Clear Sky Science · en
Benchmarking multiple instance learning architectures from patches to pathology for prostate cancer detection and grading using attention-based weak supervision
Why this matters for patients and doctors
Prostate cancer is one of the most common cancers in men, and deciding how serious it is from microscope slides directly affects treatment choices. Today, this grading work is slow, can vary between pathologists, and is hard to scale as populations age. This study shows how modern artificial intelligence can learn to judge cancer severity from digital slides using far fewer human markings than before, while still remaining understandable to doctors. That makes it a realistic candidate to support everyday diagnosis in real hospitals rather than just research labs.
From giant slides to tiny image tiles
Modern scanners turn a thin slice of prostate tissue into a gigantic digital picture, often 20,000 by 20,000 pixels in size. Most of that image is background, and even the useful tissue is far too large to feed into a typical computer model all at once. The researchers solved this by cutting each whole slide into many small square "patches" of tissue, like turning a poster into a jigsaw puzzle. They carefully detected just the true tissue regions, filtered out empty areas and artifacts, and generated coordinate maps telling the computer exactly where to sample. They tested four patch strategies, mixing large and small squares and with or without overlap, eventually producing more than 31 million patches from over 10,600 slides in a large public prostate cancer dataset called PANDA. 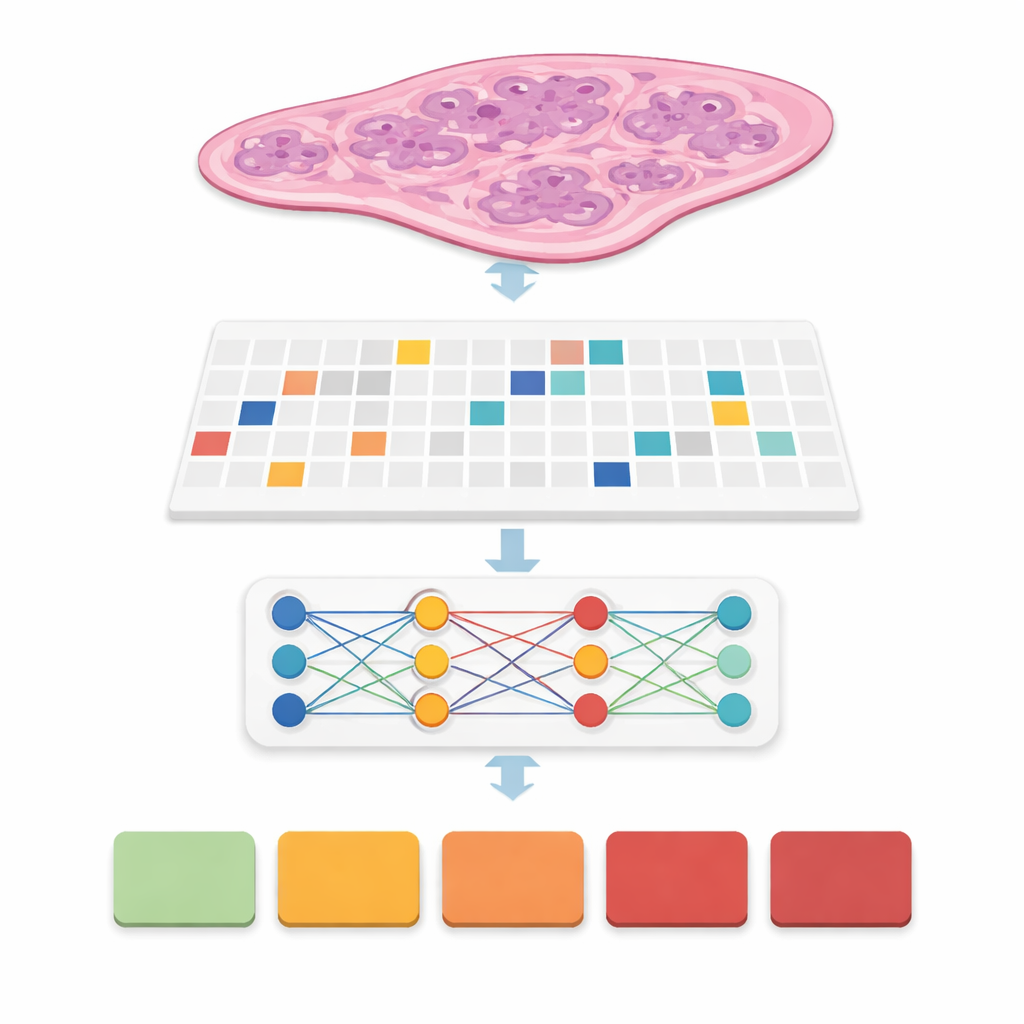
Teaching computers to see cancer without pixel-by-pixel labels
In many previous systems, experts had to draw precise outlines around every cancer focus, a time‑consuming step that makes real‑world deployment difficult. Here, the team used an approach called weak supervision: the computer only sees a slide‑level label, such as the final cancer grade group from 0 (benign) to 5 (most aggressive), without any detailed markings inside. Each slide becomes a "bag" of patches, and special models learn which patches matter most for predicting the overall grade. This family of methods, called multiple instance learning, lets the computer discover patterns of glands and cells that distinguish mild from aggressive disease while greatly reducing the workload on pathologists during training.
Comparing many AI building blocks head‑to‑head
Rather than proposing just one new model, the authors built a careful benchmark. They combined six leading multiple‑instance learning designs with three different feature extractors, or "encoders," that turn each patch into a rich numerical fingerprint. One encoder was a classic vision model trained on everyday photos, while the others were large "foundation" models trained specifically on hundreds of thousands of pathology slides. They ran all of these combinations under four patch settings and five cross‑validation folds, yielding 360 training runs and 72 core configurations. Performance was measured with accuracy, several types of F‑scores, and quadratic weighted kappa, a statistic that rewards getting the cancer grade close to the expert’s decision even when it is not perfectly exact.
Finding the sweet spot: small overlapping patches and a pathology foundation model
The clear winner emerged when the team used relatively small tissue patches (256 by 256 pixels) with 50 percent overlap, encoded by the UNI2 foundation model and aggregated by a low‑rank attention method called ILRA‑MIL. This combination reached about 79 percent accuracy and a very high quadratic weighted kappa of just over 0.90, approaching agreement levels seen between human experts in the original PANDA challenge. Smaller, overlapping patches gave the model both fine cellular detail and enough context, while the pathology‑specific encoder clearly outperformed the generic photo‑trained network by 15–20 percentage points in accuracy. Importantly, the authors showed that these gains are achievable with accessible cloud resources, using distributed computing on public platforms rather than expensive dedicated clusters. 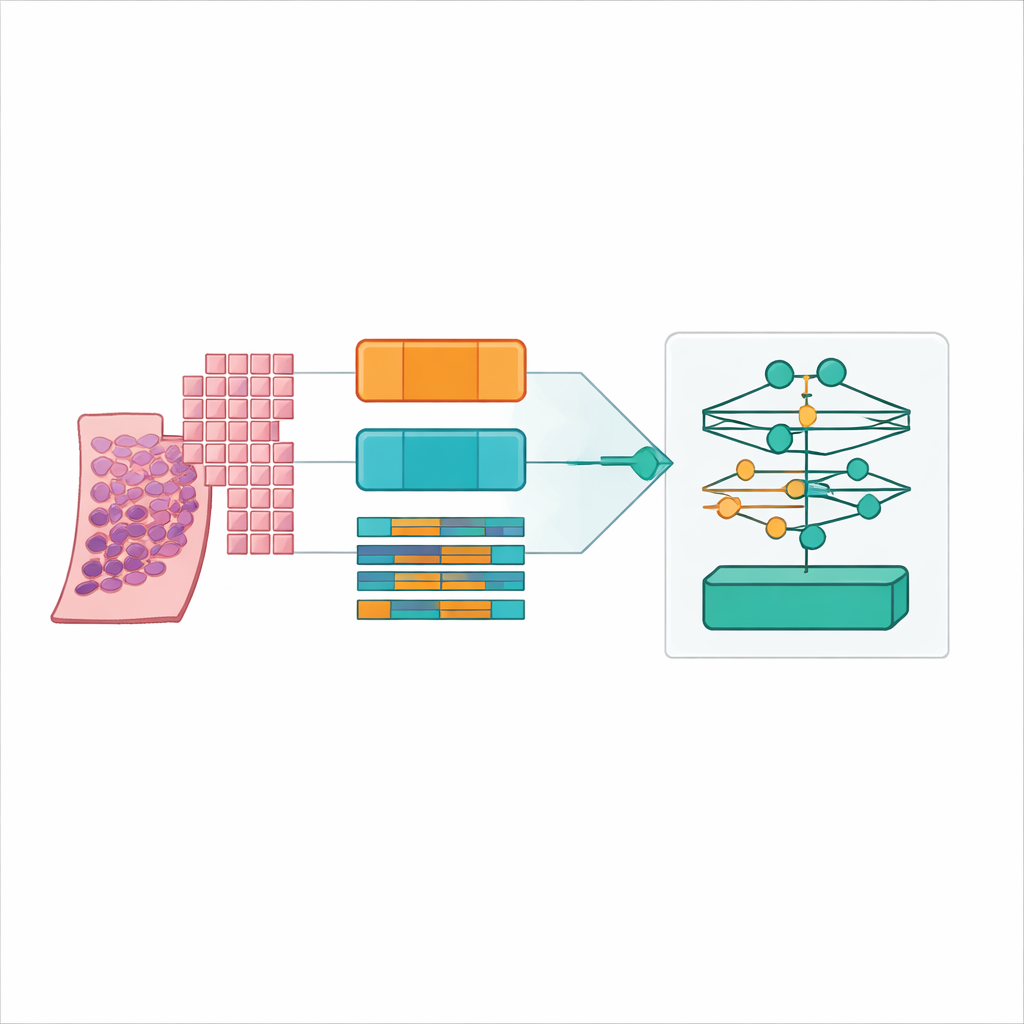
Making AI decisions visible to pathologists
For any tool that may influence treatment, it is not enough to be accurate; doctors need to understand why it makes a particular call. The researchers built in attention maps and Grad‑CAM heatmaps that highlight which regions of a slide drove the prediction. These visual explanations can be overlaid on the tissue image so that a pathologist can confirm that the model is truly focusing on cancerous glands or aggressive patterns rather than noise or benign structures. This attention to interpretability, combined with a web‑based interface and a fully described workflow from raw slide to prediction, is intended to ease adoption and independent validation in clinical laboratories.
What this means for future prostate cancer care
In plain terms, the study shows that a carefully designed AI system can grade prostate cancer on digital slides with performance close to that of specialist pathologists, without demanding exhaustive manual drawing on every training slide. The most effective recipe uses many small, overlapping image tiles and a large pathology‑trained encoder feeding into an attention‑based model that can point back to the tissue it relied on. While the work still depends on a single major dataset and needs broader hospital‑to‑hospital testing, it offers a realistic path toward faster, more consistent, and more widely available prostate cancer diagnosis, especially in regions where expert pathologists are scarce.
Citation: Butt, N.A., Sarwat, D., Noya, I.D. et al. Benchmarking multiple instance learning architectures from patches to pathology for prostate cancer detection and grading using attention-based weak supervision. Sci Rep 16, 11535 (2026). https://doi.org/10.1038/s41598-026-39196-x
Keywords: prostate cancer, digital pathology, weakly supervised learning, multiple instance learning, computational pathology