Clear Sky Science · en
Proviral NUP153 binding to viral proteins and RNA regulates structural–nonstructural protein ratios in orthoflavivirus infection
How Viruses Turn a Cell’s Gatekeeper into a Helper
Many dangerous mosquito- and tick-borne viruses, including those that cause dengue, Zika, West Nile fever, and tick-borne encephalitis, still have no specific treatments. This study reveals how these viruses hijack a human protein called NUP153—normally part of the cell’s gatekeeping system around the nucleus—to boost their own production. Understanding this takeover at a molecular level could open the door to broad new antiviral strategies that target the host instead of the virus.
A Family of Viruses Without a Cure
Orthoflaviviruses are small RNA viruses carried by mosquitoes and ticks that can cause severe brain infections, hemorrhagic disease, and lifelong complications. Their genome is a single long strand of RNA that is read into one giant polyprotein, which is then cut into ten pieces: three that form the virus shell and seven that act as internal tools for copying and assembling new particles. To multiply, these viruses remodel the cell’s endoplasmic reticulum (ER)—a membrane network near the nucleus—into a specialized “amplification site” where viral RNA is copied, proteins are made, and new virus particles are assembled. Because no approved antiviral drugs exist for this group, researchers are increasingly looking at essential host factors that viruses rely on as potential drug targets.
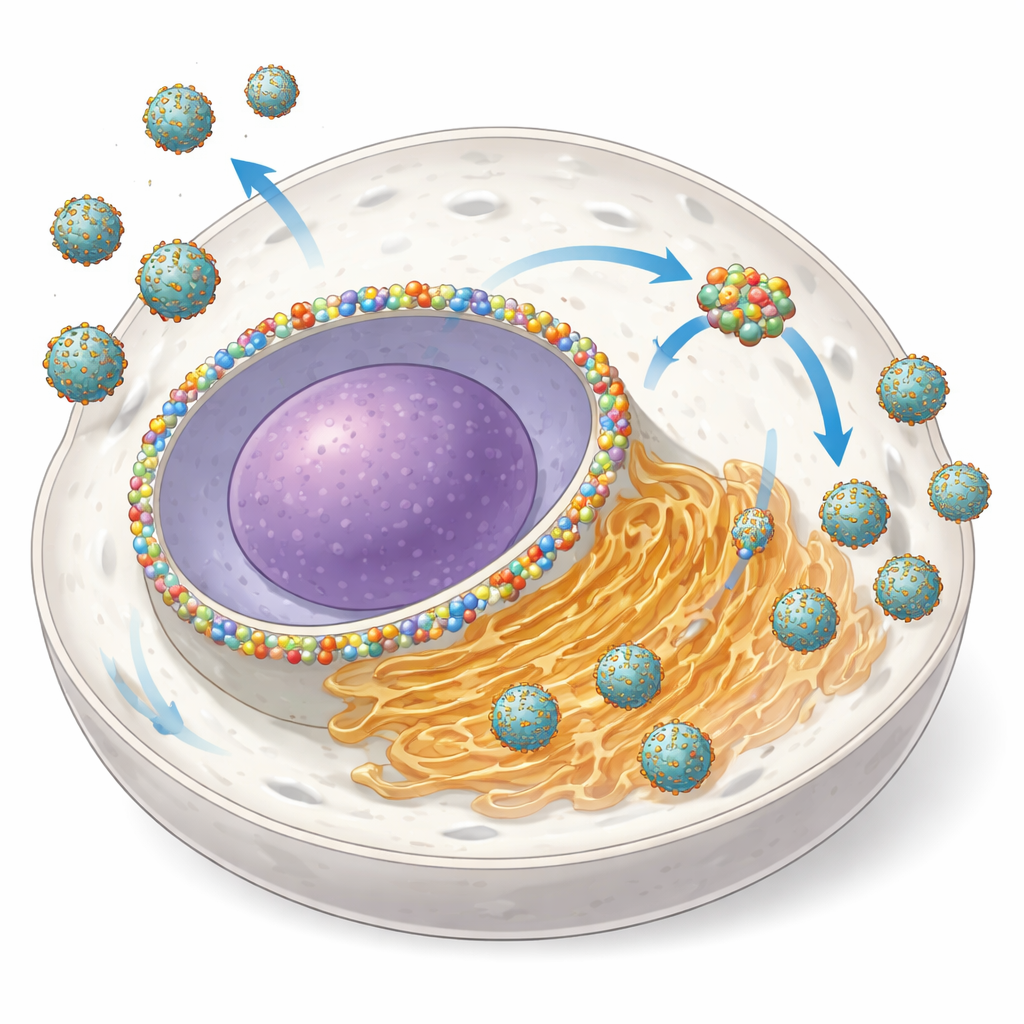
Turning a Nuclear Gate into a Virus Workshop
NUP153 is usually a component of the nuclear pore complex, a huge structure that regulates traffic of RNA and proteins between the nucleus and the surrounding cytoplasm. The authors discovered that during infection with several orthoflaviviruses, cells increase the amount of NUP153 and redistribute part of it from the nuclear envelope to the ER-based viral amplification site. Using advanced microscopy, they showed that NUP153 clusters together with viral proteins NS3 and NS5 and with viral RNA near ER membranes, while remaining largely separate from the tiny vesicles where RNA copying itself occurs. When they rapidly degraded NUP153 in engineered cells, the number of infected cells dropped and the amount of virus released fell by up to 80–95%, especially early in infection—clear signs that NUP153 acts as a “proviral” helper.
Direct Contact with Viral Proteins and RNA
To understand how NUP153 helps the virus, the researchers pulled down NUP153 from infected cells and identified its partners by mass spectrometry and RNA sequencing. They found that NUP153 directly binds two key viral enzymes, NS3 and NS5, without needing RNA as a bridge, and also latches onto specific regions of the viral RNA itself. One of these RNA regions sits at the end of the gene segment encoding the outer shell protein E and is rich in the nucleotide guanine. This stretch is strongly conserved across many orthoflaviviruses, hinting at an important shared function. Biophysical assays showed that the RNA-binding domain of NUP153 recognizes this G-rich sequence, especially when it is not folded into a compact four-stranded “G-quadruplex” shape. The same domain is also required for NUP153 to move from the nucleus to the viral amplification site and for the virus to replicate efficiently.
Rebalancing the Viral Parts List
New virus particles can only form if the cell produces the right mix of structural and nonstructural proteins from the single viral polyprotein. The team tested whether NUP153 influences this balance. In test-tube translation experiments using viral-like RNA, adding the NUP153 RNA-binding domain changed which protein products were made: it reduced production of a downstream reporter while favoring shorter products corresponding to the structural proteins. In infected cells lacking NUP153, total viral protein levels went down, but the ratios shifted: structural proteins such as E were relatively underrepresented, while several nonstructural proteins (including NS1, NS2B, NS3, and NS5) became more abundant compared with E. This shift did not stem from faster protein decay, pointing instead to altered translation of the viral polyprotein. As a result, early in infection, viruses in NUP153-deficient cells struggled to assemble and release complete particles, delaying spread to neighboring cells.
A Shared Weak Spot Across Multiple Viruses
The authors extended their observations beyond a single virus. During infection with Zika, Japanese encephalitis, West Nile, yellow fever, and dengue viruses, NUP153 again moved toward sites of viral RNA and its removal consistently reduced both the fraction of infected cells and the number of progeny particles produced. Because the G-rich RNA motif that recruits NUP153 is conserved across this virus family, and because NUP153’s role is to fine-tune the protein mix needed for particle assembly, the study suggests a common, exploitable vulnerability. Targeting the interaction between NUP153, viral RNA, and viral proteins—without necessarily destroying NUP153’s essential housekeeping roles—could provide a new, host-directed strategy for broad-spectrum antivirals.
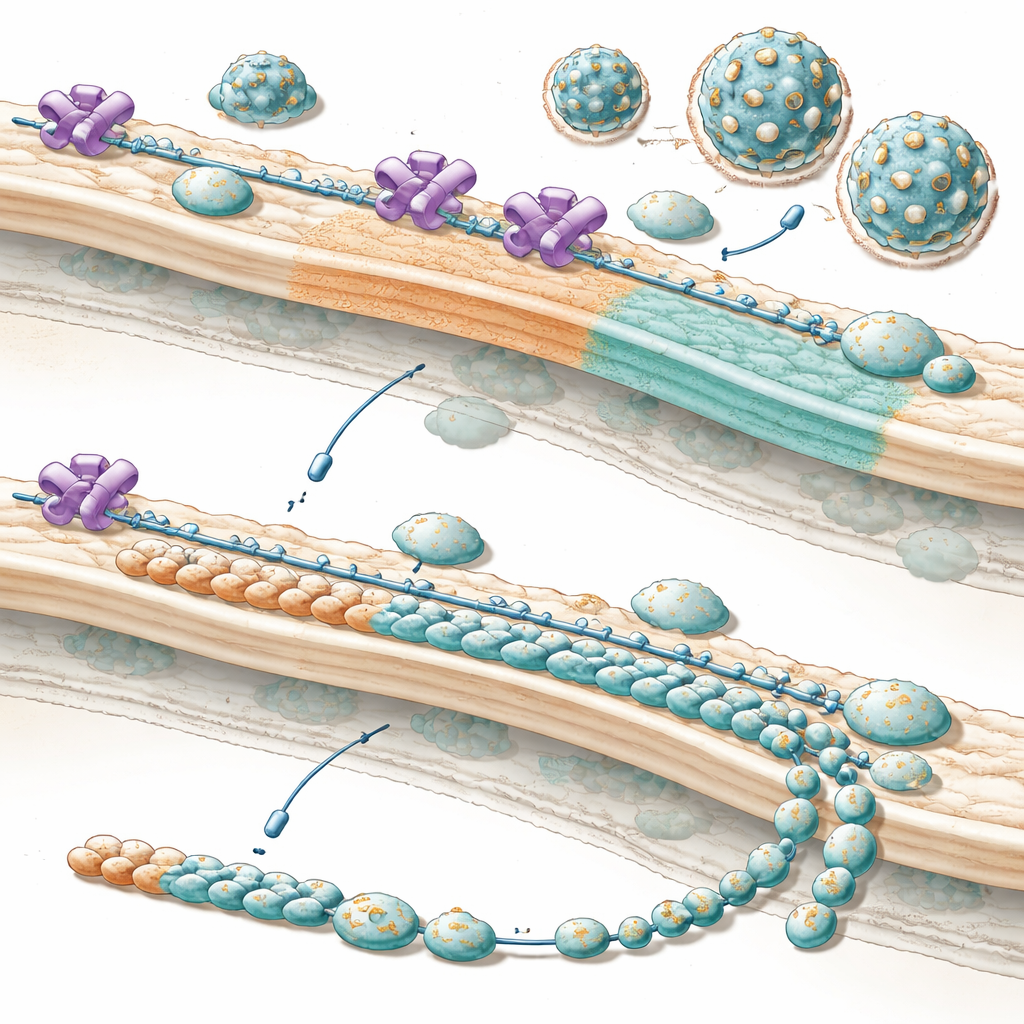
Why This Matters for Future Treatments
In everyday terms, the work shows that these viruses co-opt a cellular gatekeeper and repurpose it as a production manager, steering the cell to make more of the outer parts needed to build new virus particles at just the right time. When NUP153 is removed or blocked, the “parts list” becomes unbalanced: plenty of internal tools, but not enough shell pieces, so far fewer infectious particles emerge, especially early in infection when timing matters most. Because this strategy appears to be conserved among many medically important flaviviruses, carefully disrupting this hijacked partnership between virus and NUP153 may one day help prevent or limit severe infections from a whole family of currently hard-to-treat pathogens.
Citation: Peters, M.B.A., Lindqvist, R., Kassa, E. et al. Proviral NUP153 binding to viral proteins and RNA regulates structural–nonstructural protein ratios in orthoflavivirus infection. Nat Commun 17, 3402 (2026). https://doi.org/10.1038/s41467-026-71449-1
Keywords: orthoflavivirus, NUP153, viral protein translation, host–virus interaction, broad-spectrum antivirals