Clear Sky Science · en
Differentiation of sphingomyelin and cholesterol by hyperspectral mid-infrared detection of single-bond vibrational modes in the fingerprint region
Why the fats in our cells matter
Fats, or lipids, are more than just stored calories. They help build cell membranes, steer chemical signals and are tightly linked to conditions such as cancer, heart disease, obesity and neurological disorders. Yet watching specific kinds of lipids inside living cells has been surprisingly hard, often requiring fluorescent dyes that can disturb the very processes scientists want to observe. This study introduces a way to see and tell apart key lipids in living cells without any labels, opening a window onto how these molecules behave in real time.
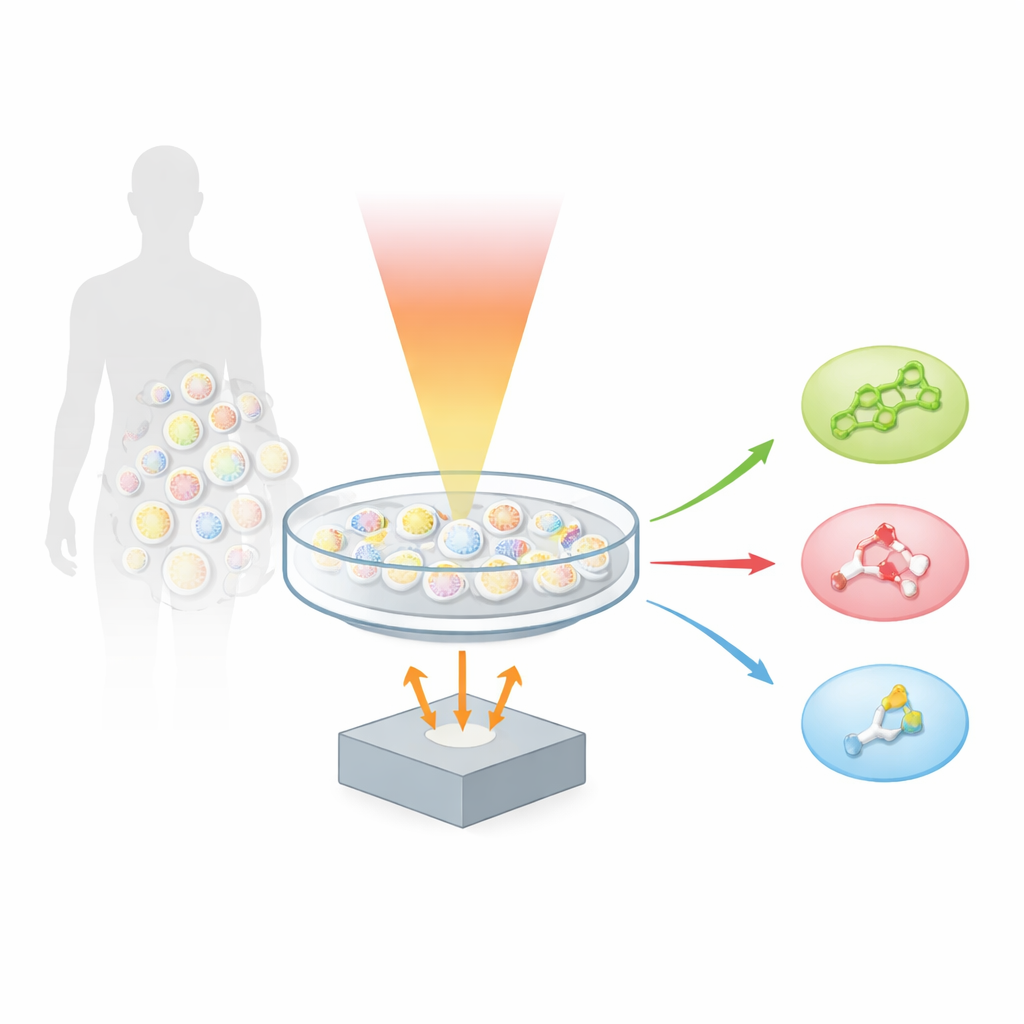
A new way to listen to molecules
The authors build on a technique called optoacoustics, which turns light into sound. When a molecule absorbs a brief pulse of infrared light, it heats and expands ever so slightly, creating a tiny ultrasonic wave. By scanning a sample with many colors of mid‑infrared light and recording the resulting sound, the team created “hyperspectral fingerprint optoacoustic microscopy,” or hyFOPM. Each type of lipid vibrates in a characteristic way when illuminated in the so‑called fingerprint region of the infrared spectrum, roughly between 1,730 and 900 inverse centimeters. These vibrations act like barcodes, allowing the system to distinguish cholesterol from sphingomyelin and from a common membrane lipid called phosphatidylcholine, even though these molecules all live together in the same cell membrane.
Testing the method on simplified samples
To find out how sharply hyFOPM could separate lipids, the researchers first tested it on controlled samples. They prepared tiny wells containing pure cholesterol, sphingomyelin, phosphatidylcholine and water, and scanned them across both the broad carbon–hydrogen stretching region and the more detailed fingerprint region. By mathematically unmixing the recorded spectra, they showed that the fingerprint region holds much richer information. There, cholesterol produced strong signals linked to its rigid ring structure, while sphingomyelin showed distinct features from its amide bonds, and phosphatidylcholine revealed a clear signature from its ester groups. Statistical analysis confirmed that these spectral fingerprints allowed the three lipids to be classified with about 96–97% accuracy, with the fingerprint region alone giving the cleanest separation and the least cross‑talk between species.
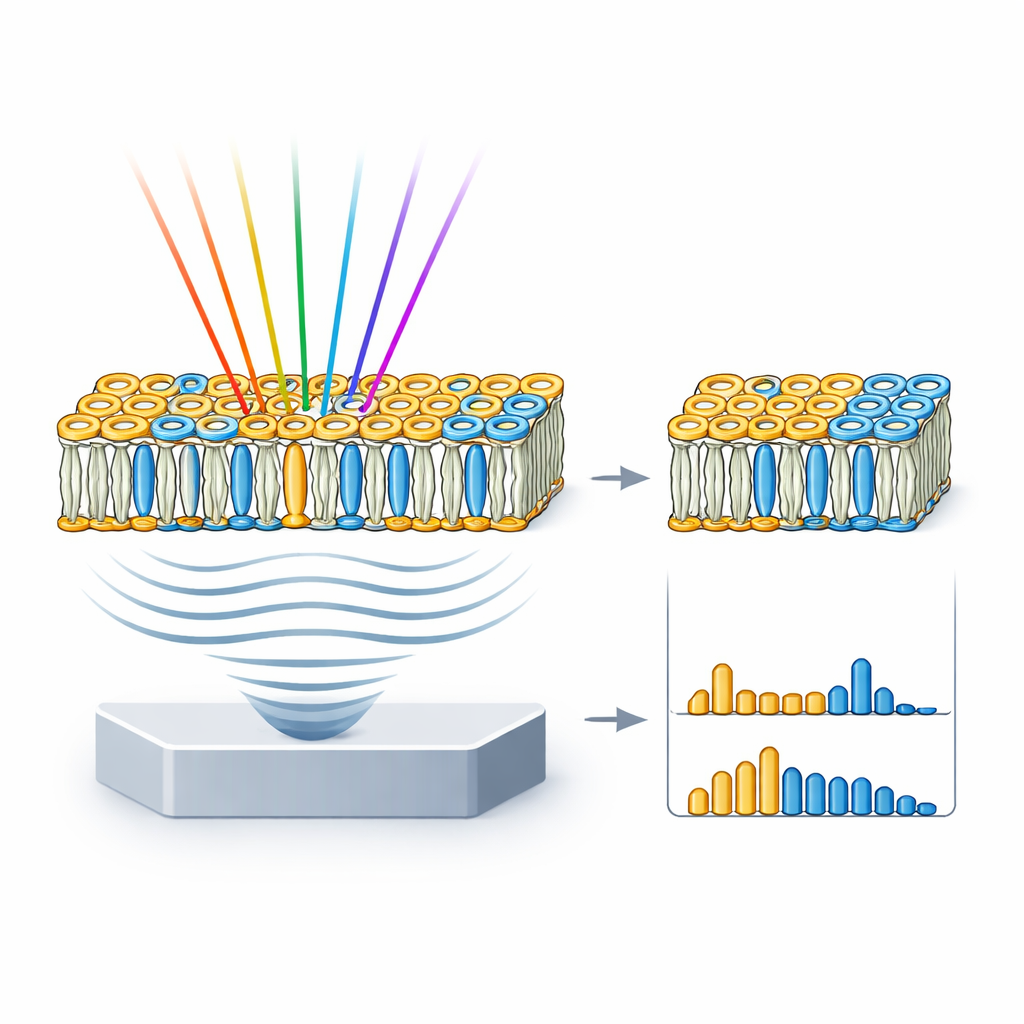
From model membranes to realistic cell surfaces
Real cell membranes are more complex than pure lipid wells, so the team next turned to giant unilamellar vesicles, or GUVs—cell‑sized bubbles made from chosen mixtures of lipids. They assembled three types of GUVs: one densely packed with just sphingomyelin and cholesterol; one that mimics the mixed, patchy nature of a typical cell membrane; and one made only of a fluid phosphatidylcholine. HyFOPM images revealed these vesicles clearly and showed that tightly packed membranes produced stronger signals than looser ones. By comparing the vesicle spectra with those of the pure lipids, and unmixing the contributions, the researchers could recover the actual lipid composition of each vesicle and even detect batch‑to‑batch variation of up to about 40%. This means the technique can double as a quality‑control tool for membrane models often used in biophysics and drug research.
Watching disease‑linked lipids change inside living cells
The real test was whether hyFOPM could track specific lipids inside living cells as they responded to treatments. In lung cancer cells exposed to an experimental anti‑tumor compound that is known to boost sphingomyelin, the method measured a more than two‑fold rise in the spectral feature linked to that lipid over 72 hours, while control cells showed only a modest drift. In a second experiment, human kidney cells were flooded with cholesterol delivered by carrier molecules. HyFOPM detected a sharp increase in cholesterol‑related signals and a small decrease in sphingomyelin, consistent with known effects of this treatment on cell membranes. Crucially, these measurements were achieved using just a handful of carefully chosen infrared colors, suggesting that future systems could be made faster by sampling only the most informative spectral bands.
What this means for health and disease
By providing label‑free, chemically specific maps of lipids in living cells, hyFOPM sidesteps many drawbacks of fluorescent stains and genetic tags. Researchers can now differentiate major lipid classes such as cholesterol and sphingomyelin directly from their vibrational fingerprints, follow how they accumulate or vanish during drug treatments, and assess the true composition of model membranes. Because mid‑infrared optoacoustic signals can be detected at depths larger than conventional microscopy allows, future versions of this approach may image lipid imbalances in tissues and even in patients. That could eventually help diagnose and monitor diseases—from atherosclerosis to neurodegeneration—that are driven in part by the hidden choreography of fats inside our cells.
Citation: Gasparin, F., Prebeck, A., Soldà, A. et al. Differentiation of sphingomyelin and cholesterol by hyperspectral mid-infrared detection of single-bond vibrational modes in the fingerprint region. Nat Methods 23, 815–822 (2026). https://doi.org/10.1038/s41592-026-03025-w
Keywords: lipid imaging, cholesterol, sphingomyelin, optoacoustic microscopy, mid infrared spectroscopy