Clear Sky Science · en
Nucleotide-resolution mapping of regulatory elements via allelic readout of tiled base editing
Why tiny DNA changes can matter for cancer treatment
Cancer doctors increasingly rely on smart immune therapies that hunt down tumor cells by recognizing specific tags on their surface. One of the best-known tags on leukemia cells is called CD19, the target of powerful CAR-T cell therapies. Yet some patients relapse because their cancer cells quietly lose or dim this tag, slipping past treatment. This study shows how scientists can now zoom in on stretches of DNA that control CD19, pinpointing which individual "letters" in those control regions matter for keeping CD19 visible—and how changes at just a few of these letters can help cancer cells escape therapy. 
Reading the control switches of our genes
Genes are not simply on or off; they are controlled by nearby DNA switches called enhancers that fine-tune how much of a given protein a cell makes. Many disease-linked DNA changes lie in these non-coding control regions rather than in protein-coding genes themselves. Traditional genetic studies can flag suspicious regions but struggle to say exactly which DNA letters within them are critical, or how they affect cell behavior. For therapies like CD19 CAR-T cells, this gap is important: a subtle change in a control switch might lower CD19 just enough for cancer cells to hide, while leaving the rest of the cell’s machinery intact.
Using molecular pencils to rewrite DNA one letter at a time
The researchers built an end-to-end experimental and computational framework to map these crucial DNA letters with unprecedented precision. They focused on a 346-letter-long enhancer just upstream of the CD19 gene in leukemia cells. Using advanced CRISPR base editors—molecular "pencils" that can change single DNA letters without cutting the DNA—they swept across this enhancer with a dense library of guide molecules. Each guide targeted a slightly different spot, collectively sprinkling millions of cells with distinct single-letter changes and small combinations of changes within the same short DNA segment.
Linking DNA variants to protein levels in bulk
To see how each edited DNA version affected CD19, the team stained the cells with a fluorescent antibody that glows more strongly when CD19 levels are high. They then used a cell-sorting machine to split the population into CD19-high and CD19-low groups. Crucially, instead of just counting which guides were present, they directly sequenced the edited enhancer region itself in each group. New analysis tools—CRISPR-Millipede and an adapted version of the DESeq2 software—allowed them to treat each distinct edited DNA sequence as its own variant and statistically infer which exact single-letter changes shifted cells toward lower or higher CD19. 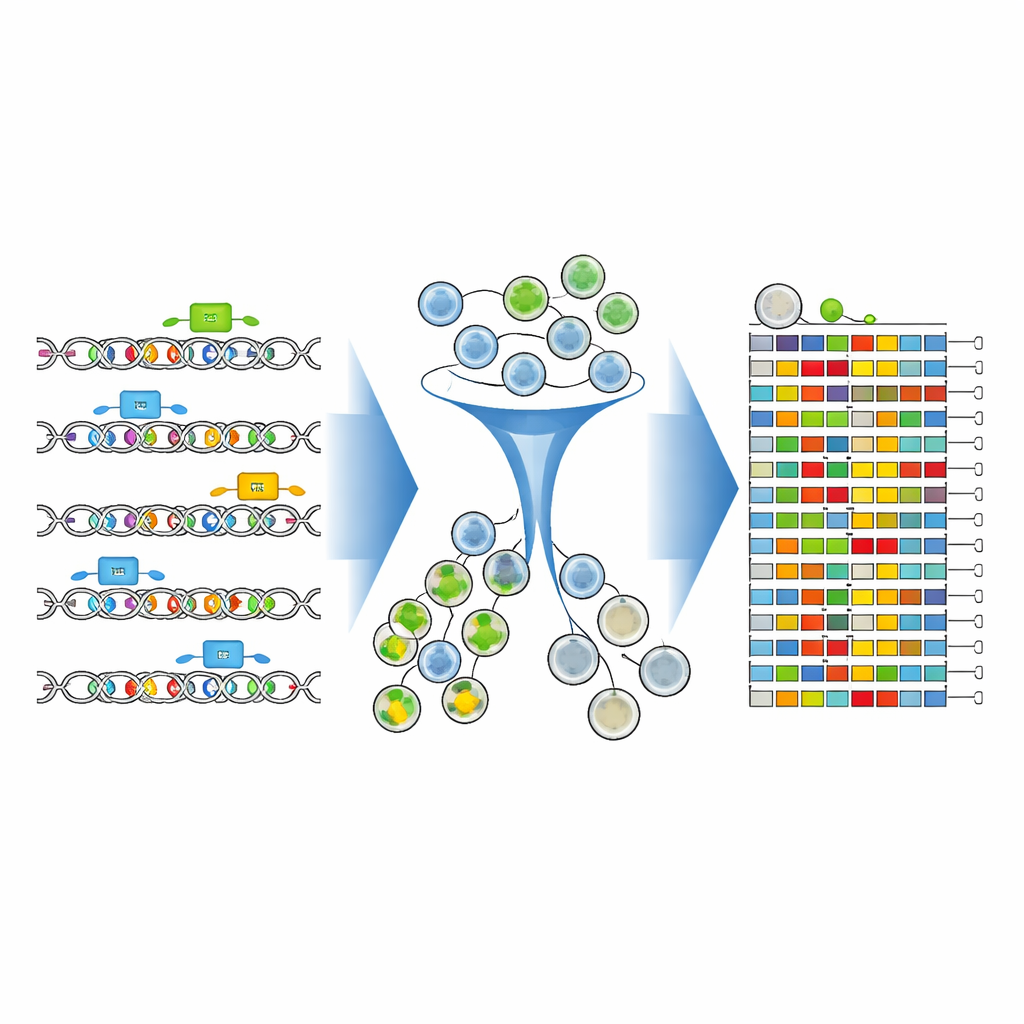
Finding the key protein docking sites in the enhancer
With this nucleotide-by-nucleotide map in hand, the researchers overlaid their results with known docking motifs for transcription factors—proteins that bind DNA to turn genes up or down. They found that the most impactful letters clustered within sites recognized by factors such as MYB, PAX5, and EBF1, which are important regulators in B cells. By deliberately recreating these specific enhancer mutations and by disabling the corresponding transcription factors, they confirmed that damaging these docking sites lowers CD19 levels. They also showed that some apparent hits from older, guide-count-based methods were false positives caused by the mere presence of the bulky editing machinery on DNA rather than genuine sequence changes, and they engineered a degradable editor to separate these effects.
How small enhancer changes can undermine CAR-T therapy
Finally, the team asked whether these subtle regulatory tweaks actually influence cancer treatment. They mixed leukemia cells with and without the enhancer mutations and exposed them to CD19 CAR-T cells. Edited cells carrying particular enhancer changes consistently outgrew their unedited counterparts under CAR-T attack, revealing that modest reductions in CD19—driven not by breaking the CD19 gene but by altering its enhancer—were enough to help cancer cells survive. The work demonstrates a practical, scalable way to chart cause-and-effect at the single-letter level in gene control regions, and it shows that non-coding variants can silently steer the success or failure of cutting-edge immunotherapies.
Citation: Becerra, B., Wittibschlager, S., Patel, Z.M. et al. Nucleotide-resolution mapping of regulatory elements via allelic readout of tiled base editing. Nat Commun 17, 3398 (2026). https://doi.org/10.1038/s41467-026-69918-8
Keywords: CRISPR base editing, gene regulation, CD19 CAR-T resistance, non-coding variants, enhancer mapping