time a cell divides, it must pass along an exact copy of its genetic material. To keep these copies together until the right moment, cells rely on a molecular “glue” that holds paired chromosomes side by side. When this glue fails, chromosomes can separate too early, leading to broken genomes, developmental disorders, and cancer. The study described here tackles a very practical problem: how to spot these subtle glue failures automatically in microscope images, using modern artificial intelligence instead of tired human eyes.
How chromosomes stay together
Before a cell splits, each chromosome is copied so there are two identical strands, known as sister chromatids. A protein machine called cohesin helps keep these sisters paired until the cell is ready to pull them apart. If cohesin or its helper proteins malfunction, the sisters can drift away from each other too soon. One such helper is DDX11, a DNA-unwinding enzyme whose defects are linked to a rare human condition called Warsaw Breakage syndrome. Under the microscope, cells with cohesion problems often show chromosomes whose two strands are oddly open or split at the center, but telling normal from abnormal shapes requires patience and judgment.
Why manual checks are not enough
Traditionally, researchers spread chromosomes on a slide, stain them, and then classify at least 50 to 100 of them per sample by hand. Each chromosome is judged by its shape and how far apart its sister strands appear. This work is slow, subjective, and easy to tire of, which increases the risk of mistakes and inconsistent results between scientists. Earlier efforts by the same research group used deep learning to classify individual chromosome images, but those systems required separate software steps to cut out single chromosomes from crowded microscopy pictures and to discard confusing overlaps. That extra image handling still depended heavily on human tuning and cleanup, blocking true “push-button” automation.
Letting the computer find chromosomes on its own
In the new study, the team turned to object detection, a branch of computer vision that can both find and categorize many items within a single image. They focused on a family of fast, widely used models called YOLO (short for “You Only Look Once”) along with several competing methods. First, they created a test system by engineering human cells that lack DDX11 and therefore show strong cohesion problems, then preparing microscope images of their chromosomes along with those from normal cells. Each visible chromosome in these images was outlined with a box and labeled into one of three shape types: tightly paired, partly separated along the arms, or clearly split at the center. These hand-made labels served as the teaching material for the algorithms.
Teaching AI to see subtle shape differences
The researchers fine-tuned several pre-trained object detection models on just over a thousand labeled chromosomes, reserving other images for checking performance. They compared how well each model could first find chromosomes at all and then assign them to the correct of the three shape categories. Among the tested systems, a version called YOLOv8 performed best. It correctly matched human labels for nearly nine out of ten chromosomes, even though the differences between categories were often quite subtle. Importantly, when applied to images from normal and DDX11-deficient cells, YOLOv8 reproduced the key biological pattern that human experts see: the faulty cells had far fewer tightly paired chromosomes and many more with clearly separated sister strands.
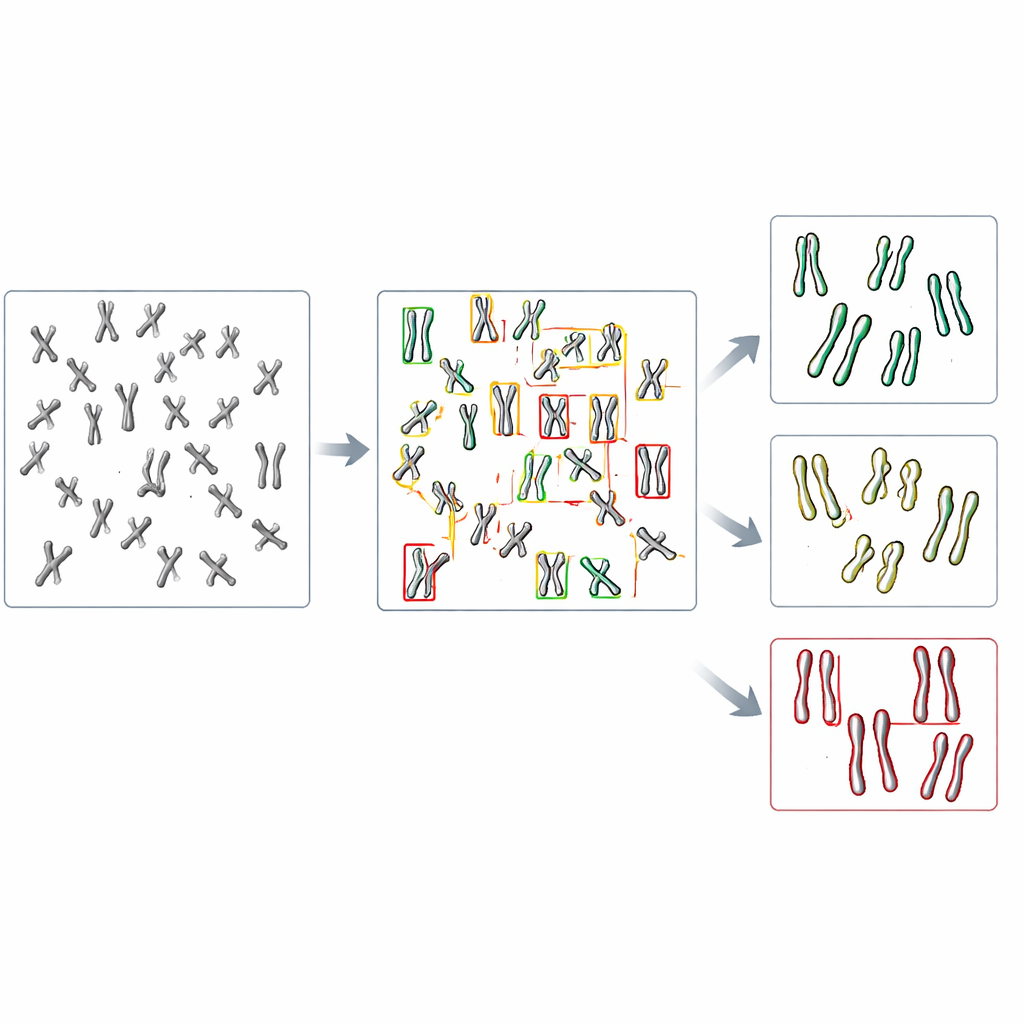
What this means for future lab work
The study shows that an AI model can scan full chromosome spreads and automatically flag cohesion defects without extra cropping steps or manual cleanup. While there is still room to improve performance—especially for overlapped chromosomes and borderline shapes—the approach already rivals earlier methods that required more human intervention. Because the models were built by adapting existing open-source tools and can be retrained with modest image sets, laboratories without deep computing expertise could develop their own tailored systems. In the long run, similar strategies may speed up screening for drugs, help compare many genetic mutants, and extend beyond cohesion to other delicate chromosome changes that matter for human health.
Citation: Matsumoto, S., Sojo, M., Oshima, K. et al. Detection and classification of chromosomes with sister chromatid cohesion defects using object detection models. Sci Rep 16, 13719 (2026). https://doi.org/10.1038/s41598-026-43009-6
Keywords: chromosome imaging, machine learning, object detection, cohesion defects, YOLOv8