Clear Sky Science · en
Single-cell thiol profiling enabled by live-cell labeling reveals metabolic heterogeneity in ferroptosis
Why tiny cell chemicals matter
Every cell in our body depends on a family of small sulfur‑containing molecules, called thiols, to keep its chemistry running smoothly. These molecules help control energy use, protect against damage from reactive oxygen, and decide whether a cell survives or dies. Yet because thiols are fragile and exist in tiny amounts, researchers have struggled to measure many different thiols in single cells at once. This study introduces a way to freeze and read out thiols inside individual living cells, and uses it to uncover why some cancer cells succumb to a type of cell death called ferroptosis while others resist.
Locking fragile cell helpers in place
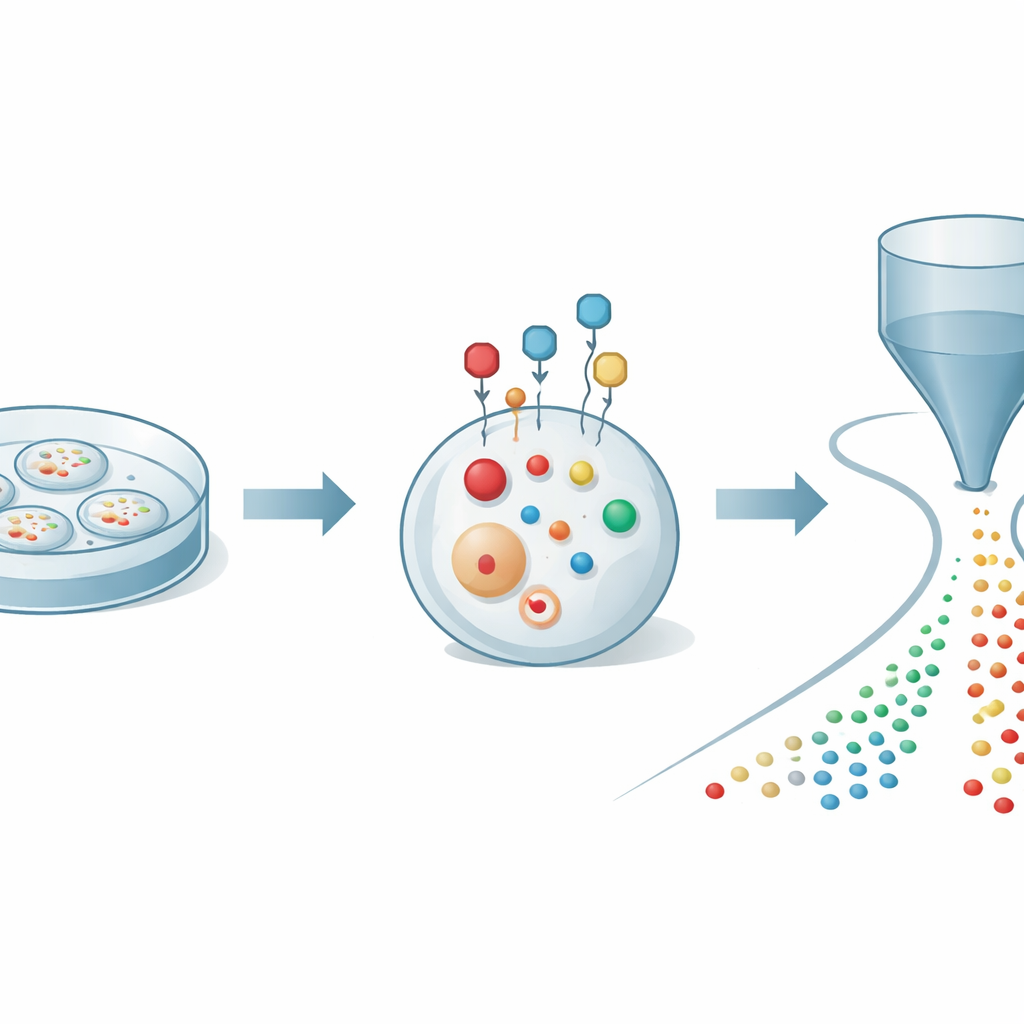
The authors began by tackling a basic problem: thiols quickly oxidize and change when cells are broken open for analysis, which can give a distorted view of what is really happening inside. They designed a small chemical probe that can slip through the cell membrane and react rapidly and selectively with thiols in living cells. Once attached, the probe acts like a protective cage and an amplifier: it stabilizes the thiols against oxidation and adds an electrical charge that makes them far easier to detect by mass spectrometry. By carefully tuning the probe’s structure, the team found conditions that label thiols within minutes while keeping cells healthy, allowing them to capture a reliable snapshot of thiol chemistry in real time.
Reading hundreds of molecules from single cells
To make use of this labeling, the researchers coupled it to an advanced detection platform known as organic mass cytometry. In this setup, thousands of labeled cells are gently pushed one by one through a thin capillary into an ionization source attached to a high‑resolution mass spectrometer. As each cell passes through, its small molecules are extracted and converted into ions, creating a brief pulse of signals unique to that cell. Computational tools then identify which pulse belongs to which cell and assign hundreds of molecular peaks to each event. With this integrated strategy, the authors could quantify 27 distinct labeled thiols along with more than 300 other metabolites in individual cancer cells, revealing a rich map of sulfur and energy metabolism at single‑cell resolution.
Watching stress ripple through sulfur networks
The team next asked whether this approach could track how cells respond when thiol pathways are deliberately disrupted. They used drugs that selectively block different steps of glutathione (GSH) synthesis, a major thiol that protects cells from oxidative damage. In single cells, they observed precise and pathway‑consistent shifts: blocking cystine import depleted cysteine and many downstream sulfur metabolites; blocking individual glutathione‑building enzymes caused upstream components to pile up while GSH levels fell. Beyond thiols, hundreds of other metabolites also changed in characteristic patterns, reflecting broader shifts in energy and redox balance. These experiments showed that the method can follow subtle, localized changes as they spread through the cell’s interconnected sulfur network.
Uncovering hidden differences in cell death sensitivity
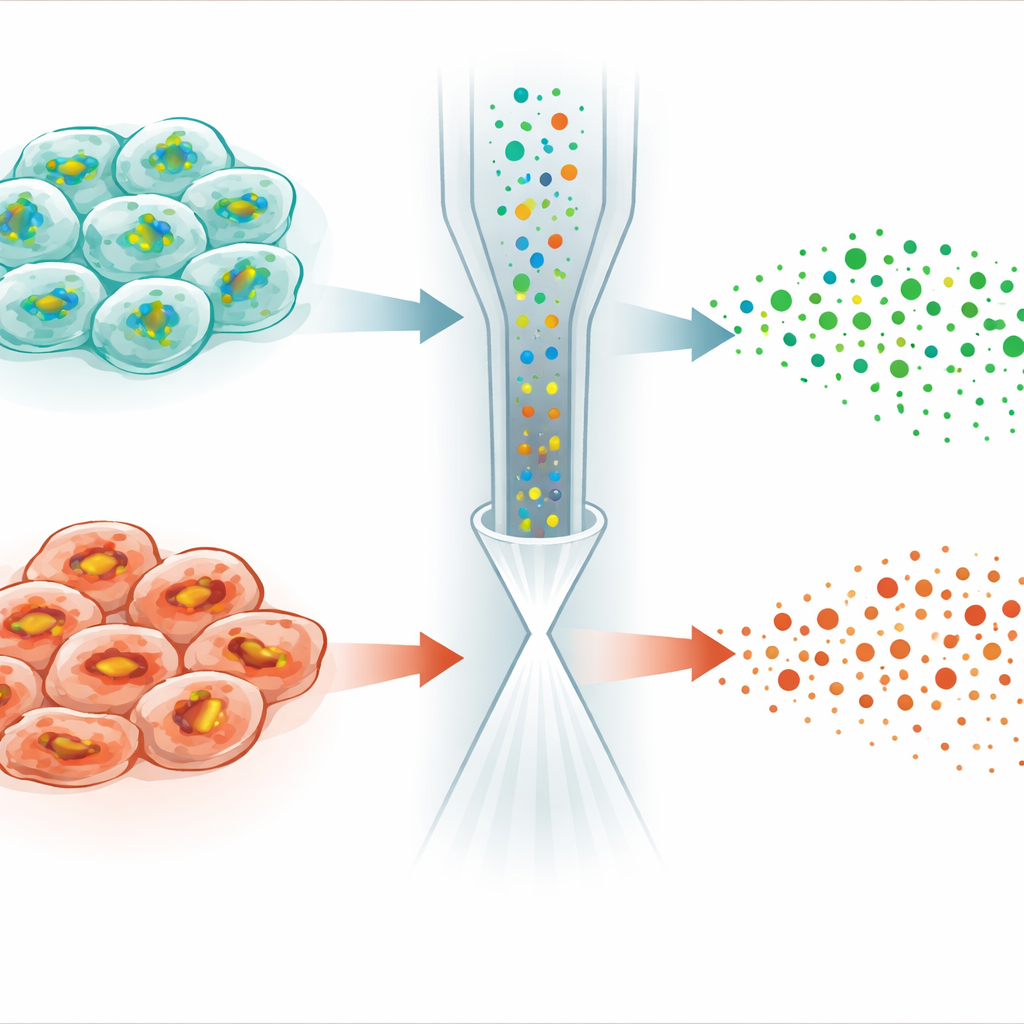
The most striking application came from studying ferroptosis, a form of regulated cell death driven by iron‑dependent damage to cell membranes. A small molecule called RSL3 kills cancer cells by disabling an enzyme that normally uses GSH to detoxify lipid peroxides. Surprisingly, when the authors treated HeLa cancer cells with RSL3, GSH levels did not simply stabilize or rise—as one might expect when its consumption is blocked—but instead dropped in many cells. Single‑cell analysis revealed that the population split into two subtypes: one retained near‑normal GSH and lipid profiles, while the other showed sharply reduced GSH, weakened GSH production, and marked changes in lipid molecules prone to peroxidation. Additional experiments linked these differences to varying levels of a stress‑response regulator called NRF2, which controls the expression of genes that import cystine and synthesize and recycle GSH.
What this means for future therapies
Taken together, the study shows that combining live‑cell thiol labeling with organic mass cytometry can stabilize and measure many fragile thiols alongside broader metabolism in individual cells. This powerful readout uncovered that RSL3 does more than block GSH use: it also suppresses GSH production through the NRF2–glutathione axis, creating distinct subpopulations of ferroptosis‑resistant and ferroptosis‑sensitive cells within the same tumor line. For a lay reader, the key message is that cells that look identical under the microscope can differ profoundly in how they manage protective chemicals like glutathione, and those differences can decide whether a cancer therapy works. By making it possible to see these hidden metabolic variations one cell at a time, the new method may help design more precise strategies to push vulnerable cells toward death while sparing healthy or resistant ones.
Citation: Miao, D., Li, Q., Zhang, Y. et al. Single-cell thiol profiling enabled by live-cell labeling reveals metabolic heterogeneity in ferroptosis. Nat Commun 17, 3378 (2026). https://doi.org/10.1038/s41467-026-70336-z
Keywords: single-cell metabolomics, glutathione, ferroptosis, redox metabolism, cancer cell heterogeneity