Clear Sky Science · en
A model for drug transport across two membranes of Gram-negative bacteria by an MFS tripartite assembly
Why tiny pumps in bacteria matter
Antibiotic resistance often comes down to a microscopic arms race between drugs and the defenses of bacteria. One powerful defense is a set of tiny pumps that kick antibiotics out of the cell before they can do harm. This study uncovers the detailed structure and working model of one such pump, called EmrAB-TolC, in Escherichia coli. By showing how this machine spans both of the cell’s protective layers and expels drugs in a single step, the work offers new ideas for disarming bacterial defenses and making existing medicines more effective.
The challenge of getting drugs into tough bacteria
Many dangerous bacteria are wrapped in not one but two membranes, with a watery space in between. This double shell makes it hard for antibiotics to reach their targets. Gram-negative bacteria also build molecular machines that recognize harmful compounds inside the cell and eject them outside. EmrAB-TolC is one of these machines, and it can handle several different antibiotics. Until now, scientists only had partial snapshots of its individual parts. The big unknowns were how the pieces fit together and how the pump manages to move drugs all the way across both membranes in a coordinated way.
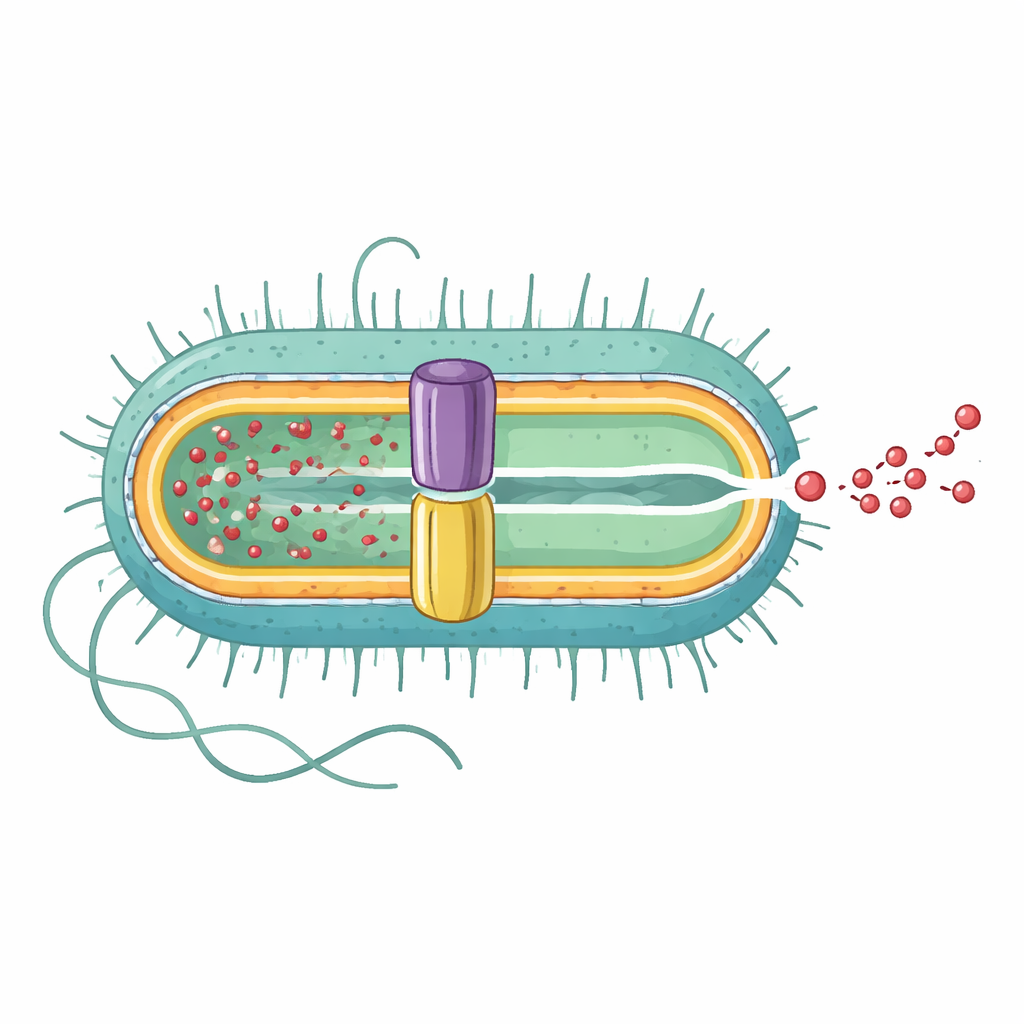
Revealing the full pump from inside to out
The research team used high-resolution cryo-electron microscopy to determine the three-dimensional structure of the complete EmrAB-TolC pump. They found that it is an elongated assembly made of three main parts in a 3:6:1 ratio: three copies of the outer membrane channel TolC, six copies of a middle adapter protein EmrA, and a single copy of the inner membrane transporter EmrB. Together they form a continuous tunnel roughly 320 angstroms long that reaches from the inner membrane, through the space between membranes, to the outside. EmrA forms a hexagonal sleeve between TolC and EmrB, shaping a central nanochannel, while TolC opens wide where it meets EmrA, creating a direct exit route to the cell exterior.
The inner engine that powers drug export
At the base of the assembly sits EmrB, a member of a large transporter family that uses the flow of protons, the charged particles that define acidity, to power movement of cargo. EmrB contains 14 membrane-spanning helices arranged into two domains that rock against each other. Structural analysis and computer simulations show that a key acidic residue, D29, and its neighbors form a proton-sensitive network that can switch EmrB between inward- and outward-open states. Mutations in D29 and nearby positions greatly reduce the cell’s resistance to several antibiotics, highlighting their importance in coupling proton movement to drug transport, even though proton flow can also involve additional residues.
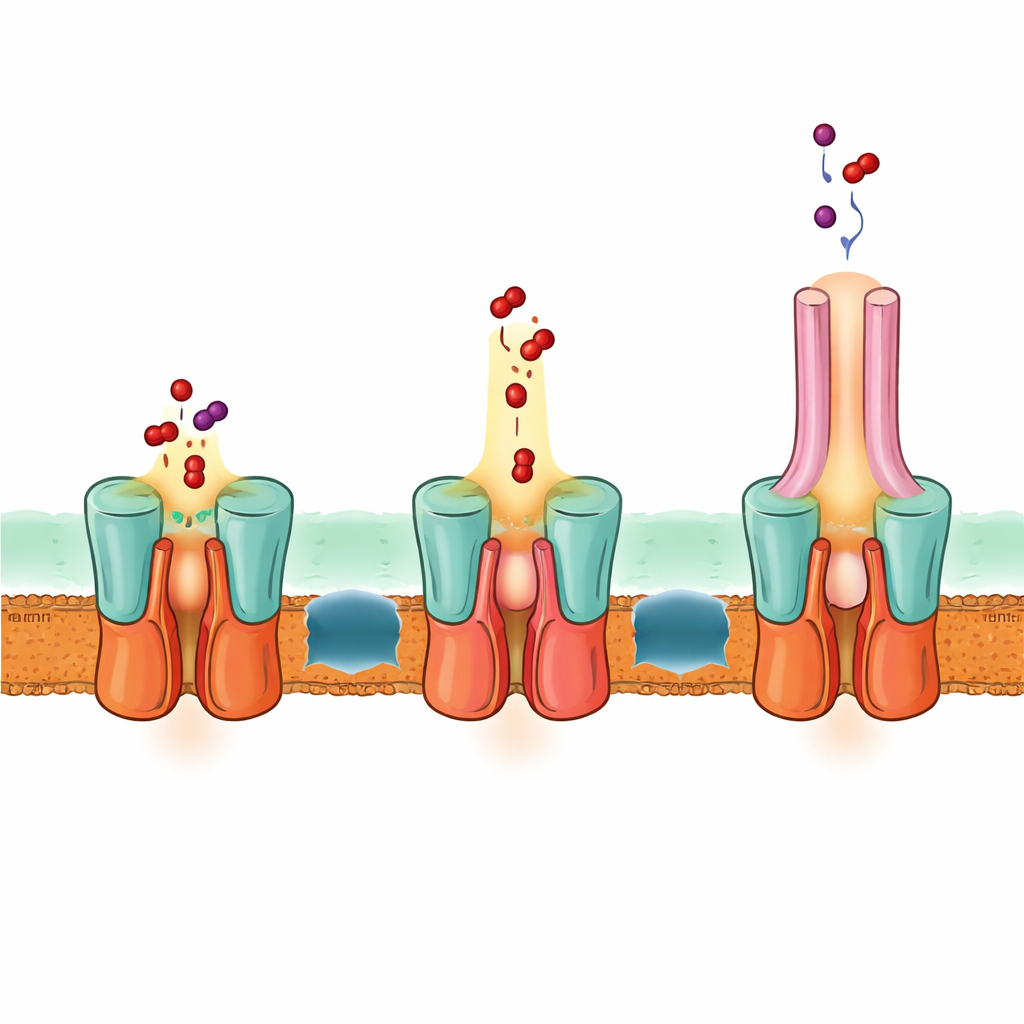
A one-step escape route across both membranes
The structure also explains how EmrA and EmrB interface to form a sealed conduit. A special extended helix bundle on the periplasmic side of EmrB plugs into a ring-shaped region of EmrA, locking the transporter to the adapter. This arrangement creates a spacious cavity that can receive drugs released from EmrB and guide them up into the EmrA channel and then into TolC. Based on the structure, computer predictions and mutational tests, the authors propose that EmrB grabs drugs from the inner membrane or cytoplasm, then rocks to open toward EmrA and release them into this shared cavity. A flexible loop within EmrA shapes a narrow path that funnels drugs upward through the adapter and out through TolC, all without the drugs ever freely diffusing in the space between the membranes.
What this means for fighting resistance
Altogether, the work supports a model in which EmrAB-TolC performs a true one-step export of antibiotics directly from inside the cell to the outside world. The pump uses proton flow to drive repeated shape changes in EmrB, which in turn push drugs into a continuous channel formed by EmrA and TolC. By pinpointing structural features that are essential for assembly and function, such as hydrophobic rings in EmrA and key residues in EmrB, the study highlights potential weak spots where small molecules could jam the pump. Targeting these spots could help restore the potency of existing antibiotics against Gram-negative pathogens that currently withstand treatment.
Citation: Zhong, Z., Maimaiti, T., Jackson, M.L. et al. A model for drug transport across two membranes of Gram-negative bacteria by an MFS tripartite assembly. Nat Commun 17, 4039 (2026). https://doi.org/10.1038/s41467-026-70500-5
Keywords: antibiotic resistance, efflux pump, Gram-negative bacteria, drug transport, EmrAB-TolC