Clear Sky Science · en
Hierarchical small molecule inhibition of MYST acetyltransferases
Why tiny switches on DNA matter for cancer treatment
Inside every cell, chemical switches on our DNA help decide which genes turn on or off. A group of enzymes called MYST acetyltransferases fine-tune these switches and have been linked to cancers such as breast cancer and leukemia. This study explores how new drug molecules can turn these enzymes up or down in a controlled, stepwise way, offering clues for designing more precise cancer treatments and for interpreting ongoing clinical trials.
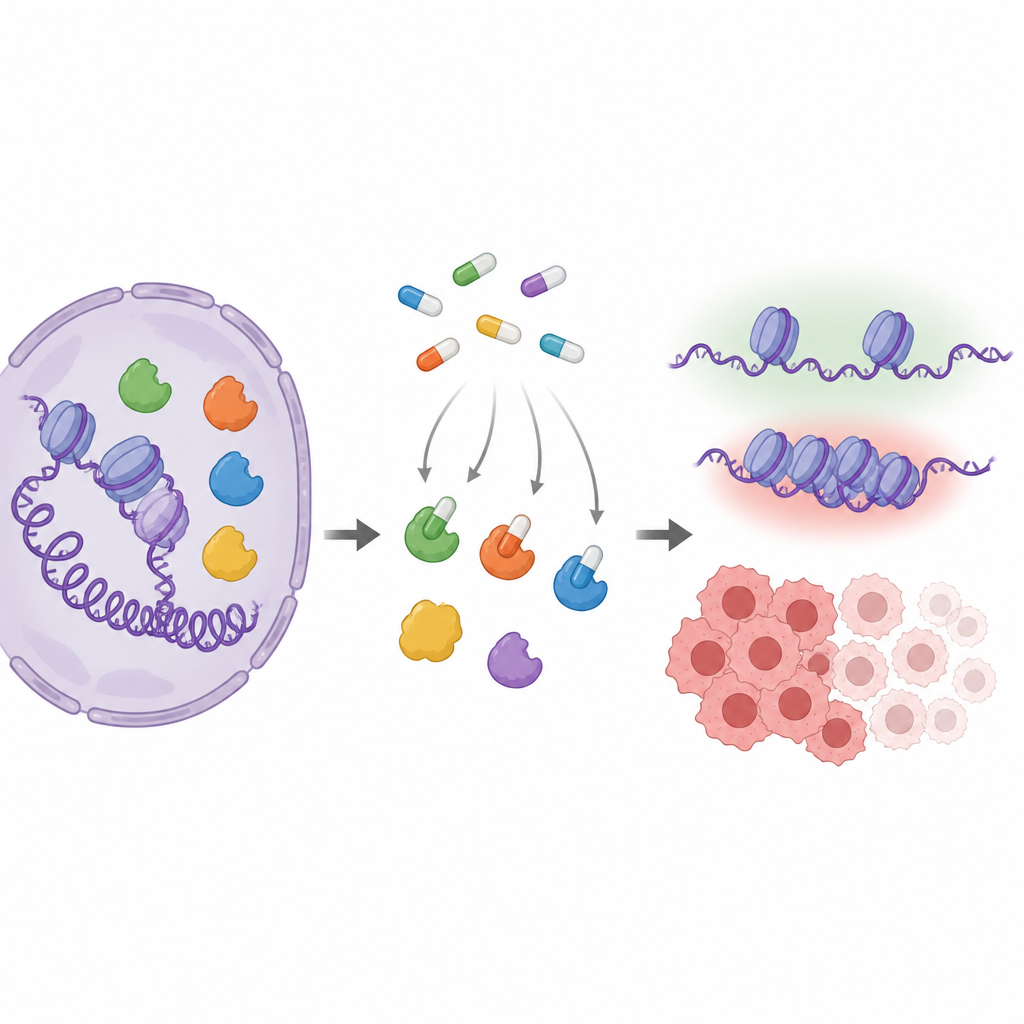
Key players that tune gene activity
MYST enzymes subtly alter how DNA is packaged by adding small chemical tags to proteins called histones. That packaging helps decide whether genes involved in growth, repair, or hormone response are active. Several members of this family, especially KAT6A and KAT6B, are altered or overproduced in cancers, which makes them attractive drug targets. Drug companies have developed pill-like compounds that fit into the working site of these enzymes, but because the different MYST enzymes look very similar at that site, it is hard to know exactly which ones each drug hits inside real cells.
Measuring which enzymes each drug really hits
The authors built a layered testing strategy to map how a leading drug, PF-9363, and related compounds interact with MYST enzymes. First, they used a "bait" molecule attached to magnetic beads that grabs enzymes using the same pocket where they use their natural fuel. If a drug blocks that pocket, the enzyme no longer sticks to the bait. By adding increasing drug doses to nuclear extracts from cells and then measuring, by mass spectrometry, which proteins still bound the bait, they found a clear hierarchy: PF-9363 first displaces KAT6A and KAT6B, then KAT7, then KAT8, and only at the highest levels KAT5. The same pattern appears for the clinical drug PF-8144, now in trials for estrogen receptor positive breast cancer.
Finding hidden partners and clear cellular readouts
Because blocking the enzyme pocket also weakens the grip of entire protein assemblies that sit on the bait, the team could see which other proteins travel together with each MYST enzyme. By clustering proteins that were lost from the bait with the same dose pattern as known complex members, and then combining this with 3D structure prediction and cell experiments, they flagged the transcription factor FOXK2 as a previously unrecognized partner, especially linked to the KAT8-containing NSL complex. Next, they moved into living cancer cells and examined hundreds of chemical marks on histones as they raised the PF-9363 dose. They pinpointed a small set of acetylation marks that reliably report which MYST enzyme has been blocked: one mark on histone H3 (H3K23Ac) for KAT6A/B, another on H3 (H3K14Ac) for KAT7, one on H4 (H4K16Ac) for KAT8, and marks on the variant histone H2A.Z for KAT5. These marks can be measured either by advanced mass spectrometry or by standard antibody-based blots, making them practical for many labs.
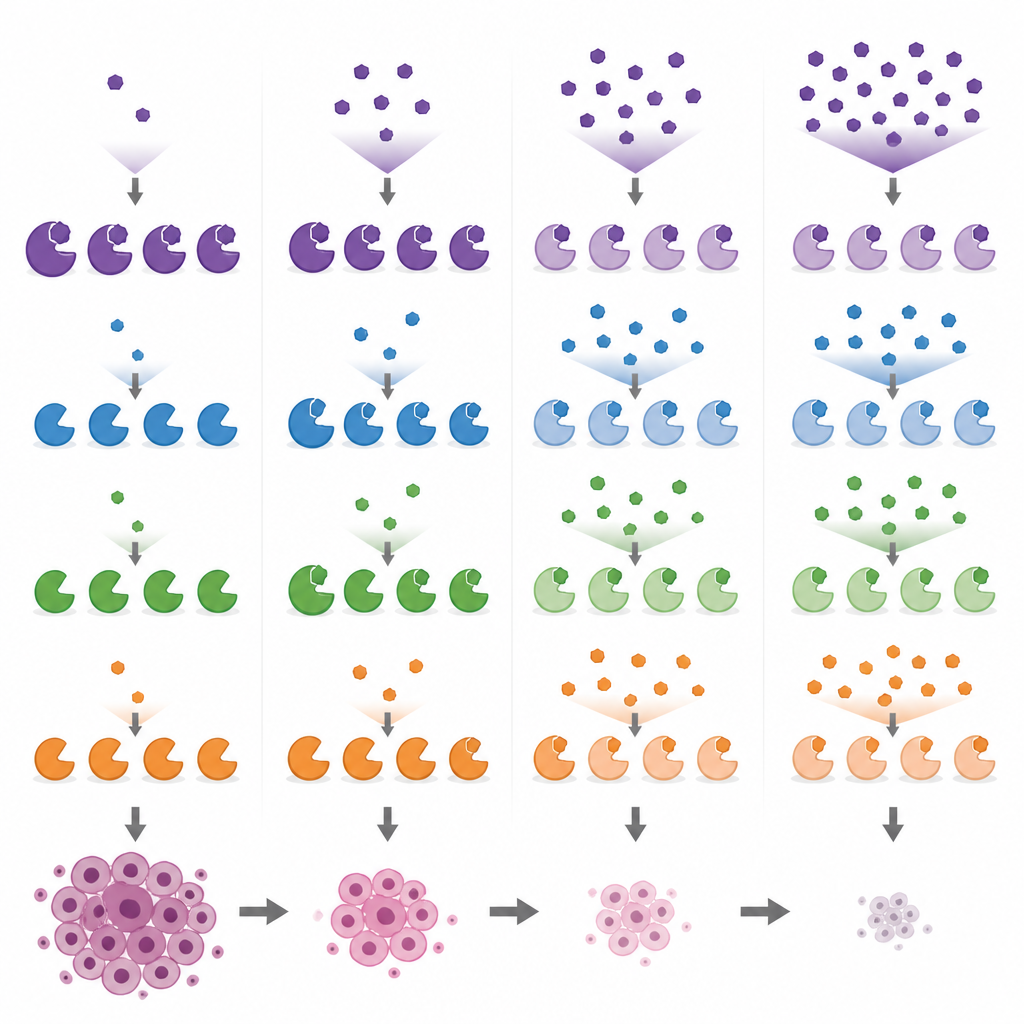
Comparing popular research drugs and their side targets
Armed with these biomarker readouts, the researchers compared several widely used MYST-targeting compounds in breast cancer cells across a range of doses. They showed that PF-9363 and its clinical cousin PF-8144 both follow the same stepwise pattern, eventually affecting KAT8 at higher doses. In contrast, WM-1119 turned out to be highly selective for KAT6A/B even when used at high concentrations, while WM-8014 and WM-3835 inhibit KAT6A/B first and only then KAT7. One compound, MOZ-IN-3, which looks potent in test-tube assays, showed little effect in cells, likely because it does not enter them well. These comparisons reveal that some tools marketed as specific for one enzyme actually hit several, which can complicate the interpretation of past and future experiments.
Linking dose, enzyme engagement, and cancer cell growth
The team then asked whether this layered inhibition pattern shapes how cancer cells respond. Screening PF-9363 across the NCI-60 panel of cancer cell lines showed that short exposures to doses that block only KAT6A/B and KAT7 have modest effects on growth. Strong, rapid growth slowdowns typically appeared only at higher doses where KAT8, an enzyme essential for most cells, was also engaged. In a triple-negative breast cancer line that especially depends on KAT8, the dose that shrank cell numbers matched the dose that fully suppressed the KAT8-linked histone mark while also blocking KAT6A/B and KAT7. This suggests that for some tumors, higher doses of these drugs may act through broader MYST inhibition rather than through KAT6A/B alone.
What this means for future cancer drugs
Overall, the study shows that small-molecule blockers of MYST enzymes behave much like many kinase drugs: at low doses they mainly hit their preferred targets, but as the dose rises they spread to closely related enzymes in a predictable order. By providing simple histone-based readouts for each MYST member and revealing where popular compounds are truly selective, the work offers researchers and clinicians a toolkit to match dose to desired enzyme coverage. This should help design clearer lab studies, interpret clinical trial results, and ultimately guide the development of more precise therapies that reshape gene control in cancer cells while limiting unwanted side effects.
Citation: Chen, X., Castroverde, A., Perez, M. et al. Hierarchical small molecule inhibition of MYST acetyltransferases. Nat Commun 17, 4329 (2026). https://doi.org/10.1038/s41467-026-70574-1
Keywords: epigenetic enzymes, histone acetylation, cancer drug targeting, chemoproteomics, lysine acetyltransferase inhibitors