Clear Sky Science · en
A Subphase-Labeled Mitotic Dataset for AI-powered Cell Division Analysis
Why counting dividing cells matters
When doctors look at cancer under the microscope, one of the most important clues they use is how many cells are in the act of dividing. Fast‑dividing tumors often behave more aggressively and can demand different treatment. Yet spotting these tiny dividing cells is slow, tiring work for pathologists and even experts often disagree. This study presents a new, richly annotated image dataset and an improved artificial intelligence (AI) method designed to help computers find and understand these dividing cells more reliably across many cancer types and hospitals.
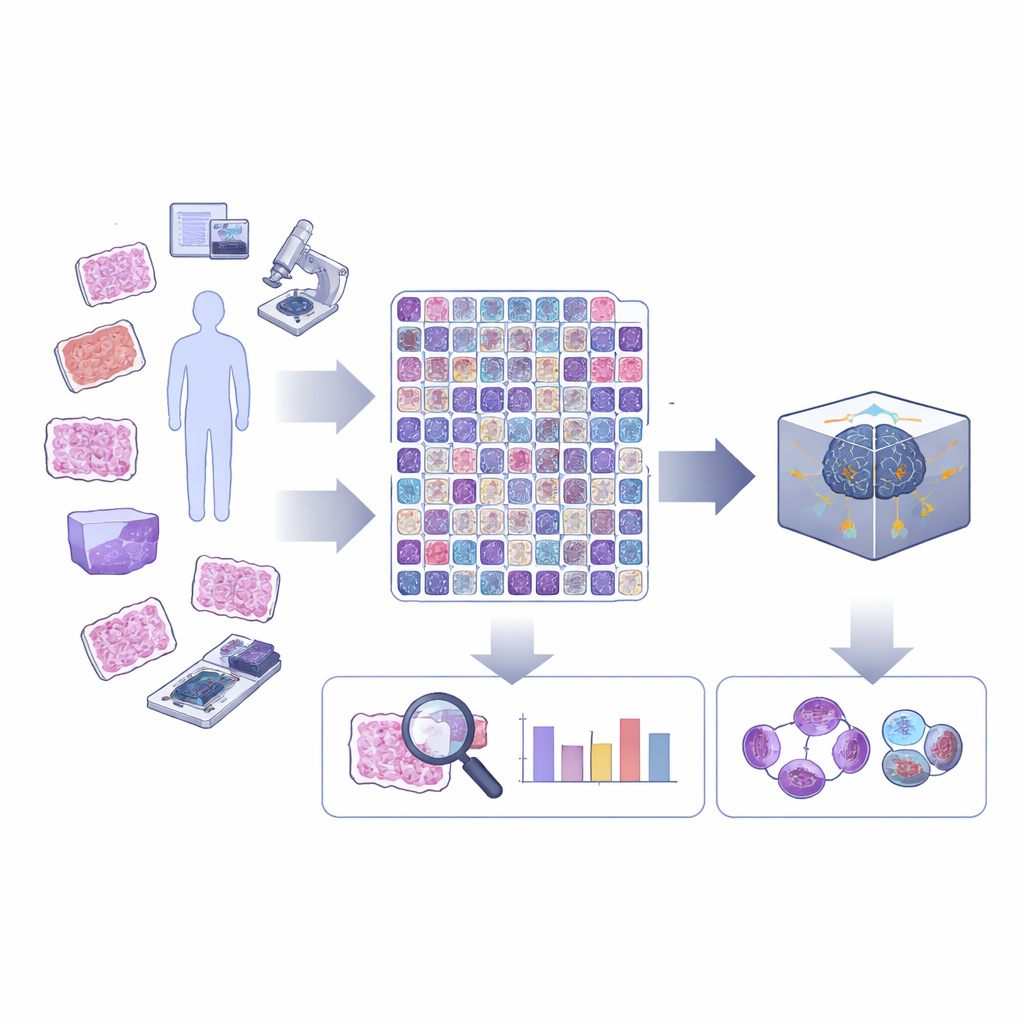
A closer look at cell division in cancer tissue
Cell division, or mitosis, unfolds in a series of stages as the cell’s genetic material is duplicated and pulled apart. In routine cancer diagnosis, most computer tools simply mark “mitotic figures” as a single category, ignoring which stage they are in or whether they look abnormal. However, unusual or “atypical” divisions, and the balance between early and late stages, can carry important information about how dangerous a tumor is. The authors argue that AI will only reach its full potential in pathology if it can see these finer distinctions that human specialists already look for under the microscope.
Building a richer library of images
To move beyond a simple yes‑or‑no view of dividing cells, the team extended an existing large benchmark known as MIDOG++, which already covered multiple tumor types, species, and scanning devices. Expert annotators revisited more than ten thousand mitotic figures and assigned each one to one of the five main stages of normal cell division, plus a separate class for atypical mitoses. They also drew precise outlines around each cell, not just rough boxes. This careful work turns the dataset into a resource that can support both more exact AI training and future studies of cell shape and structure during division.
Adding a new lung cancer dataset
Recognizing that AI systems often stumble when moved from one hospital or organ type to another, the researchers also created a new dataset from lung adenocarcinoma, a common and deadly form of lung cancer. Instead of focusing only on the “hottest” regions with the most activity, they selected the entire tumor area on each slide, tiled it into manageable image patches, and painstakingly labeled dividing cells, atypical divisions, and look‑alike but non‑dividing cells. This LUNG‑MITO dataset provides a tough, real‑world test of whether an algorithm trained elsewhere can still perform well when faced with new tissue and scanner conditions.
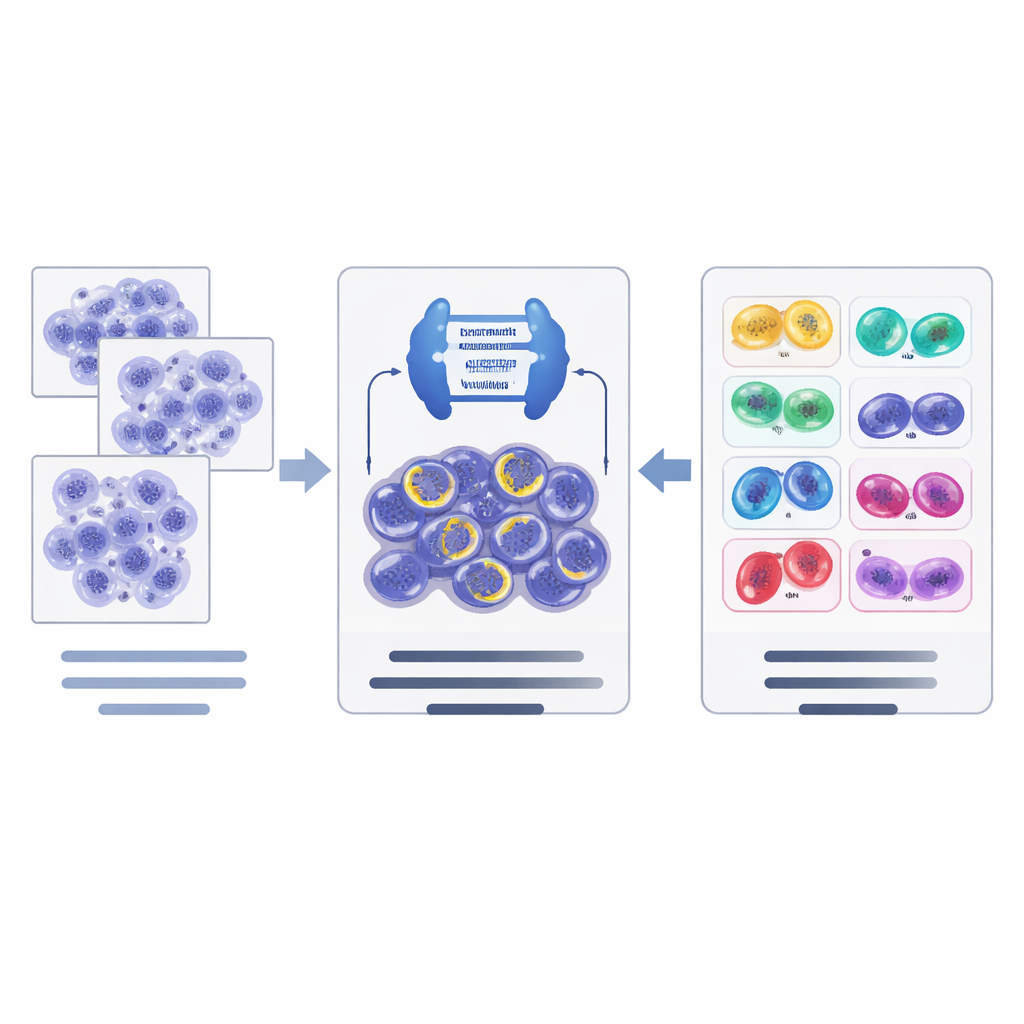
How the new AI pipeline works
On top of these data resources, the authors designed a two‑step AI pipeline. First, a powerful image segmentation network, based on a modern architecture called ConvNeXt combined with Mask R‑CNN, scans tissue tiles and proposes candidate dividing cells, outlining each one and giving an initial guess of its stage. Second, a separate classification network (EfficientNet) takes a closer look at these candidates and refines the decision in a hierarchical way: it first judges whether a candidate is truly dividing or just an imposter, and then, for confirmed mitoses, chooses the most likely stage. The system uses data augmentation tricks tailored to pathology images to help it cope with differences in staining and scanners across laboratories.
Testing performance across different settings
To fairly judge how well their approach works, the researchers followed the same evaluation rules used in previous mitosis detection challenges. They measured how often the algorithm’s predicted cells matched expert‑drawn cells within a small distance, accounting for both missed detections and false alarms. Training was done on a portion of the extended MIDOG++ data, while the new LUNG‑MITO slides were held back as a separate test to probe how robust the method is to changes in tumor type and imaging hardware. The enhanced backbone and refinement step clearly improved performance over a more traditional system, boosting both overall detection and the accuracy for each subtype of cell division.
What this means for cancer care and research
For non‑specialists, the main message is that the study delivers both a detailed public dataset and a stronger AI model for recognizing how and when cancer cells divide. By teaching algorithms to tell apart different stages of mitosis and to flag atypical divisions, and by showing that this can be done more reliably across diverse tumors and scanners, the work lays the groundwork for future tools that support pathologists in grading tumors and studying their biology. In the long run, such tools could help make cancer diagnoses more consistent from one hospital to another and open new research directions that link the appearance of dividing cells to their molecular behavior and patient outcomes.
Citation: Ivan, Z.Z., Hirling, D., Grexa, I. et al. A Subphase-Labeled Mitotic Dataset for AI-powered Cell Division Analysis. Sci Data 13, 680 (2026). https://doi.org/10.1038/s41597-026-07007-7
Keywords: digital pathology, cell division, mitosis detection, cancer imaging, medical AI