Clear Sky Science · en
A pipeline of machine learning-driven multi-modal data fusion methods for prognostic risk analysis in bevacizumab-treated metastatic colorectal cancer
Why this research matters
For people living with advanced bowel cancer, one of the biggest questions is whether a powerful but costly drug will actually help them. This study explores how patterns in a patient’s tumor DNA, combined with clinical information, can be used with modern machine learning to predict who is likely to benefit from a common targeted treatment, bevacizumab, and who is not. In the future, such tools could spare some patients from side effects and ineffective therapy, while guiding others toward the most promising options.
A closer look at bowel cancer treatment
Metastatic colorectal cancer—bowel cancer that has spread to other organs—is a major cause of cancer death worldwide. Many patients whose tumors carry specific gene changes (RAS mutations) receive standard chemotherapy combined with bevacizumab, a drug that blocks blood vessel growth to starve tumors. While this combination improves survival on average, only a fraction of patients see meaningful benefit. Others endure months of treatment, side effects, and financial cost with little gain. At present, doctors have no reliable test to tell in advance who will not respond to bevacizumab, creating a pressing need for better decision tools.
Bringing many data types together
The researchers built a multi-step analysis pipeline that uses machine learning to fuse several kinds of information from each patient. They drew on a well-characterized European cohort called ANGIOPREDICT, which includes 117 people with metastatic colorectal cancer treated with bevacizumab plus chemotherapy. For each patient they had: regions of the genome that were either gained or lost (copy number alterations), a small set of important gene mutations, and standard clinical details such as age, tumor stage, and tumor location. A specialized tool called PhenMap was then used to uncover hidden patterns—called meta-variables—that summarize how these genetic changes and clinical features vary together across patients.
Finding the DNA signature linked to outcome
Among the ten patterns identified by PhenMap, two were strongly tied to how long patients lived without their disease worsening, a measure called progression-free survival. The team then focused on which specific DNA changes drove these two key patterns. Using additional statistical and machine-learning steps, they narrowed hundreds of genomic regions and mutations down to just three features: losses in two chromosomal regions (15q21.1 and 1p36.31) and mutation in a gene called BRAF. These three features together formed a compact genetic signature that was tightly linked to poorer outcomes in patients who received bevacizumab.
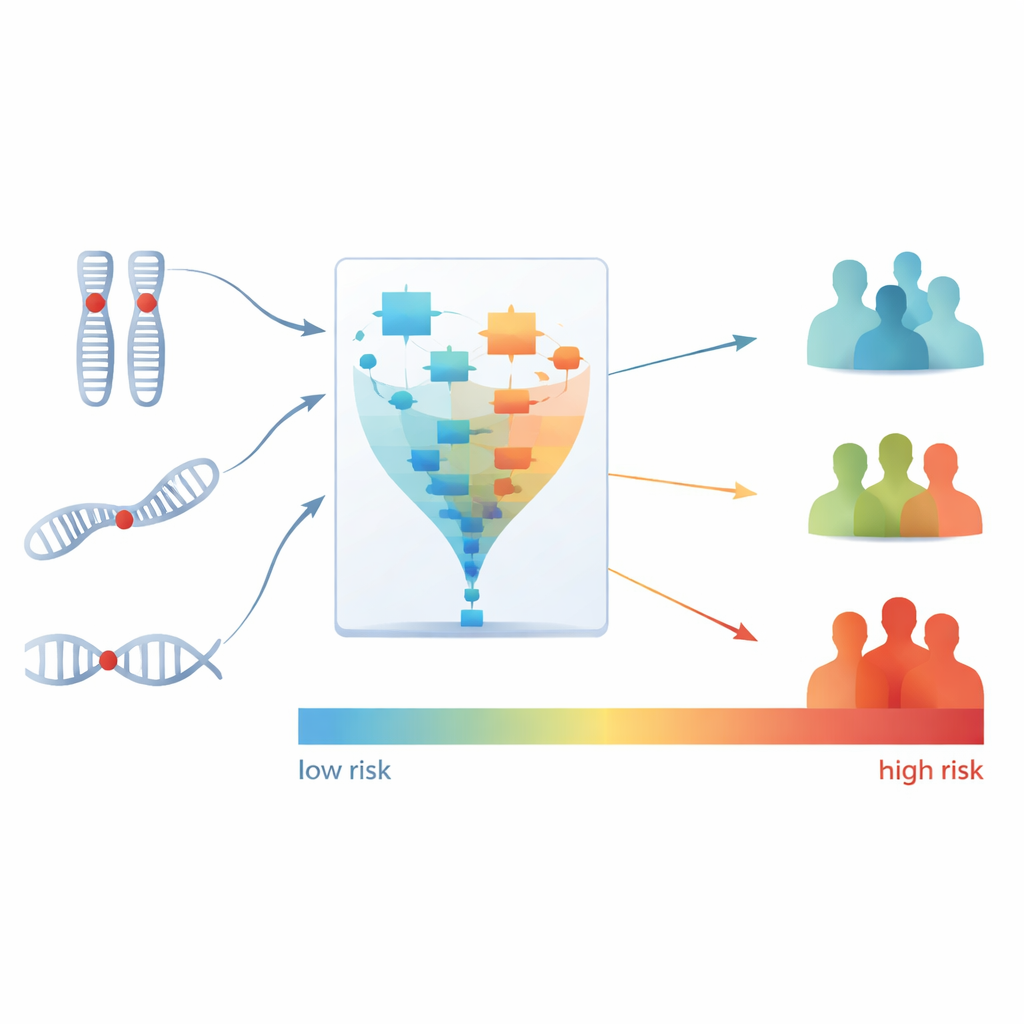
Turning a signature into risk groups
Next, the scientists converted this three-part signature into a single risk score for each patient, reflecting their estimated risk of death while on bevacizumab-based therapy. They then divided patients into three groups—low, medium, and high risk—based on their scores. The differences were striking: every patient in the high-risk group failed to respond to bevacizumab, while most patients in the low-risk group showed a response. The high-risk group also had a much higher chance of early disease progression compared with the low-risk group. Importantly, this risk score offered prognostic information beyond what doctors could already infer from standard clinical factors or previous genomic subtyping alone.
What this could mean for patients
Although this work still needs to be validated in larger and independent patient cohorts, it points toward a future in which complex tumor and clinical data can be integrated into a single, actionable risk score. If confirmed, a simple test that reads out the presence of the two chromosomal losses and the BRAF mutation could help identify metastatic colorectal cancer patients who are unlikely to benefit from bevacizumab combination therapy. Those patients could then be directed toward alternative strategies or clinical trials sooner, while others continue to receive a drug from which they are more likely to benefit. More broadly, the machine learning pipeline demonstrated here could be adapted to other cancers and treatments, advancing the goal of truly personalized cancer care.
Citation: Thomas, V., Nyamundanda, G., Lärkeryd, A. et al. A pipeline of machine learning-driven multi-modal data fusion methods for prognostic risk analysis in bevacizumab-treated metastatic colorectal cancer. Sci Rep 16, 8843 (2026). https://doi.org/10.1038/s41598-026-39189-w
Keywords: metastatic colorectal cancer, bevacizumab resistance, machine learning, genomic biomarkers, precision oncology