Clear Sky Science · en
3d-OT: a deep geometry-aware framework for heterogeneous slices alignment of spatial multi-omics
Seeing Tissues in Three Dimensions
Biologists are increasingly able to measure not just which molecules are active in a cell, but exactly where that cell sits inside a tissue. These new “spatial” maps promise clearer views of how brains are wired, how tumors spread and how embryos build organs. Yet most experiments still capture only thin slices, each using different measurement types and resolutions. This paper presents 3d-OT, a computational method that knits those slices together into coherent 3D views, while making the most of the rich molecular information in every slice.
Why Mapping Space Matters
Traditional genetics and genomics tell us what molecules cells contain, but not where those cells live. Spatial technologies change that by recording both the molecular profile of each tiny spot in a tissue and its physical coordinates. Different methods emphasize different molecular layers: some profile RNA, others proteins, others the chemical marks on DNA. Each layer reveals a different side of tissue organization. The challenge is that researchers often end up with multiple, partially overlapping slices of tissue measured with different tools. Aligning those heterogeneous slices in a consistent 3D space, without losing fine anatomical detail, has been a major unsolved problem.
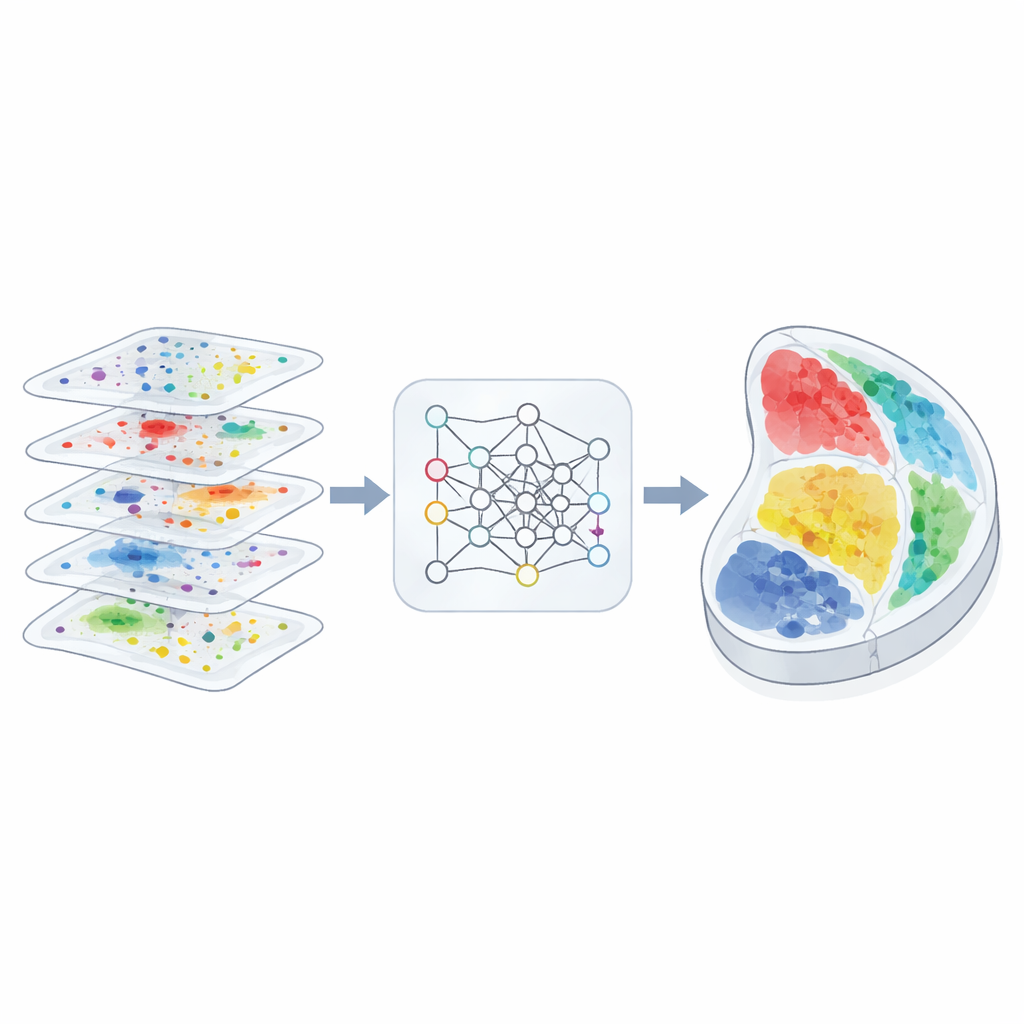
A Geometry-Aware View of Tissues
The authors designed 3d-OT to treat each tissue slice as a “point cloud,” a cloud of spots in physical space, each carrying multi-omics measurements. Instead of relying mainly on a grid or graph to describe neighborhood relationships, 3d-OT uses a neural network architecture called PointNet++ that was originally created for 3D object recognition. This network repeatedly feeds each spot not only its molecular signals but also its exact coordinates, allowing the model to learn subtle geometric patterns such as layered cortex structures or curved organ boundaries. For slices with more than one molecular layer, a fusion module blends the modalities into a shared representation while still keeping the distinctive contribution of each layer.
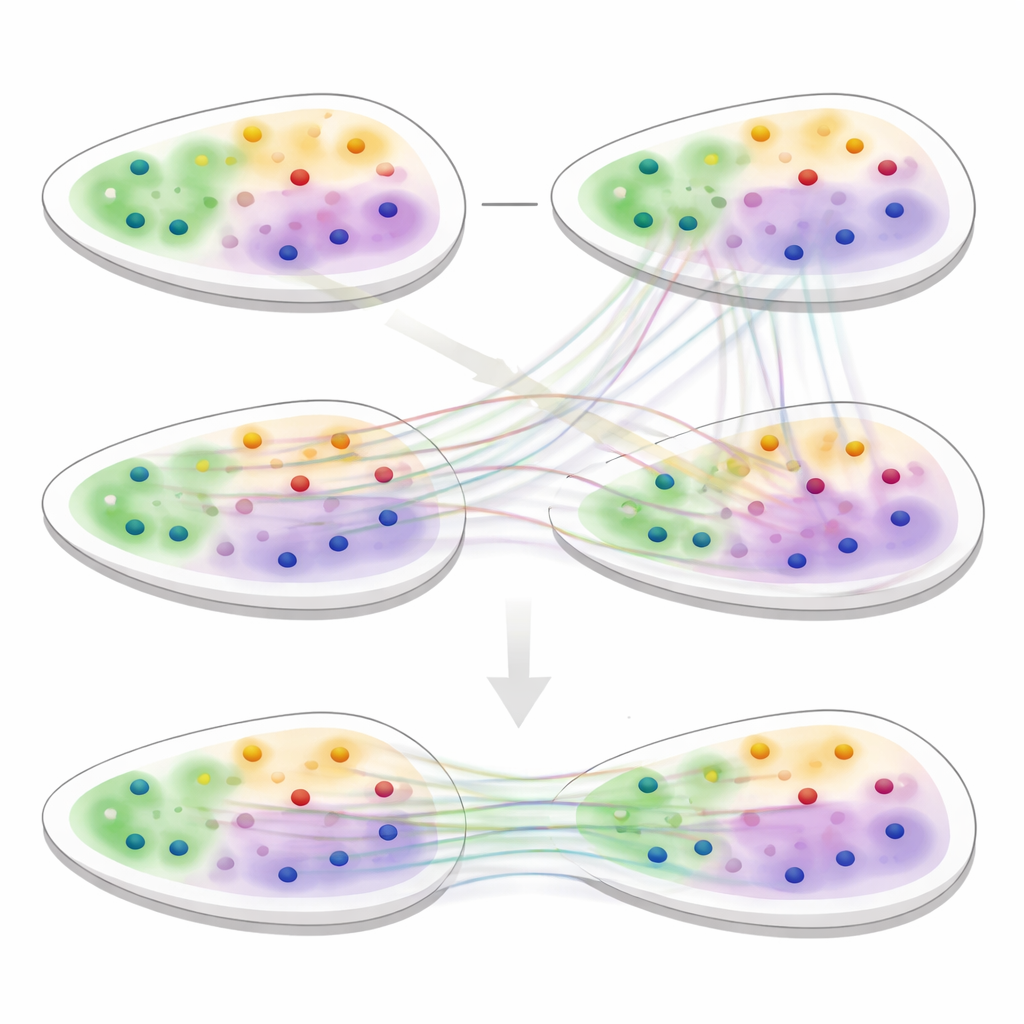
Aligning Slices with Soft Matches
The second core of 3d-OT is an alignment module based on “optimal transport,” a mathematical framework for matching two distributions while preserving their overall shape. Rather than forcing a rigid one-to-one pairing between spots in two slices, the method computes soft correspondences: each spot in a source slice can partially match several candidates in a target slice. A reconstruction loss based on chamfer distance rewards alignments that make the reconstructed positions closely resemble the real target positions. Additional smoothness and “zero-divergence” constraints encourage the flow of spots between slices to follow realistic, continuous tissue deformations, such as bending, stretching or growth, instead of implausible jumps or twists.
Revealing Hidden Tissue Structure
To test 3d-OT, the team compared it against leading spatial analysis tools on a variety of datasets, including human brain, breast cancer, lymph node and multiple mouse organs. In single-slice clustering tasks, 3d-OT more accurately recovered known layers of the human and mouse cortex and better distinguished cancer regions than competing methods. In multi-omics mouse brain data that combined RNA with chromatin marks, it separated two sublayers within the deepest cortical layer that often appear as a single band, and highlighted marker genes for these fine regions. Across multiple technologies and resolutions, including challenging heart and neural crest tissues, 3d-OT provided smoother and more anatomically faithful alignments than existing alignment methods.
Following Organs Through Time
Perhaps the most striking demonstration came from reconstructing mouse embryo development. Using spatial data from several embryonic days, 3d-OT aligned slices across time to build 3D “movies” of organ growth. It captured, for example, how the heart changes from a compact form to a ring-like structure and back, and how the liver expands steadily. The method also linked early precursor regions to later organs, correctly tracing the urogenital ridge into kidney and ovary, and dermomyotome into muscle and connective tissue. These trajectories matched known developmental biology, while also suggesting additional differentiation paths for further study.
What This Means for Future Biology
In simple terms, 3d-OT turns scattered, multi-layered snapshots of tissues into consistent, high-resolution 3D maps that respect both molecular identity and physical shape. By explicitly encoding geometry and allowing flexible, soft matching between slices, it outperforms earlier tools at identifying fine tissue regions and aligning data from different technologies and time points. As spatial multi-omics datasets grow, methods like 3d-OT will help researchers move from flat pictures to dynamic 3D atlases of organs in health, disease and development.
Citation: Dai, B., Yi, L., Wang, P. et al. 3d-OT: a deep geometry-aware framework for heterogeneous slices alignment of spatial multi-omics. Nat Methods 23, 760–771 (2026). https://doi.org/10.1038/s41592-026-03034-9
Keywords: spatial omics, multi-omics integration, tissue 3D reconstruction, embryo development, computational biology