Clear Sky Science · en
Human neuronal differentiation under Aβ exposure: a single-cell transcriptomic and epigenomic dataset
Why this brain cell study matters
Alzheimer’s disease slowly robs people of memory and thinking, but we still do not fully understand what happens to newborn brain cells as the disease unfolds. This study creates a detailed “atlas” of how human nerve cells develop in the lab when they are exposed to amyloid‑beta, a small protein fragment that builds up in Alzheimer’s brains. By tracking thousands of cells one by one, and looking not only at which genes are active but also how the DNA is packaged, the researchers offer a rich reference map that other scientists can use to probe early changes linked to dementia.
Building brain cells in a dish
To study early nerve cell development in a controlled way, the team used human induced pluripotent stem cells, which can be coaxed to become neural progenitor cells—the starting material for many brain cell types. These progenitors were grown in dishes and then switched into a special medium that encourages them to turn into more mature neurons over 20 days. Some dishes followed this program under normal conditions, while others were exposed from the start to amyloid‑beta 1–42, the same peptide that forms plaques in Alzheimer’s disease. Cells from both groups were collected at four key moments—before the change (day 0) and during the transformation (days 7, 13, and 20)—to capture the step‑by‑step unfolding of their developmental paths. 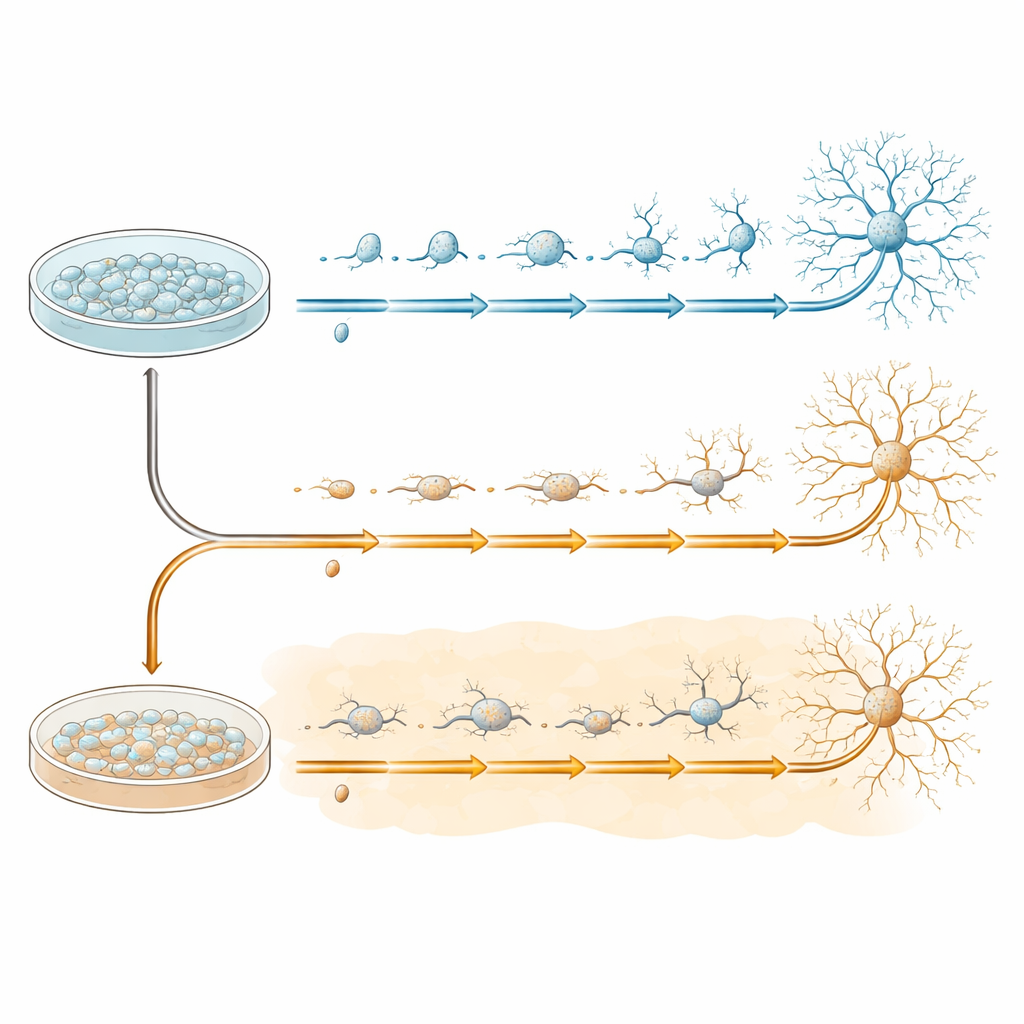
Listening to genes and opening the genome
What makes this work stand out is that each cell was examined in two complementary ways. First, the researchers used single‑cell RNA sequencing to measure which genes were turned on in more than 42,000 cells. Second, they used single‑cell ATAC sequencing in over 30,000 cells to see which parts of the DNA were physically accessible, a sign that these regions can be used as switches to control genes. Sophisticated computer methods grouped cells into clusters, ordered them along developmental “trajectories” from progenitors to more mature neurons and astrocyte‑like cells, and linked patterns of gene activity to changes in chromatin, the DNA‑protein material that packages our genes.
Tracing the journey from stem cells to neurons
Within the developing neuronal cells, the team identified a series of five broad stages, labeled A through E, that reflect a gradual shift from dividing stem‑like cells toward maturing neurons with complex branches. Classic marker genes for early and late neuronal states changed in an orderly fashion along this path, and an independent “pseudotime” analysis confirmed the same sequence. At later stages, they zoomed in further and found sub‑paths among more mature cells, showing that some cells follow slightly different routes as they refine their identity. Across these steps, the scientists cataloged genes that change together, genes that respond differently when amyloid‑beta is present, and sets of genes that seem to be altered only under amyloid‑beta exposure.
What amyloid‑beta changes—and what it leaves intact
Although the overall structure of the developmental paths was similar with and without amyloid‑beta, the team detected distinct shifts in which genes were more or less active at particular stages when the peptide was present. Many of these genes are involved in nerve cell communication, growth of axons and dendrites, and other functions critical for healthy brain circuits. The researchers also combined the gene‑activity and chromatin data to build networks that link key control proteins, called transcription factors, to sets of target genes. They discovered thousands of such “regulons” and showed that some of them behave differently between the normal and amyloid‑beta conditions, even though large‑scale DNA accessibility changes remain modest. 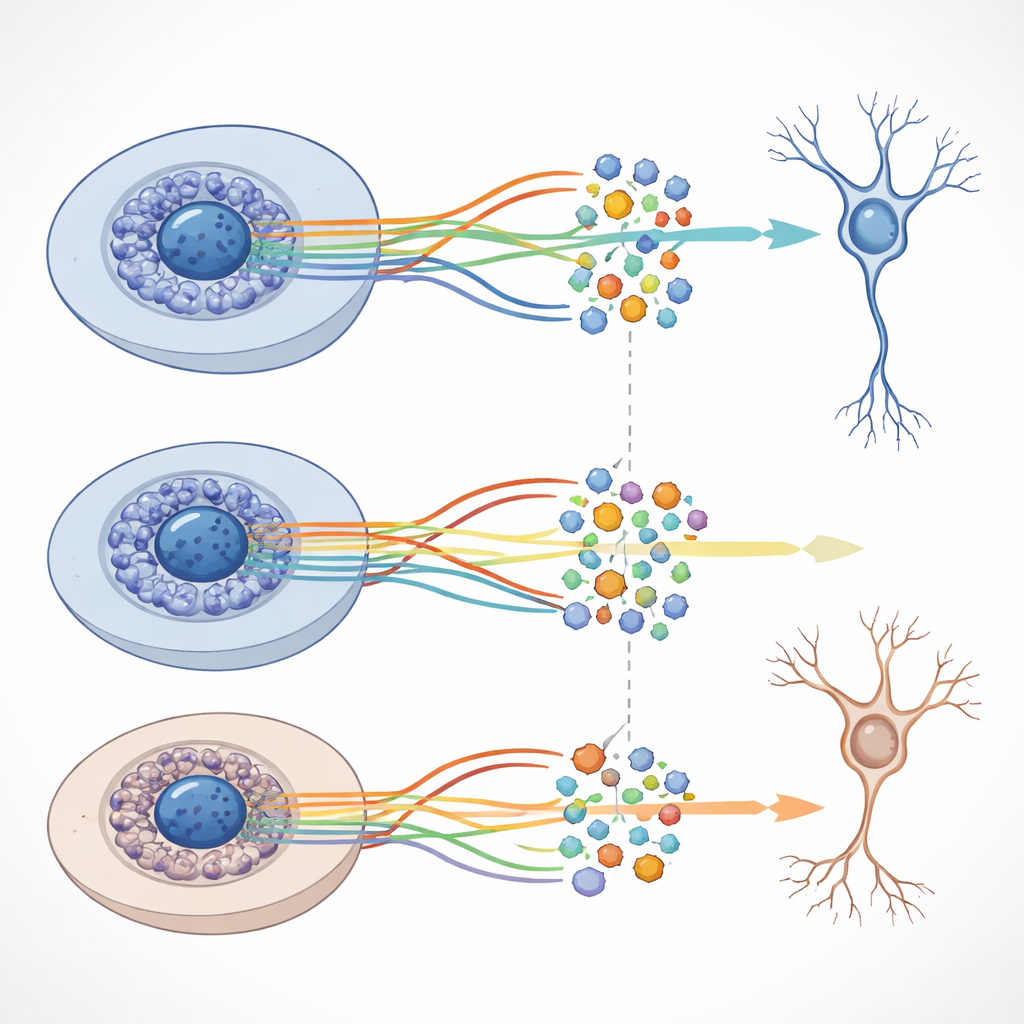
Connecting lab findings to human Alzheimer’s brains
To see how relevant their cell‑culture system is to real disease, the authors compared their gene lists to an existing dataset of brain tissue from people with and without Alzheimer’s. Several of the same genes that differed between normal and amyloid‑beta‑treated cells also differed between control and Alzheimer’s brains. When they treated groups of development‑stage‑specific genes as “signatures,” these signatures were enriched among the changes seen in patient hippocampus samples, especially for genes tied to neuronal communication and structure. This suggests that the in‑vitro model captures meaningful pieces of the biology occurring in the vulnerable memory centers of the aging brain.
What this resource offers for the future
The main product of this work is not a single headline result but a public, reusable dataset. All raw and processed information, along with analysis code and an interactive web browser, are freely available for other scientists. Researchers can use this map to benchmark new computational tools, explore how different lab protocols affect neuron development, or test new ideas about how Alzheimer’s‑related processes disrupt the birth and maturation of brain cells. In simple terms, the study provides a detailed “wiring diagram” of how young human brain cells grow—with and without a key Alzheimer’s‑linked stressor—that will help the field dissect when and where things go wrong.
Citation: Blanco-Luquin, I., Martínez-de-Morentin, X., Vilas-Zornoza, A. et al. Human neuronal differentiation under Aβ exposure: a single-cell transcriptomic and epigenomic dataset. Sci Data 13, 638 (2026). https://doi.org/10.1038/s41597-026-06971-4
Keywords: Alzheimer’s disease, neurogenesis, single-cell sequencing, amyloid beta, neuronal differentiation