Clear Sky Science · en
NorQD AAA+ complex drives metal insertion by a twisting mechanism
How Cells Thread Metals into Key Enzymes
Many of the proteins that keep microbes – and ultimately ecosystems – running must carry tiny metal ions in just the right spot to work. This study uncovers how a specialized helper machine, built from two proteins called NorQ and NorD, physically twists and stretches another enzyme so that an iron atom can be slotted into place. Understanding this microscopic choreography helps explain how bacteria control greenhouse-gas chemistry in soils and could inspire new ways to tune metal-containing catalysts.
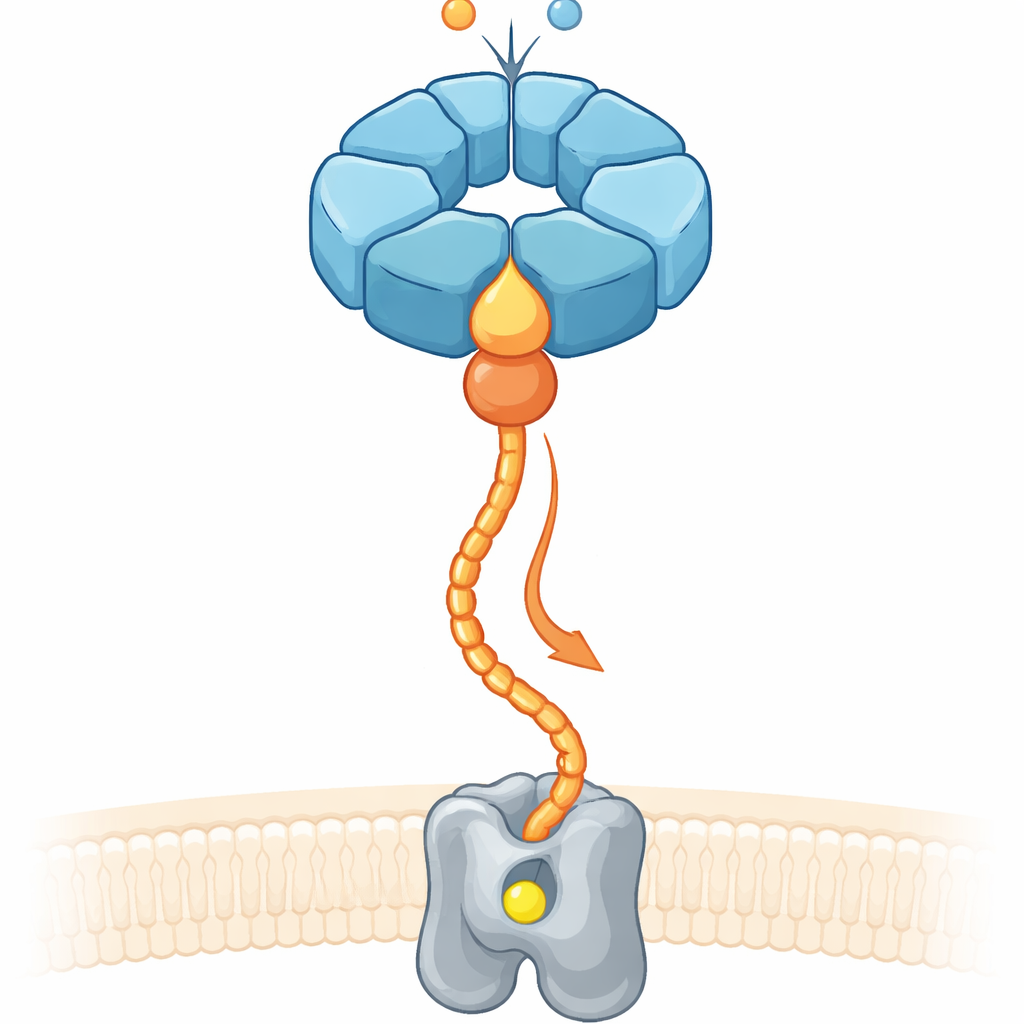
The Problem of Putting Metals in the Right Place
Roughly a third of all known proteins need metals such as iron or magnesium to function. Yet simply flooding a cell with metal ions does not guarantee that the right metal lands in the right pocket. Cells therefore rely on “chaperone” systems that bind to unfinished enzymes, reshape them, and guide metal ions into their active sites. In denitrifying soil bacteria, which convert nitrogen compounds and influence nitrous oxide emissions, a membrane enzyme called nitric oxide reductase (cNOR) requires a non‑haem iron atom (called FeB) to become active. Earlier work showed that two accessory proteins, NorQ and NorD, are essential for installing this iron, but how they cooperated and used chemical energy to do mechanical work remained unclear.
A Molecular Ring and Its Helper Partner
NorQ belongs to a large family of ring‑shaped molecular motors, known for forming six‑subunit rings that burn the energy-rich molecule ATP and pull on protein chains threaded through their central pores. NorD carries a so‑called VWA domain, a compact latching module often used by proteins to grab other partners via a metal-based contact site. Using high‑resolution cryo‑electron microscopy on NorQ and NorD from two bacterial species, together with biochemical tests and modern structure prediction, the authors found that NorQ and NorD meet at two main contact points. First, a finger‑like projection from the VWA domain of NorD inserts deeply into the central pore of the NorQ ring, nestling against a set of conserved loops. Second, the tail end of NorD reaches sideways to touch a loop on one NorQ subunit that sits just past a key sensor region for ATP.
Twisting, Stretching, and a Critical Flexible Link
By comparing several structural states, the study shows that when NorD binds, the NorQ ring shifts from a relatively flat shape into a spiral staircase. Different subunits hold ATP or its spent form, ADP, and as ATP binding and hydrolysis proceed, the active subunit at the bottom of the spiral steps upward, in line with a known hand‑over‑hand mechanism. Because NorD’s finger is clamped inside the pore while its tail is anchored to a specific loop, these changes rotate and tug the entire VWA domain. NorD also contains a long, previously mysterious linker connecting its VWA domain to an N‑terminal domain that likely contacts cNOR. The authors observe multiple conformations in which this linker must span different distances, implying that it can be progressively stretched. When they deleted parts of this linker, cNOR completely lost its non‑haem iron and activity, even though the chaperone complex could still form, underscoring that controlled stretching of this region is central to the remodeling job.
From Rotary Motion to Metal Insertion
Tying the structural snapshots together, the authors propose a model in which NorQ behaves like a molecular winch acting through NorD. In the first step, the NorQ ring associates with NorD via the finger‑in‑pore and tail‑to‑loop contacts, raising its ATPase activity and locking the VWA domain into position. Next, NorD engages cNOR at two “grip points”: one involving its N‑terminal domain and the other involving the VWA region or the linker near a metal‑binding motif. As NorQ cycles through ATP binding and hydrolysis, successive subunits climb the spiral, dragging the NorD finger around inside the pore. This motion twists the VWA domain relative to the N‑terminal grip and pulls the linker taut, transmitting torque and tension to cNOR. The resulting distortion is proposed to open an otherwise shielded region of cNOR, allowing an iron ion to be inserted into the FeB site, after which the complex can release and reset.
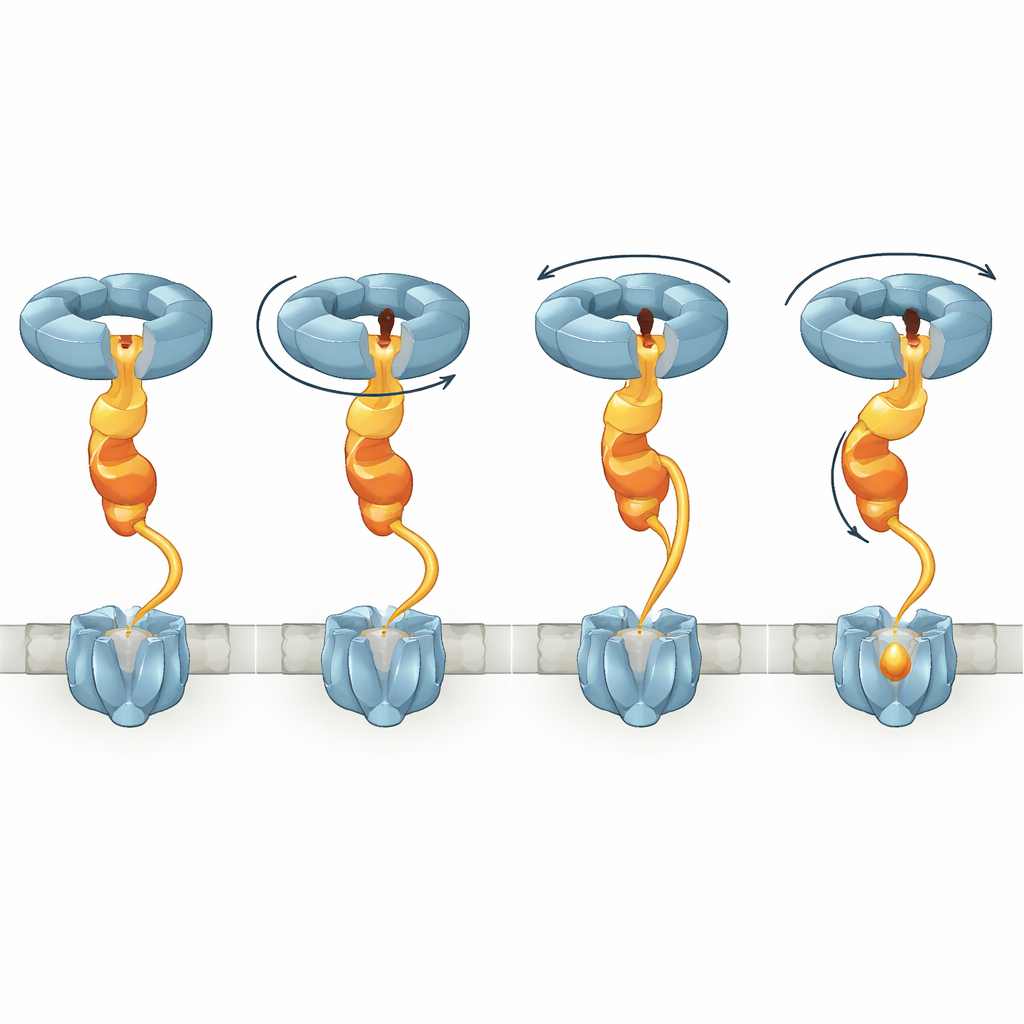
Why This Matters Beyond One Bacterial Enzyme
To a non‑specialist, the upshot is that the NorQ–NorD pair acts like a nanoscale pipe‑wrench, grabbing an enzyme at two points and twisting it just enough to slip an iron atom into place. This work not only clarifies how a key step in bacterial nitrogen chemistry occurs, but also reveals a general “finger‑in‑pore” strategy that similar motor–partner systems may use across biology. By showing how chemical fuel is converted into a precise twisting force on a target protein, the study offers a blueprint for understanding – and perhaps one day engineering – machines that control where and when metals are built into complex molecular devices.
Citation: Kahle, M., Appelgren, S., König, F. et al. NorQD AAA+ complex drives metal insertion by a twisting mechanism. Nat Commun 17, 3032 (2026). https://doi.org/10.1038/s41467-026-71044-4
Keywords: AAA+ ATPase, metallochaperone, nitric oxide reductase, cryo-EM structure, metal cofactor insertion