Clear Sky Science · en
Technical and biological sources of noise confound multiplexed enhancer AAV screening
Finding the Right Genetic Keys
Scientists increasingly rely on gene-delivering viruses to turn specific brain cell types on or off, with hopes for both basic research and future therapies. But to make these tools precise, researchers must attach the right “on switches,” called enhancers, so each virus activates only in its intended cell type. This study shows that when many of these enhancer-equipped viruses are tested together in a single experiment, hidden technical and biological noise can badly blur the results, making promising tools look far less accurate than they really are.
Why Cell-Specific Gene Control Matters
Enhancers are short stretches of DNA that help turn genes on in particular cell types or brain regions. By coupling these enhancers to adeno-associated viruses (AAVs), scientists can deliver fluorescent markers or therapeutic genes specifically to, say, a certain kind of neuron or supporting glial cell. Traditionally, each enhancer-AAV combination is tested one at a time in animals—slow, expensive, and difficult to scale. Multiplexed approaches promise a shortcut: mix many differently barcoded enhancer-AAVs together, inject them into a mouse, then use single-cell RNA sequencing to read out which enhancer was active in which cell. In theory, this should reveal dozens of enhancer behaviors in a single experiment.
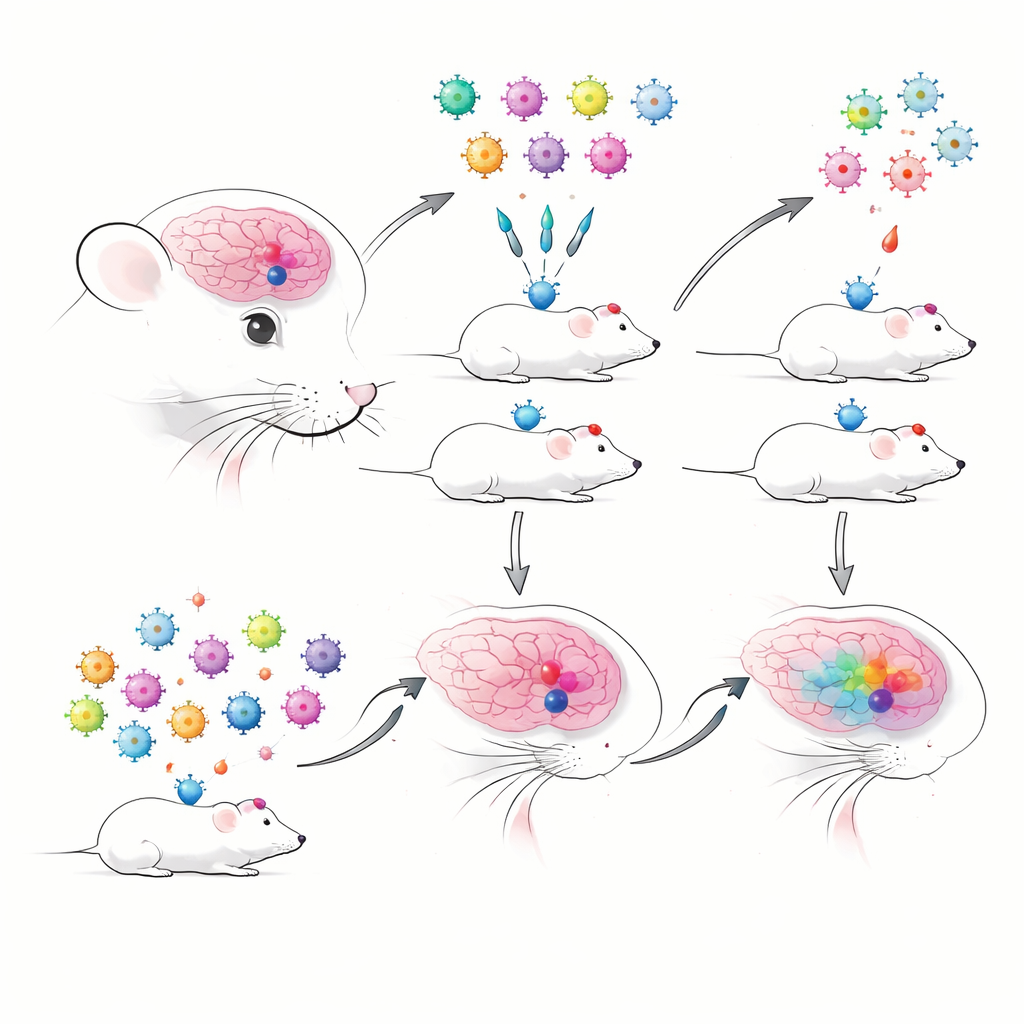
When Many Experiments Share One Mouse
The authors began by building barcoded AAVs that carried previously well-characterized enhancers known to label specific brain cell types with high accuracy. Each enhancer drove a fluorescent protein and a short RNA barcode tag, allowing tracking by sequencing. They compared traditional one-by-one injections with pooled “multiplex” injections, where many enhancer-AAVs were delivered at once and read out using droplet-based single-cell RNA sequencing. Right away, they ran into a practical hurdle: in nuclei-based preparations—useful for frozen or delicate tissues—the viral barcodes were detected very poorly, whereas whole-cell preparations captured viral transcripts much more reliably. This forced them to favor whole cells, which can be more fragile and may underrepresent some cell types.
Hidden Mix-Ups in Viral Genomes
Even with better transcript capture, multiplex screens did not reproduce the crisp, cell-specific patterns seen in one-by-one tests. Many enhancers appeared far less selective, especially those that were naturally weaker or targeted rare cell types. To investigate, the team compared viruses that were packaged individually and then mixed, versus viruses packaged together from a single plasmid pool. Using long-read sequencing, they discovered that pooled packaging produced a high rate of “chimeric” genomes—viral particles in which enhancer and barcode had been scrambled and mismatched. These chimeric particles add misleading barcode signals and lower apparent specificity. The authors also explored a dual-barcoding system (“scQers”), where each virus carried both an expression-linked barcode and a separate constitutive barcode. Matching these pairs allowed some erroneous reads to be filtered out and improved specificity, but not enough to recover the performance of one-by-one validation.
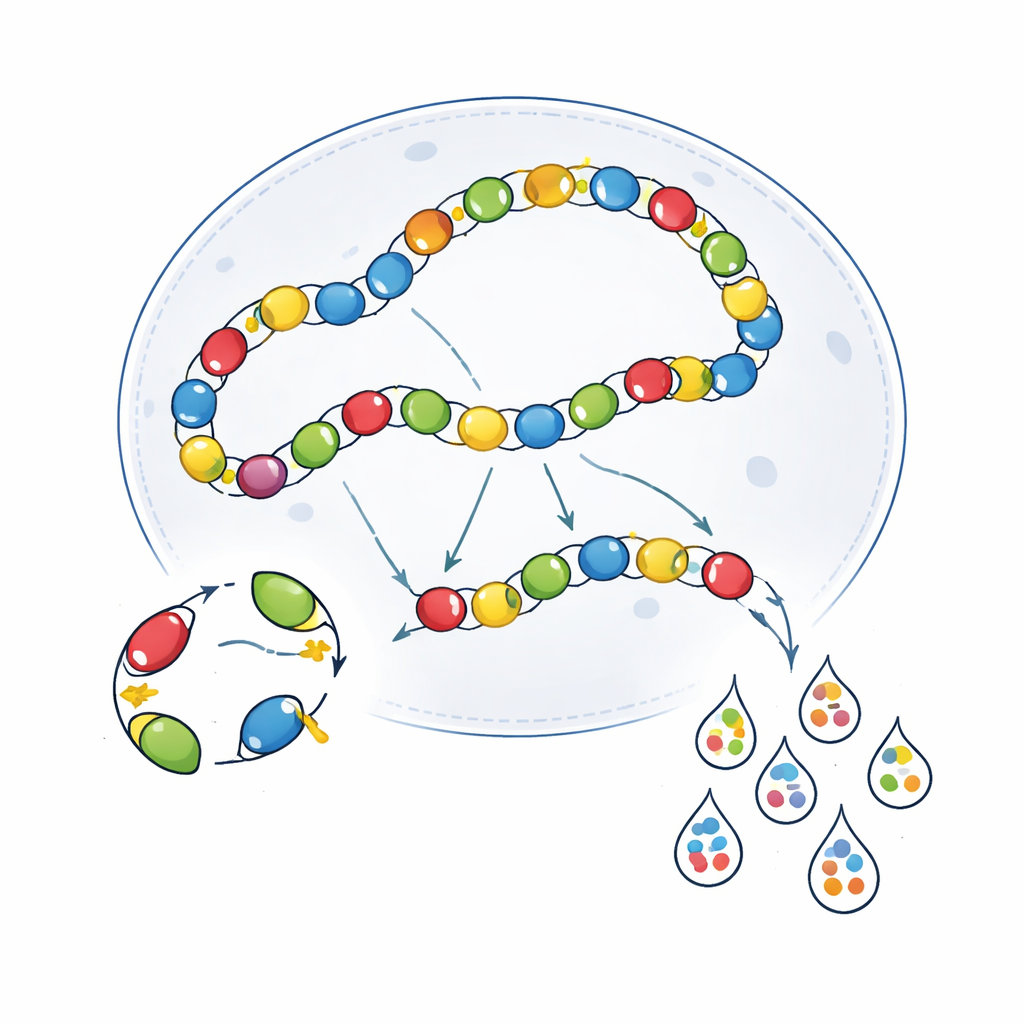
Biological Crosstalk Inside Cells
Technical artifacts were only part of the problem. The viruses themselves interact inside cells in ways that blur the picture. Once inside the nucleus, AAV genomes often link end to end, forming long chains. In this arrangement, an active enhancer on one viral genome can inadvertently drive expression from a neighboring genome—a phenomenon called transcriptional crosstalk. The authors demonstrated that when two different enhancer-AAVs were co-injected, they could spur each other’s activity in cell types where one alone would normally be silent. Using immunodeficient SCID mice, which have reduced AAV concatenation, partially reduced this overlap for some enhancers but also weakened overall expression, and did not consistently improve specificity across the board. Attempts to insulate viral genomes with barrier DNA elements likewise showed only context-dependent benefits, helping in some brain regions but not others.
Why Noise Hits Some Enhancers Harder
By comparing many enhancers side by side, the study found that those targeting abundant cell types and producing strong fluorescent signals fared better under multiplexed conditions. Weak enhancers or those active in rare cell populations were disproportionately drowned out by background noise from chimeric genomes, crosstalk, and stray barcodes. Importantly, counts of barcode transcripts in multiplex screens did not correlate well with the protein levels measured when each enhancer was tested alone, suggesting that much of the barcode signal in pooled experiments does not faithfully reflect true enhancer-driven activity.
What This Means for Future Brain Tools
For non-specialists, the bottom line is that combining many enhancer-equipped viruses into a single test animal is more complicated than it appears. The viruses can rearrange during production, talk to each other inside cells, and generate misleading barcode signals that make precise enhancers look sloppy. The authors conclude that current multiplexed enhancer AAV screening cannot yet replace careful one-by-one validation, especially when searching broadly for new, finely targeted tools. However, small and thoughtfully designed pools may still be useful for ranking similar enhancers or optimizing designs before committing to full validation. Substantial technical and biological improvements will be needed before large, unbiased pooled screens can reliably reveal the best genetic “keys” for targeting specific brain cell types.
Citation: Hunker, A.C., Mich, J.K., Taskin, N. et al. Technical and biological sources of noise confound multiplexed enhancer AAV screening. Nat Commun 17, 3738 (2026). https://doi.org/10.1038/s41467-026-72147-8
Keywords: enhancer AAV, gene delivery, single-cell RNA sequencing, viral barcoding, brain cell types