Clear Sky Science · en
Lid loop-mediated proton transfer revealed in the Fe/αKG-dependent decarboxylase TraH
Why a tiny moving loop matters
Inside microbes, specialized proteins quietly build and reshape complex molecules, including potential antibiotics. This study zooms in on one such protein, called TraH, from a common food‑spoiling fungus. The researchers reveal that a small flexible "lid" in TraH acts like a movable gate that controls how the enzyme removes a piece of a molecule and, in some cases, helps determine what kind of chemical change happens at all. Understanding this moving part not only deepens our picture of how nature’s catalysts work, but could also help scientists design new enzymes for making medicines and other valuable compounds.
A fungal helper in molecule making
TraH belongs to a large family of enzymes that use iron and a helper molecule called α‑ketoglutarate to carry out powerful oxidation reactions. These enzymes are central players in microbial chemistry, where they help build “secondary metabolites” such as pigments, toxins, and drug‑like molecules. In the fungus Penicillium crustosum, TraH helps produce terrestric acid, a compound with promising antibacterial and antiviral properties. TraH’s main job is to clip off a carbon dioxide group from a precursor called crustosic acid, while at the same time introducing a double bond into the molecule. Intriguingly, TraH can also act on a closely related methyl‑ester version of the same compound, but in that case it simply introduces a double bond without removing carbon dioxide.
A closing lid that reshapes the active site
Using X‑ray crystallography, the authors captured high‑resolution snapshots of TraH in different states: empty, bound to metal only, and bound to metal plus either crustosic acid or its methyl ester. All forms share a rigid core shaped like a double‑layered beta‑helix, which holds the iron atom where chemistry happens. Wrapped around this core are extra segments at the protein’s ends, including a loop near the front that behaves like a lid. In the unbound state, this lid is floppy and partly invisible in the crystal structure, indicating that it wanders. When the true carboxylate substrate binds, the lid swings down, becomes well ordered, and, together with nearby helices, seals off the active site so the molecule sits deep inside a closed pocket. For the methyl‑ester substrate, however, much of the lid remains disordered and pulled back, showing that its precise closure depends on the nature of the bound molecule.
A water‑based relay for moving protons
Closer inspection of the closed‑lid structure with crustosic acid revealed an intricate network of hydrogen bonds that links the lid to a conserved lysine (K191) in the enzyme’s core and to the substrate’s carboxyl group. Key lid residues help position one or two water molecules between K191 and the reacting end of the substrate. To test how important this network is, the team changed individual amino acids in the lid and nearby core. Mutations that disrupted the network, especially at positions Q99, D112, K114, K191, and Y214, sharply reduced or abolished the decarboxylation reaction, while changes in neighboring, non‑network residues had little effect. In contrast, most of these disruptive mutations hardly affected the enzyme’s ability to carry out desaturation on the methyl‑ester substrate, suggesting that the water‑mediated network is crucial specifically for carbon‑dioxide loss.
Following the reaction step by step
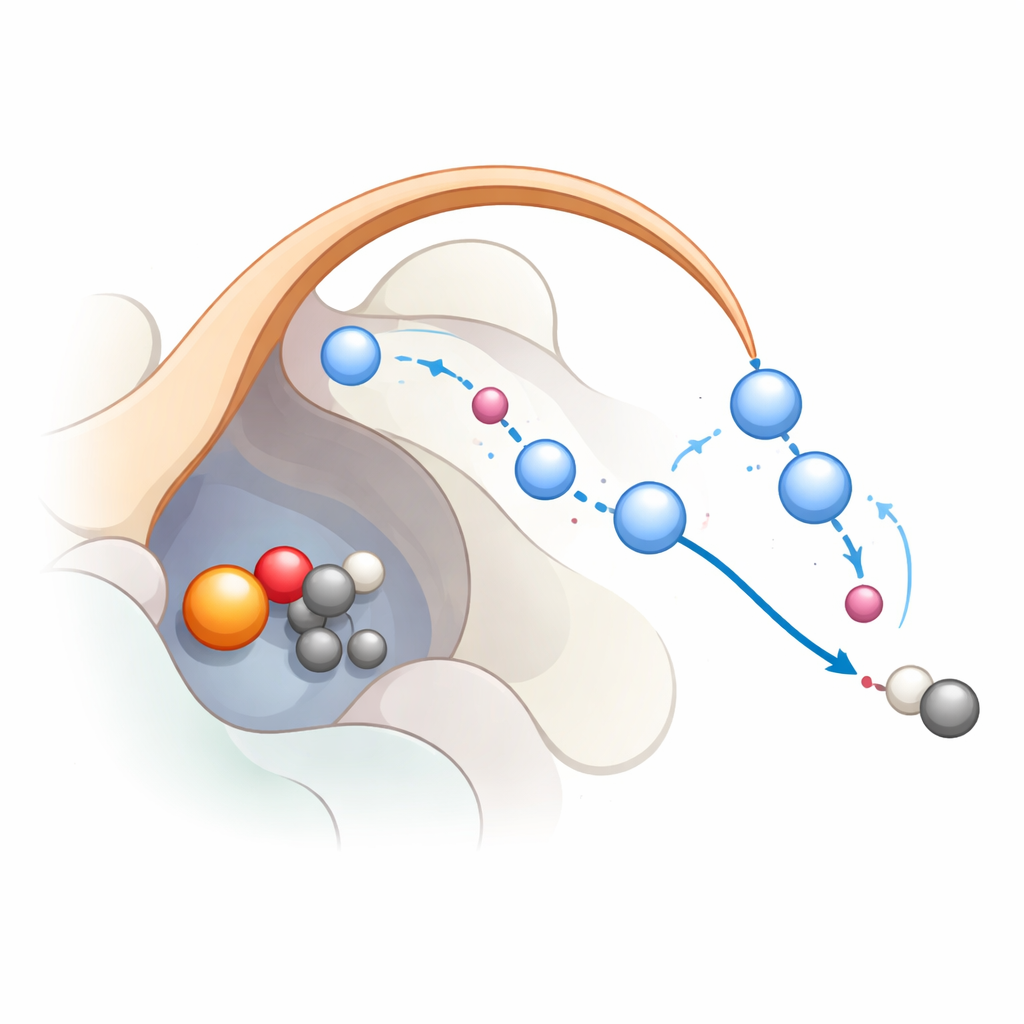
To see how this network actually works during the reaction, the researchers combined classical molecular dynamics with more advanced quantum‑mechanical simulations. They started from a state where the iron has already formed a highly reactive iron‑oxo species, a common intermediate in this enzyme family. The calculations support a three‑step pathway: first, the iron‑oxo pulls a hydrogen atom from the substrate, creating a radical; second, an electron shifts back to the iron, helping the substrate shed carbon dioxide; third, the iron‑bound hydroxyl group is reprotonated so the iron returns to its resting state. The simulations show that K191 and the chain of water molecules stabilized by the lid loop form a proton relay that shuttles protons efficiently during the last steps. When this relay is broken, the reaction becomes energetically unfavorable, neatly aligning with the mutational data.
An adaptable loop that guides enzyme evolution
Beyond TraH itself, the authors compared related enzymes that share the same iron‑binding core but handle different chemical tasks. Many of these enzymes also carry lid‑like loops, but their lengths, sequences, and degrees of motion vary widely. In some, the loop closes firmly over substrates and forms key hydrogen bonds; in others it remains open and does not appear to participate in chemistry. By building evolutionary trees and analyzing sequence patterns, the team found that certain lid residues involved in TraH’s hydrogen‑bond network are strongly conserved, whereas other positions vary in a way that tracks with different substrate preferences. This suggests that evolution tunes both the rigidity and the chemistry of these loops to recognize new molecules and to bias reactions toward particular outcomes.
What this means for future chemistry
To a non‑specialist, the central message is that a small flexible loop on the surface of an enzyme can control where protons and water molecules travel inside, and that this control can determine whether a molecule simply loses a hydrogen or also loses carbon dioxide. In TraH, the lid’s job is not just to close the active site but to assemble a tiny proton‑transfer machine out of amino acids and trapped water. This loop‑mediated strategy differs from that of many related enzymes, which use a fixed basic residue placed right next to the departing group. By revealing this alternative design, the work broadens our understanding of how nature builds versatile catalysts and offers a blueprint for engineering enzymes whose moving parts can be rewired to make new reactions and new bioactive molecules possible.
Citation: Zheng, X., Ge, R., Guo, Z. et al. Lid loop-mediated proton transfer revealed in the Fe/αKG-dependent decarboxylase TraH. Commun Chem 9, 159 (2026). https://doi.org/10.1038/s42004-026-01986-9
Keywords: enzyme mechanisms, nonheme iron enzymes, proton transfer, fungal secondary metabolism, enzyme evolution