Clear Sky Science · en
Enabling multi-target drug discovery through latent evolutionary optimization and synthesis-aware prioritization (EVOSYNTH)
Why designing smarter medicines matters
Many serious illnesses, from Alzheimer’s disease to ovarian cancer, do not arise from a single faulty protein but from entire webs of disrupted pathways. Treating such conditions with one drug aimed at one target often falls short, while giving patients multiple drugs can create toxic side effects and complex dosing schedules. This study introduces EVOSYNTH, a computational framework that aims to design single molecules capable of safely and effectively hitting several disease-related targets at once—and, crucially, that chemists can actually make in the lab.
From single targets to many switches
Traditional drug discovery has largely chased “magic bullet” compounds that bind tightly to one protein and leave everything else alone. Over time, it has become clear that this approach struggles with complex diseases involving multiple feedback loops and backup routes. Adding more drugs—the polypharmacy strategy—can improve coverage but raises the risk of dangerous drug–drug interactions, unpredictable behavior in the body, and burdensome treatment plans. Multi-target drugs offer a middle path: one thoughtfully designed molecule that modulates several key proteins at once, ideally nudging an entire network back toward health. The challenge is that finding such molecules, and ensuring they remain practical to synthesize, is extremely difficult by hand.
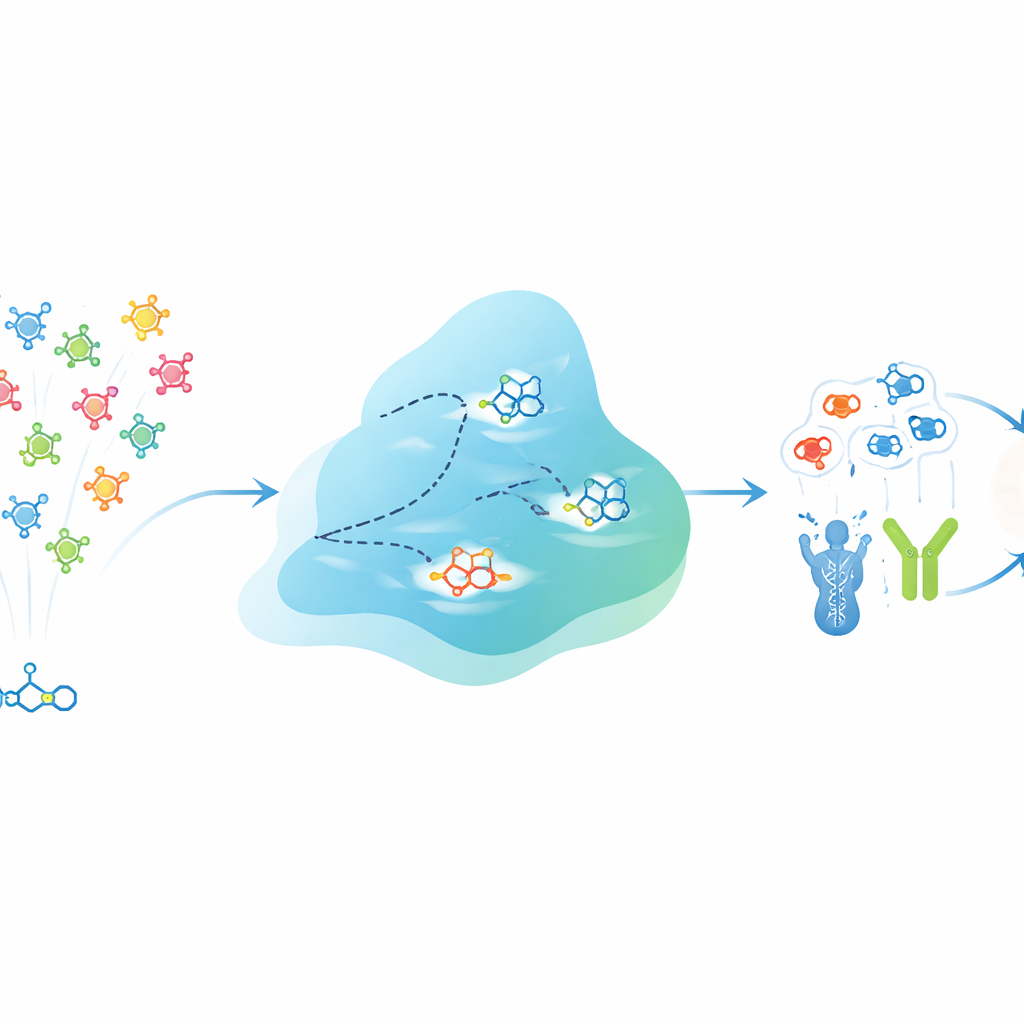
A digital evolution engine for molecules
EVOSYNTH tackles this challenge by treating potential drugs as a population that can “evolve” inside a computer. Instead of altering real molecules in a test tube, the system works in an abstract, compressed space that encodes both the structure of each molecule and its predicted behavior against different protein targets. A specialized neural network learns this space so that nearby points tend to correspond to molecules with similar multi-target activity. A diffusion process then plays the role of mutation, gently perturbing these points to generate new virtual molecules. After each round, candidates are evaluated for how strongly they are predicted to bind several chosen proteins at once, while hard filters discard any that fail basic requirements for drug-likeness, toxicity, or, for brain diseases, the ability to cross the blood–brain barrier.
Designing drugs that can actually be made
An important innovation of EVOSYNTH is that it does not stop at virtual activity. Many proposed molecules from artificial intelligence models look promising on screen but are prohibitively difficult or expensive to synthesize. To avoid this pitfall, the authors integrate a synthesis planning system called SPARROW into their workflow. This tool builds reaction networks that connect each candidate molecule back to readily purchasable building blocks, estimating the number of steps, cost of starting materials, and reliability of each reaction. Instead of simply rejecting impossible molecules after the fact, EVOSYNTH uses these network-wide calculations to prioritize candidates that offer the best trade-off between therapeutic promise and real-world manufacturability, including opportunities to reuse intermediates and share steps across several top compounds.
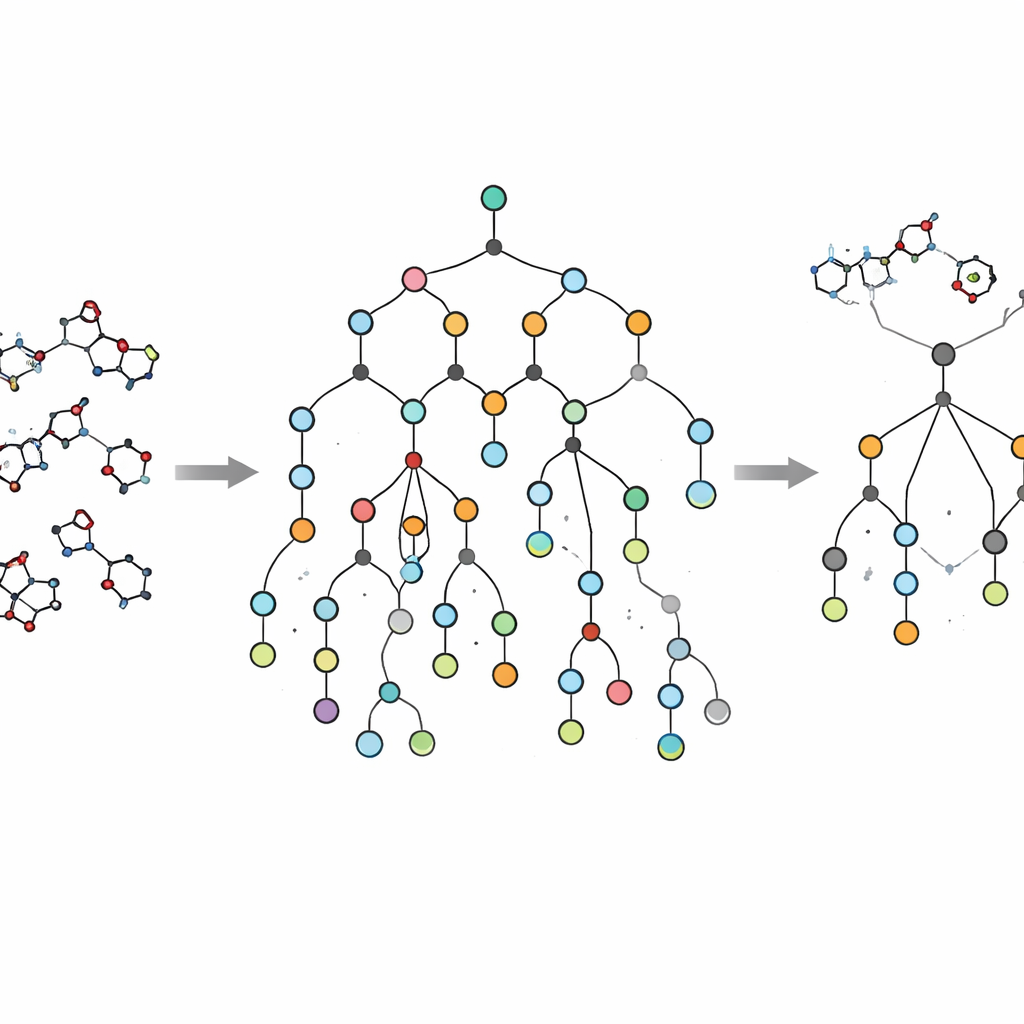
Realistic tests in brain disease and cancer
To demonstrate what EVOSYNTH can do, the researchers applied it to two demanding test cases. For Alzheimer’s disease, they sought molecules that could block both JNK3 and GSK3β, two enzymes implicated in nerve cell death and abnormal protein buildup in the brain. For ovarian cancer, they targeted a pair of proteins involved in growth signaling and DNA repair, PI3K and PARP1, which have been shown to work especially well when inhibited together. Starting from known single-target drugs as “seed” scaffolds, EVOSYNTH generated and refined new structures over many generations. Compared with a closely related method and another advanced generative model, EVOSYNTH consistently produced candidate molecules with stronger predicted activity at both targets, greater structural variety, and lower similarity to the original scaffold—signs that it is exploring genuinely new chemical territory rather than endlessly tweaking the same motif.
Balancing power, diversity, and cost
When the team examined the best molecules through the lens of synthesis, the advantages of their approach became even clearer. Under realistic limits—only about ten compounds can go forward to experimental testing—EVOSYNTH’s top selections combined higher expected therapeutic payoff with fewer reaction steps, fewer distinct starting materials, and substantially lower estimated material costs than those proposed by competing methods. Interestingly, this cost-effectiveness emerged even though the system was not explicitly trained to minimize synthetic difficulty. Instead, its tendency to generate families of related yet diverse molecules allowed the synthesis planner to reuse building blocks and reaction sequences, naturally favoring designs that would scale better in the lab.
What this means for future medicines
For a non-specialist, the key takeaway is that EVOSYNTH offers a way to search intelligently through the immense space of possible molecules, looking not just for those that hit several disease switches at once but also for those that chemists can make without heroic effort. By tightly coupling predictions of multi-target activity with checks on safety, brain penetration, and synthetic practicality, the framework moves computational drug design a step closer to producing real, testable compounds. While the current work focuses on pairs of targets and relies on computer-estimated binding rather than full experimental data, the approach is general. With further refinement, similar systems could help design a new generation of multi-purpose drugs that are both biologically sophisticated and manufacturable at scale.
Citation: Nguyen, V.T.D., Pham, P. & Hy, TS. Enabling multi-target drug discovery through latent evolutionary optimization and synthesis-aware prioritization (EVOSYNTH). Commun Chem 9, 133 (2026). https://doi.org/10.1038/s42004-026-01945-4
Keywords: multi-target drug design, AI-driven drug discovery, evolutionary optimization, synthetic accessibility, polypharmacology