Clear Sky Science · en
Pangenome-based design of strain-specific primers enables precise monitoring of bacteria in human microbiome intervention trials
Why Tracking Helpful Germs Matters
Many of the bacteria we hear about are troublemakers, but some are quiet helpers that turn milk into yogurt, protect our skin, and may even calm inflammation in our airways. These “good bacteria” are sold as probiotics or more strictly regulated live biotherapeutic products, and they are now being tested in human trials for skin problems, allergies, gut issues, and more. Yet a basic question is surprisingly hard to answer: when we give people a specific bacterial strain, can we actually prove that exact strain survives and settles in the body, rather than just its close cousins? This study presents a new way to track individual bacterial strains with high precision, using the growing wealth of genome data.
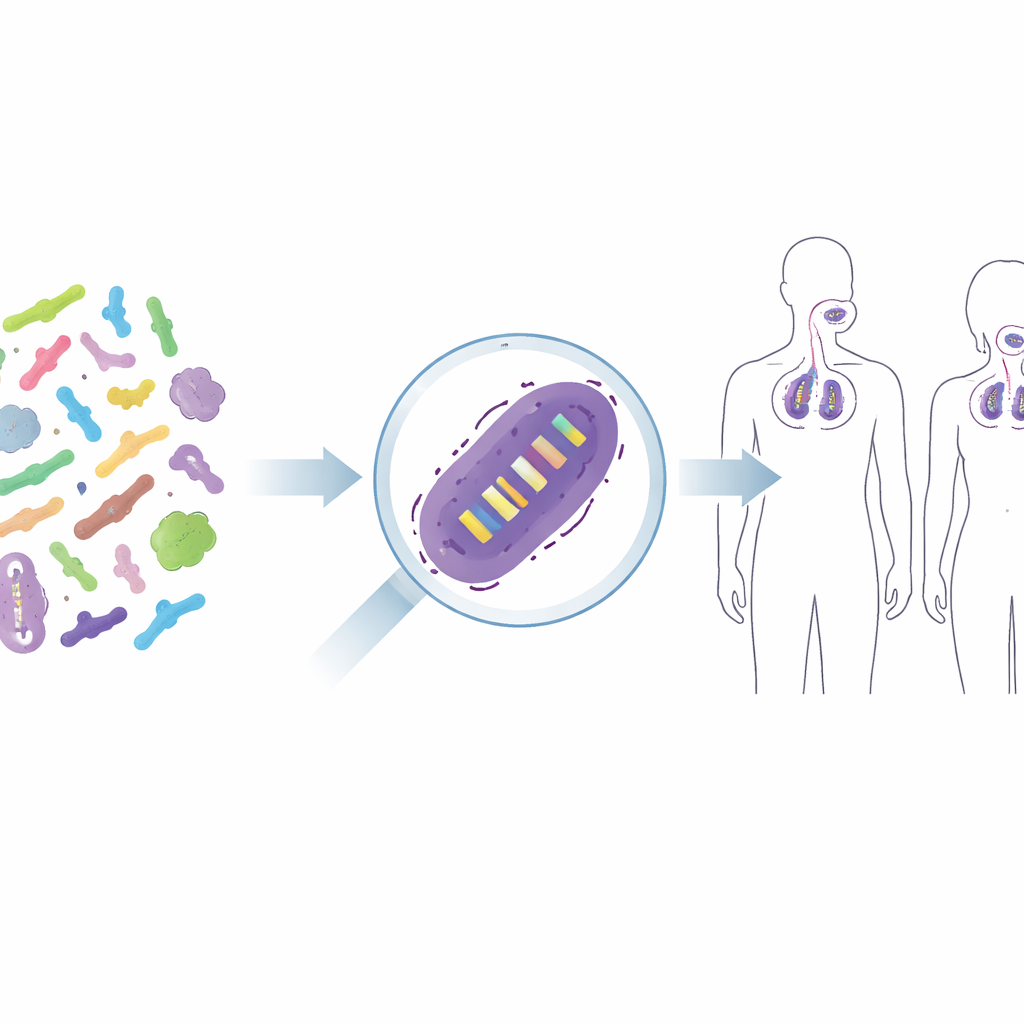
From Mixed Microbes to Individual Strains
Probiotic products rarely contain just “bacteria” in the abstract; they contain carefully chosen strains, like a particular version of Lacticaseibacillus rhamnosus GG or Lactiplantibacillus plantarum WCFS1. These closely related strains can differ in how they interact with our immune system or how well they adhere to skin or throat tissue, so researchers and regulators want to know exactly which strain is present after treatment. Traditional methods, such as growing bacteria on selective media, only tell us about broad groups—like total lactic acid bacteria—and often fail when helpful strains are outnumbered by others. More high‑tech solutions, such as tagging strains with fluorescent markers or DNA barcodes, would turn them into genetically modified organisms, which is often not acceptable for foods or human therapies. The challenge is to distinguish nearly identical strains without altering them.
Mining the Pangenome for Unique Genetic Fingerprints
The authors turned to the concept of a “pangenome,” the full collection of genes found across many genomes of the same species. Some genes are shared by all members (the core), while others are present only in certain strains (the accessory part). By comparing hundreds of genomes for several species of lactic acid bacteria and a common mouth bacterium, they searched for genes that appear in only one strain and not in any other known relative. These rare genes act like genetic fingerprints. The team built a bioinformatics pipeline that automatically assembles the pangenome, removes nearly identical duplicate genomes, and flags strain‑unique genes. They then double‑checked these candidates against broader genome databases to make sure similar sequences were not hiding in other species.
Designing Precise Molecular “Barcode Scanners”
Once strain‑unique genes were identified, the next step was to build highly specific molecular tests around them. The researchers designed short DNA pieces, called primers, that bind only to these unique genes. In a technique known as quantitative PCR (qPCR), these primers act like barcode scanners, copying and detecting only DNA from the target strain. The team created and optimized primers for six strains spanning different species, including widely used probiotics and new candidates isolated from the human body. In the lab, they tested each primer set not only on its intended strain but also on several closely related strains from the same species. The tests showed strong amplification only for the target strain, with minimal signal from the others, confirming that the chosen genetic fingerprints were truly distinctive.
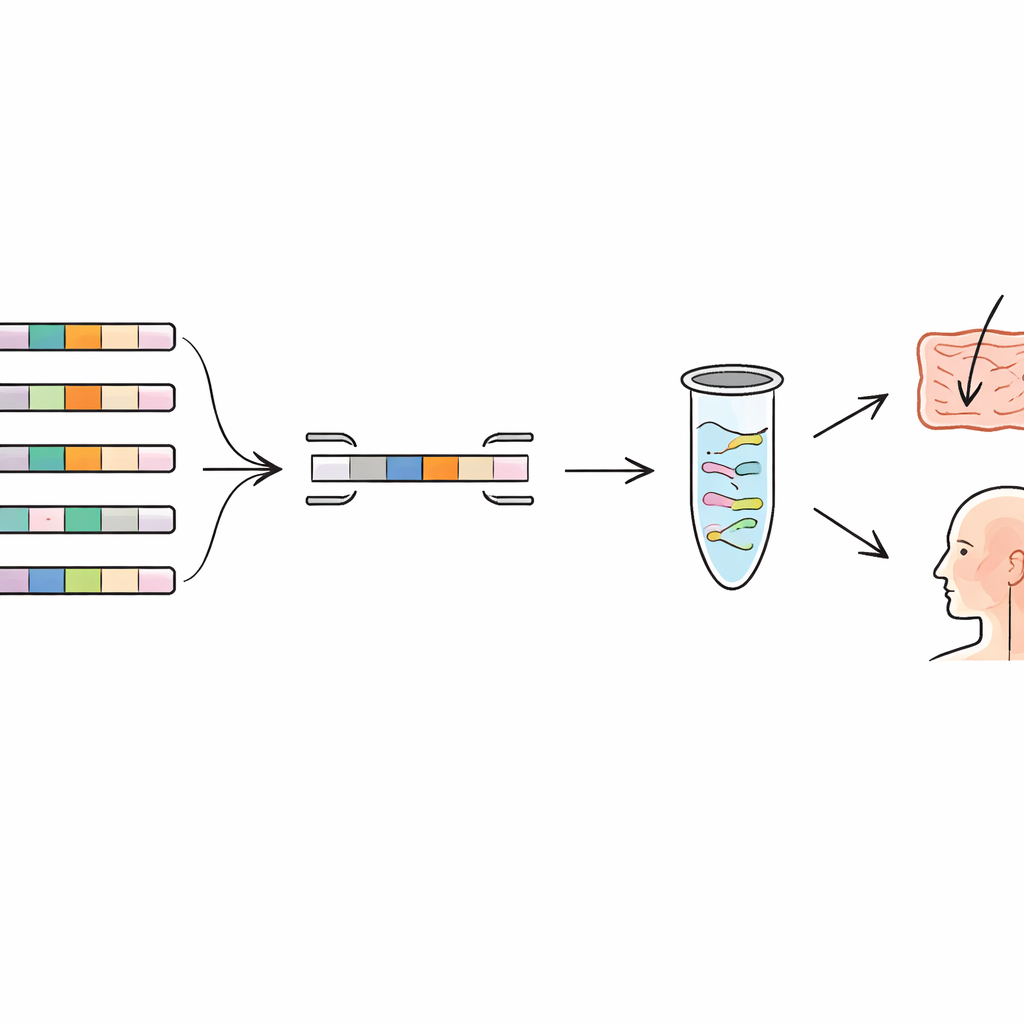
Following Probiotics on Skin and in the Airway
To see whether this approach works in real people, the authors applied their primers to samples from three placebo‑controlled human intervention trials. In one study, volunteers used a face cream containing two probiotic strains; in others, people received a probiotic throat spray or a chewable tablet. Using the new qPCR tests, the scientists could reliably detect the added strains on the skin and in the upper respiratory tract of treated participants, while almost never seeing them in placebo groups. They also observed how the signal rose during use and faded after treatment stopped, giving a clear picture of temporary colonization. In a few cases low‑level signals appeared in controls, reminding researchers to consider natural background bacteria and the risk of very minor contamination, especially in body sites with low bacterial numbers.
What This Means for Future Microbiome Therapies
In everyday terms, this work shows how to build a very sharp “strain radar” without genetically tinkering with the bacteria themselves. By exploiting subtle genetic differences uncovered by large‑scale genome comparisons, scientists can craft qPCR tests that pick out one probiotic strain from a crowd of near look‑alikes. This makes it far easier to answer practical questions: Did the strain survive the product’s shelf life? Did it reach the target body site? How long did it stay, and at what levels? Because the method is transparent, inexpensive, and adaptable to other species, it can become a standard tool for designing and testing next‑generation probiotics and live biotherapeutics, bringing more rigor and safety to the rapidly growing field of microbiome‑based interventions.
Citation: Eilers, T., Delanghe, L., De Boeck, I. et al. Pangenome-based design of strain-specific primers enables precise monitoring of bacteria in human microbiome intervention trials. Sci Rep 16, 11274 (2026). https://doi.org/10.1038/s41598-026-41449-8
Keywords: probiotics, microbiome, strain tracking, qPCR, pangenome analysis