Clear Sky Science · en
mViSE: A visual search engine for analyzing multiplex IHC brain tissue images (spatial proteomics)
Seeing Patterns in the Brain
Modern microscopes can now capture breathtakingly detailed images of entire brain slices, showing dozens of different proteins at once. These “multiplex” images promise clues about how brain cells are organized, how they talk to each other, and how disease disrupts these patterns. But the images are so huge and complex that even powerful computers struggle to make sense of them. This paper introduces mViSE, a visual search engine that lets researchers explore these vast brain images by simply clicking on cells and neighborhoods of interest, rather than writing custom code.
Why Big Brain Images Are Hard to Use
Each multiplex brain image is like a giant city map where every building, street, and utility line is labeled in many colors at once. Different proteins mark different cell types, cell states, blood vessels, and wiring patterns. Traditional analysis pipelines break this map into a long chain of programmed steps: clean the image, detect cells, segment them, label cell types, and then summarize results by region. While powerful, this approach is rigid and hard to adapt when new questions arise—especially for the brain, whose mix of neurons, glia, and blood vessels has immense molecular and spatial complexity. Scientists increasingly need a more flexible, interactive way to pose new questions to these data without becoming full-time programmers.
A Search Engine for Brain Cells
mViSE treats a brain image more like an online photo library than a static dataset. Instead of predefining every analysis step, the user clicks on a cell or a small tissue patch that looks interesting. The system then searches the entire brain image for cells or neighborhoods that look and “behave” similarly across many protein channels. The matches are highlighted directly on the whole-brain view and can be summarized as protein expression profiles. This lets researchers rapidly uncover where similar cells or microenvironments appear, delineate brain regions and cortical layers, and compare patterns across different parts of the brain—all driven by visual queries, not code. 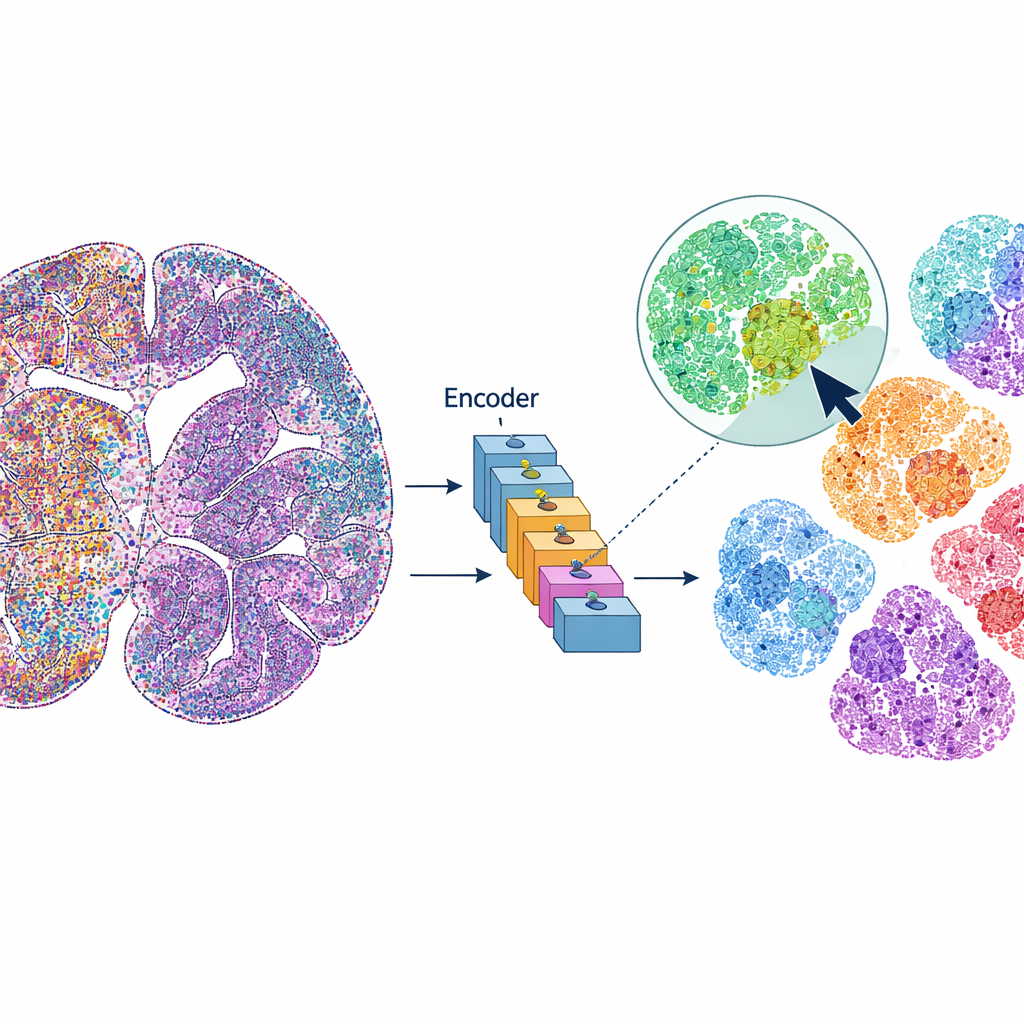
Teaching the Computer to Understand Tissue
To make this work, mViSE first goes through a learning phase entirely in the background. The authors divide the many protein channels into biologically meaningful panels, such as markers that highlight cell types, glial cells, nerve fibers, or blood vessels. For each panel, a powerful vision transformer—a modern deep-learning model—looks at many small patches across the brain and learns to represent each patch as a point in a high-dimensional “feature space.” Patches that look and act alike should end up close together in this space. An information-theoretic community detection method then groups nearby points into communities that reflect repeating patterns of tissue architecture. Importantly, this learning is self-supervised: it does not require human labeling, but it still produces color-coded maps that visually confirm that known layers and regions are being captured.
Zooming In on Cells and Neighborhoods
Once trained, mViSE can respond to different types of queries. For very small patches containing single cells, it retrieves the most similar cells across the brain and shows where they are located. The authors demonstrate that the system reliably finds major brain cell types—such as neurons, astrocytes, oligodendrocytes, microglia, and vascular cells—as well as more specific neuronal subtypes, and even cell pairs located next to blood vessels. For larger patches that include multiple cells and local wiring, mViSE returns the full community of similar neighborhoods, often tracing out known brain regions or tracts. By combining information from several marker panels at once, it can also tease apart subtle differences between cortical layers and small subregions that are otherwise hard to distinguish.
Outperforming General-Purpose AI Models
The researchers compared mViSE with several state-of-the-art “foundation” models originally trained on large collections of natural or pathology images. Because those models expect simple three-color images, the many protein channels had to be squeezed into red, green, and blue, causing loss of information. Even after this adaptation, these general models produced fuzzier boundaries between layers and missed fine-grained subregions. In contrast, mViSE, which is designed to handle many channels directly and encodes each channel separately before combining them, achieved higher accuracy in matching known brain atlas regions. It produced sharper maps of cortical layers and more coherent communities of similar patches, indicating that its representations better reflect true biological organization. 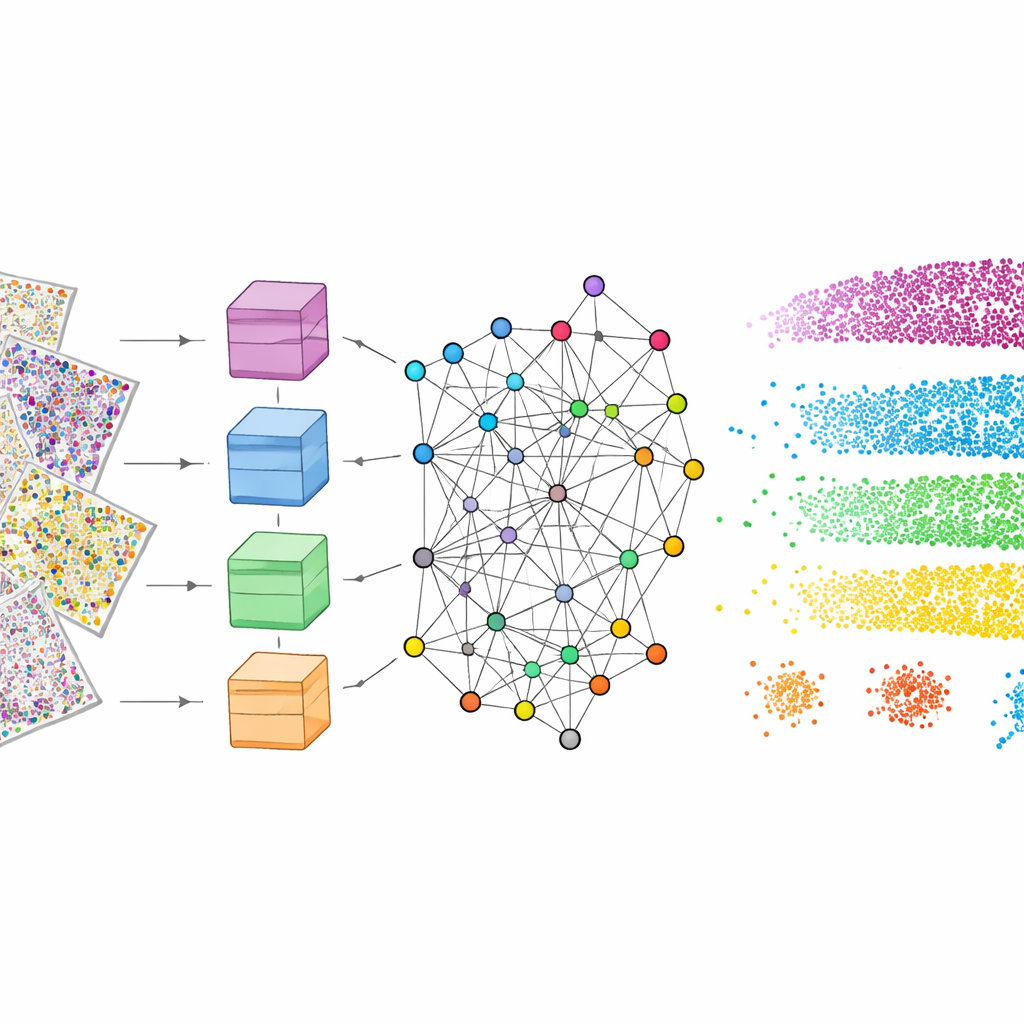
A New Way to Explore the Living Map
In essence, mViSE turns enormous, many-colored brain images into an interactive map that researchers can search visually. Instead of laboriously scripting every analysis, scientists can click on cells or microenvironments that catch their eye and instantly see where similar structures appear and how their protein profiles compare. The method requires no manual annotation during training and scales to very large datasets, making it a practical addition to the spatial proteomics toolkit. As imaging technologies add even more molecular channels, and as similar approaches are extended to diseased brains and other organs, tools like mViSE could help convert raw image complexity into intuitive, navigable views of tissue architecture—bringing us closer to reading the brain’s intricate molecular atlas like a searchable map.
Citation: Huang, L., Mills, R., Mandula, S. et al. mViSE: A visual search engine for analyzing multiplex IHC brain tissue images (spatial proteomics). Sci Rep 16, 10245 (2026). https://doi.org/10.1038/s41598-026-40620-5
Keywords: spatial proteomics, brain imaging, visual search engine, multiplex microscopy, computational pathology