Clear Sky Science · en
Versatile and portable Cas12a-mediated detection of antibiotic resistance markers
Why tracking invisible germs matters
Antibiotic resistance sounds abstract until a routine infection no longer responds to medicine. Bacteria that can shrug off our drugs are emerging not only in hospitals, but also on farms and in rivers and soils. To keep ahead of this threat, scientists need fast, affordable ways to spot the genetic warning signs of resistance wherever they appear. This paper describes a new laboratory toolbox, called C12a, that uses CRISPR gene‑cutting technology to detect key resistance genes in bacteria from people, animals, and the environment.
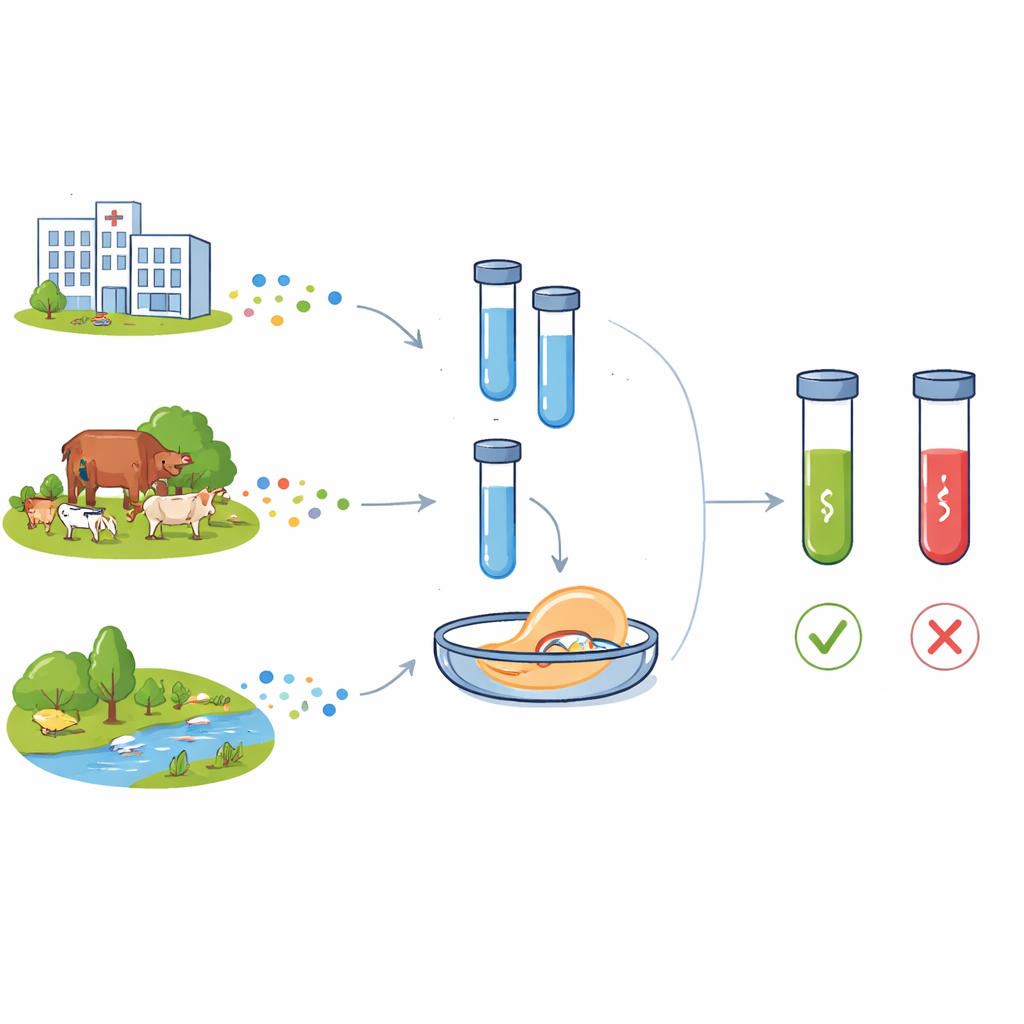
A growing problem that crosses hospital walls
Modern antibiotics transformed medicine, turning once deadly infections into treatable illnesses and enabling complex surgery and intensive care. But heavy use in human medicine, livestock production, and poorly treated waste has sped up the evolution and spread of antimicrobial resistance. Two resistance genes are especially worrying: blaCTX-M-15, which protects bacteria from a broad class of beta‑lactam antibiotics used in people, and floR, which guards against amphenicol drugs widely used in animals. Both genes now appear in bacteria from hospitals, communities, farms, and water, often moving between species on mobile genetic elements. Another genetic element, the class 1 integron (tracked by the intI1 gene), frequently carries clusters of resistance genes and is considered a marker of strong human impact on microbial ecosystems.
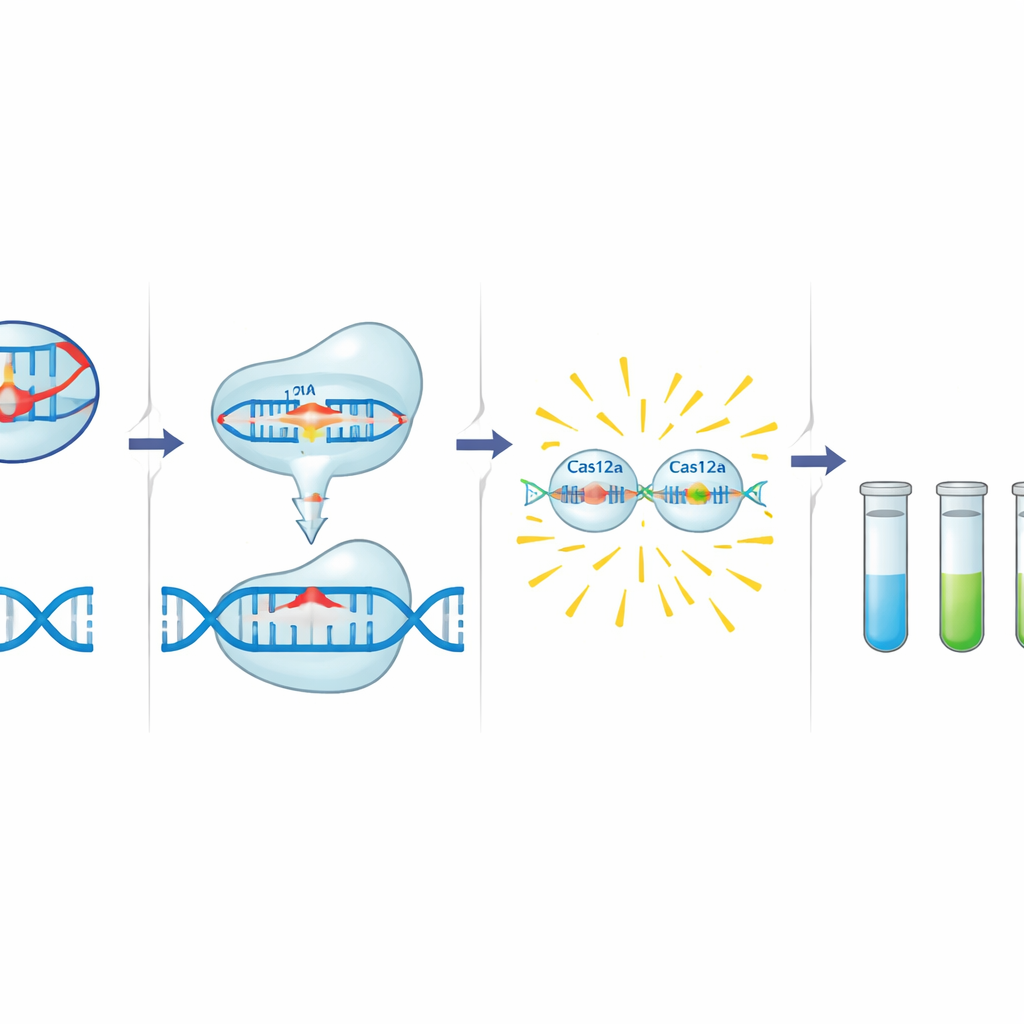
Turning CRISPR into a resistance detector
The C12a toolbox builds on Cas12a, a CRISPR‑associated enzyme that can be programmed with a short guide RNA to recognize a specific DNA sequence. When Cas12a finds its target next to a short trigger motif, it first cuts that DNA and then begins rapidly chopping nearby single‑stranded DNA. The authors harness this behavior by adding a tiny DNA probe tagged with a fluorescent dye and a quencher. In the presence of a resistance gene, activated Cas12a slices the probe, releasing a bright signal. Before this detection step, a standard PCR reaction amplifies a small region of the target gene that includes both the Cas12a trigger motif and the guide’s binding site, adding a second layer of specificity. The team designed and optimized guides and primers for three targets: blaCTX-M-15 (C12abCTX), floR (C12aFLO), and the class 1 integron integrase gene intI1 (C12aINT).
How sensitive and reliable is the toolbox?
To judge performance, the researchers measured how little DNA the system could reliably detect. Using purified fragments of the target genes, C12abCTX and C12aFLO detected concentrations as low as 70 and 50 attomolar, respectively—far below what can be seen by simply running PCR products on a gel. When they tested real bacterial cultures, the assays could detect around 77 cells per milliliter for blaCTX-M-15 and 173 cells per milliliter for floR. In collections of Escherichia coli from children’s stool samples, results from the C12a toolbox matched standard antibiotic susceptibility tests and, where available, whole‑genome sequencing: samples genetically positive for the resistance genes gave strong fluorescence signals, while negative samples stayed at background levels.
Linking resistance genes to broader risk
The authors also explored whether their tools could flag bacteria that carry many resistance traits at once. Using C12aINT, they searched the same isolates for the class 1 integron integrase gene. They found intI1 in nearly all samples that carried either blaCTX-M-15 or floR. Genome analyses showed that these integrons often hosted clusters of genes conferring resistance to multiple antibiotic families and even to disinfectants, although the main target genes of this study were usually located elsewhere in the genome. This supports the idea that tracking intI1 can serve as a convenient proxy for human‑driven antibiotic pressure and multidrug resistance, while still underscoring the need to directly test for specific high‑risk genes.
From high‑tech lab to field kit
A key strength of the C12a toolbox is its adaptability to different laboratory realities. In a fully equipped setting, reactions were measured with a microplate reader, producing quantitative curves over time. In simpler laboratories, the same reactions could be viewed with a basic blue‑light transilluminator, where only tubes containing resistance genes glowed. For portable, on‑site testing, the team replaced fluorescence readout with lateral flow strips, similar in format to a pregnancy test, which showed bands only when the CRISPR reaction had been activated. Across these setups and sample types—including total DNA directly from feces—the presence or absence of the target genes was consistently identified, with only modest losses in sensitivity in low‑resource modes.
What this means for fighting resistant infections
Together, the results present C12a as a versatile proof‑of‑concept platform for watching antibiotic resistance genes move through people, animals, and the environment. The approach is not yet a clinical diagnostic, and it detects genetic potential for resistance rather than how a specific infection will respond to treatment. Still, its speed, low detection limits, and compatibility with both sophisticated instruments and simple field tools make it a promising building block for future surveillance systems. By making it easier to spot resistance hotspots in hospitals, farms, and waterways, CRISPR‑based toolboxes like C12a could help public health officials and researchers react faster and design smarter strategies to preserve the power of our remaining antibiotics.
Citation: Vargas-Reyes, M., Alcántara, R., Alfonsi, S. et al. Versatile and portable Cas12a-mediated detection of antibiotic resistance markers. Sci Rep 16, 11509 (2026). https://doi.org/10.1038/s41598-026-42073-2
Keywords: antimicrobial resistance, CRISPR diagnostics, Cas12a, antibiotic resistance genes, One Health surveillance