Clear Sky Science · en
Structural insight into IscB’s RNA-lid-based inactivation mechanism
Smaller Gene Scissors with a Built-In Safety Lock
Gene-editing tools like CRISPR–Cas9 have transformed biology, but their large size makes them hard to deliver safely into human tissues. This study explores IscB, a much smaller cousin of Cas9, to understand how it stays safely switched off until it meets exactly the right DNA sequence. By watching IscB’s tiny moving parts at near-atomic detail, the authors reveal how it guards against stray cuts and how its performance can be tuned for future medical applications. 
A Tiny Ancestor of CRISPR’s Workhorse
IscB comes from ancient mobile genetic elements and is thought to be an evolutionary ancestor of Cas9. Like Cas9, it uses an RNA guide to home in on matching DNA and then slice it, but it is less than half the size of the standard Cas9 used in laboratories. That compactness is attractive for therapies that rely on viral vectors, which have strict cargo limits. Until now, however, researchers had only seen one snapshot of IscB bound to DNA. The key missing pieces were how the protein looks when it is idle, how it tests potential targets, and how it flips from a harmless bystander into an active cutter.
Catching the Protein in the Act
To fill in this picture, the team engineered a slightly modified, more stable version of IscB and examined it using cryo–electron microscopy. They captured four distinct states: a resting form with no DNA, two intermediates where the RNA guide pairs with just 6 or 10 DNA letters, and a primed state where 16 letters are paired and the DNA is ready to be cut. These high-resolution images trace a continuous pathway from fully off to fully on. They show that IscB’s two cutting centers are packed unusually close together and connected by short “hinges” rather than long floppy linkers, helping explain how such a small protein can carry out a complex task.
RNA Lids and a Molecular Car Pedal
The structures reveal that IscB uses an elegant double-lock system made from its own RNA. One part of the RNA folds over the first cutting center like a lid, while another section bends across the second, blocking access to stray DNA strands. As the RNA guide gradually pairs with a correct DNA target, it does not move all at once; instead it shifts in stages, like a car pedal being slowly pressed. When about 11 matching letters are paired, this motion finally pushes the RNA lids out of the way and reshapes a key region called the HNH domain, which swings by roughly 90 degrees into an active position. This threshold helps ensure IscB responds only to strongly matching sequences, limiting off-target cuts. 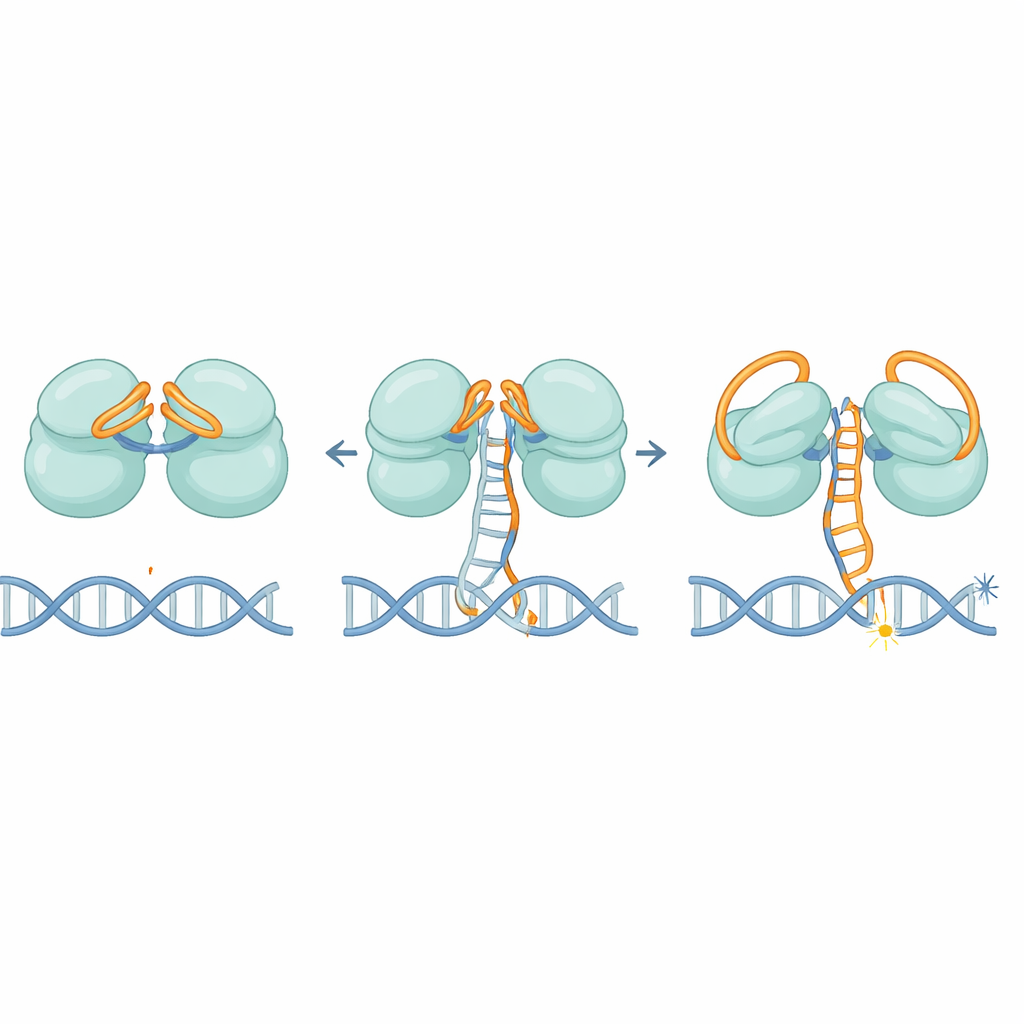
How IscB Pries Open DNA and Holds It Steady
Once activated, IscB must separate the two strands of the DNA double helix and keep them apart long enough to cut. The authors identify a small loop on the protein that wedges into the DNA, forcing the strands apart in the region where the RNA and DNA form a hybrid structure. Positively charged patches along the hinges then grip the displaced strand, stabilizing this bubble-like “R-loop.” Mutations that weaken this wedge or the charged band sharply reduce DNA-cutting activity, confirming that they are essential for both opening and holding the DNA in place.
Tuning the Hinges to Build Better Editors
Because the hinges control how easily the HNH domain can swing into action, the researchers tested whether subtle changes there could boost performance. By replacing certain bulky amino acids with smaller, more flexible ones, they created IscB variants whose hinges move more freely. In human cells, two of these engineered versions, dubbed Hig1 and Hig2, showed noticeably higher editing activity. One variant approached the efficiency of an advanced Cas9 tool while causing fewer unintended DNA breaks across the genome, suggesting that carefully loosening these hinges can make IscB both more powerful and more precise.
Why This Matters for Future Gene Therapies
Together, these findings show how a compact gene-editing enzyme can stay safely locked until it encounters a near-perfect DNA match, then rapidly rearrange itself to cut at the right spot. By mapping each step—from RNA lids that shield the cutting centers to the car-pedal–like motion that triggers activation and the hinges that steer the final slice—the work offers a blueprint for designing smaller, safer genome editors. Such tools could be easier to package into medical delivery systems and may ultimately enable more precise treatments with fewer side effects.
Citation: Wang, F., Guo, R., Zhang, S. et al. Structural insight into IscB’s RNA-lid-based inactivation mechanism. Nat Struct Mol Biol 33, 603–614 (2026). https://doi.org/10.1038/s41594-026-01761-3
Keywords: IscB, genome editing, CRISPR, RNA-guided nuclease, structural biology