Clear Sky Science · en
$${\bf{Micro}}{{\mathbb{S}}}{\bf{plit}}$$ : semantic unmixing of fluorescent microscopy data
Seeing More with Less Light
Modern microscopes let scientists watch living cells in action, but every image comes at a cost. Each flash of light can damage delicate cells, and using many different glowing tags quickly runs into physical limits. This paper introduces MicroSplit, a computational trick that lets researchers capture several cell structures in a single, simpler image and then separate them later on the computer, easing pressure on both the microscope and the cells themselves.
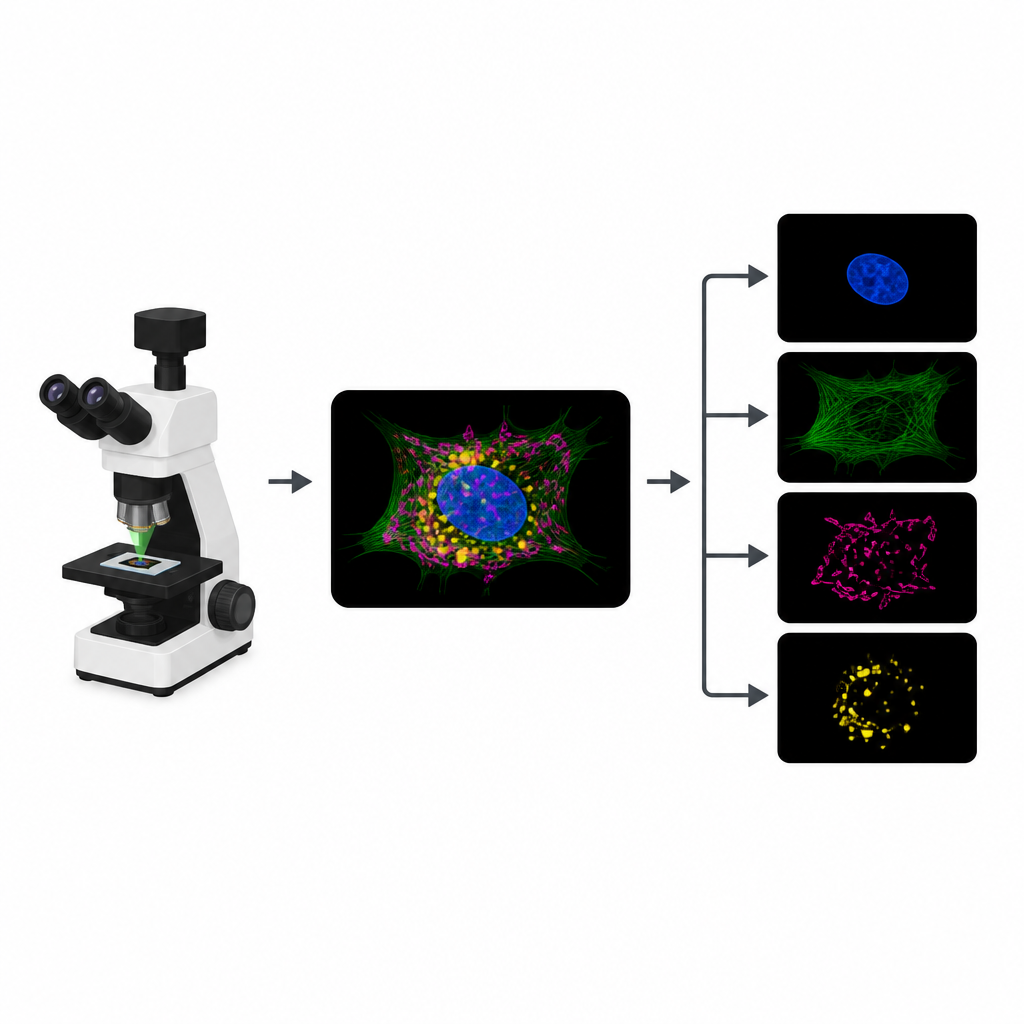
Why Glowing Cells Hit a Wall
To track different parts of a cell, biologists attach distinct fluorescent dyes to structures like the nucleus, skeleton or energy factories. In a typical experiment, the microscope records a separate image channel for each dye. However, the dyes often respond to similar colors of light, so their signals overlap. That overlap limits how many structures can be tracked at once. On top of that, each extra channel means another round of illumination, which eats into a limited “photon budget” and can slow movies, blur details or harm living samples.
Turning One Crowded Picture into Many Clear Ones
MicroSplit tackles this tradeoff by changing how information is collected and processed. Instead of taking several separate pictures, scientists can label multiple structures as usual but record them together in a single fluorescent channel. This single, crowded image is then fed into a deep learning model that has been trained to “unmix” the patterns it sees. The network outputs several cleaned-up images, each one highlighting a different structure as if it had been imaged in its own channel from the start.
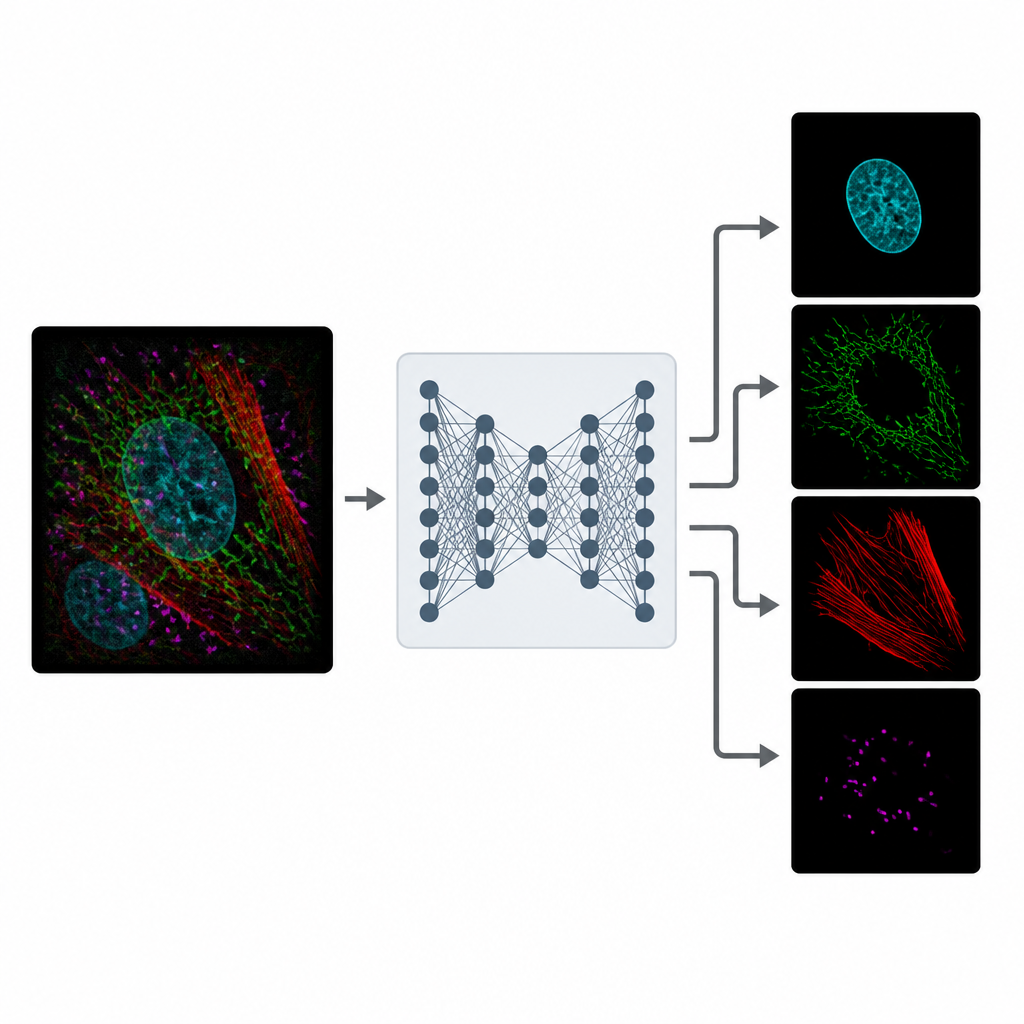
Learning from Noisy Reality
Training MicroSplit does not require pristine reference images. The authors show that the model can learn from ordinary, noisy microscope data, which are much easier to obtain. During training, the system is given examples where separate channels and their summed combination are available. It learns two things at once: how to separate overlapping structures and how to remove noise. Using a special type of neural network that samples many plausible solutions, MicroSplit can also estimate where its own predictions are uncertain, flagging regions that should be treated with caution or checked by an expert.
Working Across Datasets and Uses
The researchers tested MicroSplit on ten different microscopy datasets, spanning two and three dimensions and involving up to four overlapping cellular structures. Across 30 main tasks, the separated images were accurate enough to support standard downstream analyses such as segmenting cell parts. In careful tests, human analysts produced equally consistent segmentations on MicroSplit outputs and on conventional multichannel images. The method can also be turned around to remove unwanted features: by teaching MicroSplit to distinguish true signal from recurring specks or blotches, the model can subtract these imaging artifacts while preserving real structures.
Knowing the Limits
MicroSplit is not magic, and the authors openly explore when it struggles. Performance drops if the original images are extremely noisy, if one structure is much dimmer than another, or if two structures look almost identical. In such cases, the model tends to smooth away very fine details rather than invent false sharpness. Nonetheless, even with short exposure times, MicroSplit often delivers separated images that are good enough for quantitative work, while using far fewer photons than traditional approaches.
What This Means for Future Imaging
For non-specialists, the key message is that MicroSplit lets microscopes “see” more structures in less time and with less light by shifting part of the job from hardware to software. Instead of being limited by how many colors a microscope can separate optically, researchers can combine signals in one channel and let MicroSplit do the sorting. This frees up light to make movies faster, capture dim structures more gently, or add extra labels that were previously impractical. By releasing the code, data and trained models openly, the authors aim to make this kind of computational multiplexing a routine part of biological imaging.
Citation: Ashesh, A., Carrara, F., Zubarev, I. et al. \({\bf{Micro}}{{\mathbb{S}}}{\bf{plit}}\): semantic unmixing of fluorescent microscopy data. Nat Methods 23, 1047–1057 (2026). https://doi.org/10.1038/s41592-026-03082-1
Keywords: fluorescence microscopy, deep learning, image unmixing, computational multiplexing, bioimage analysis