Clear Sky Science · en
KDM3A catalyses the oxidation of acetyl-lysine to hydroxyacetyl-lysine on histone H3K9
How Cells Tune Genes with Tiny Chemical Switches
Every cell in your body carries the same DNA, yet heart cells, brain cells and skin cells behave very differently. One reason is that cells decorate their DNA-packaging proteins, called histones, with small chemical tags that act like switches for turning genes on or off. This paper reveals a surprising new kind of switch on a key histone site and shows that it is directly controlled by oxygen, linking how cells read their genes to how they sense the air they breathe.
A New Twist on a Well-Known Histone Tag
Histones are spools around which DNA is wrapped. Cells attach chemical groups such as acetyl or methyl groups to specific spots on histones to loosen or tighten DNA, influencing which genes are active. A famous hotspot is lysine 9 on histone H3 (H3K9), where acetylation tends to promote gene activity, while certain methyl marks are associated with gene silencing. Enzymes called histone acetyltransferases add acetyl groups, and deacetylases remove them; other enzymes add or erase methyl groups. Until now, these changes were thought to be “redox-neutral” – they did not explicitly depend on oxygen. 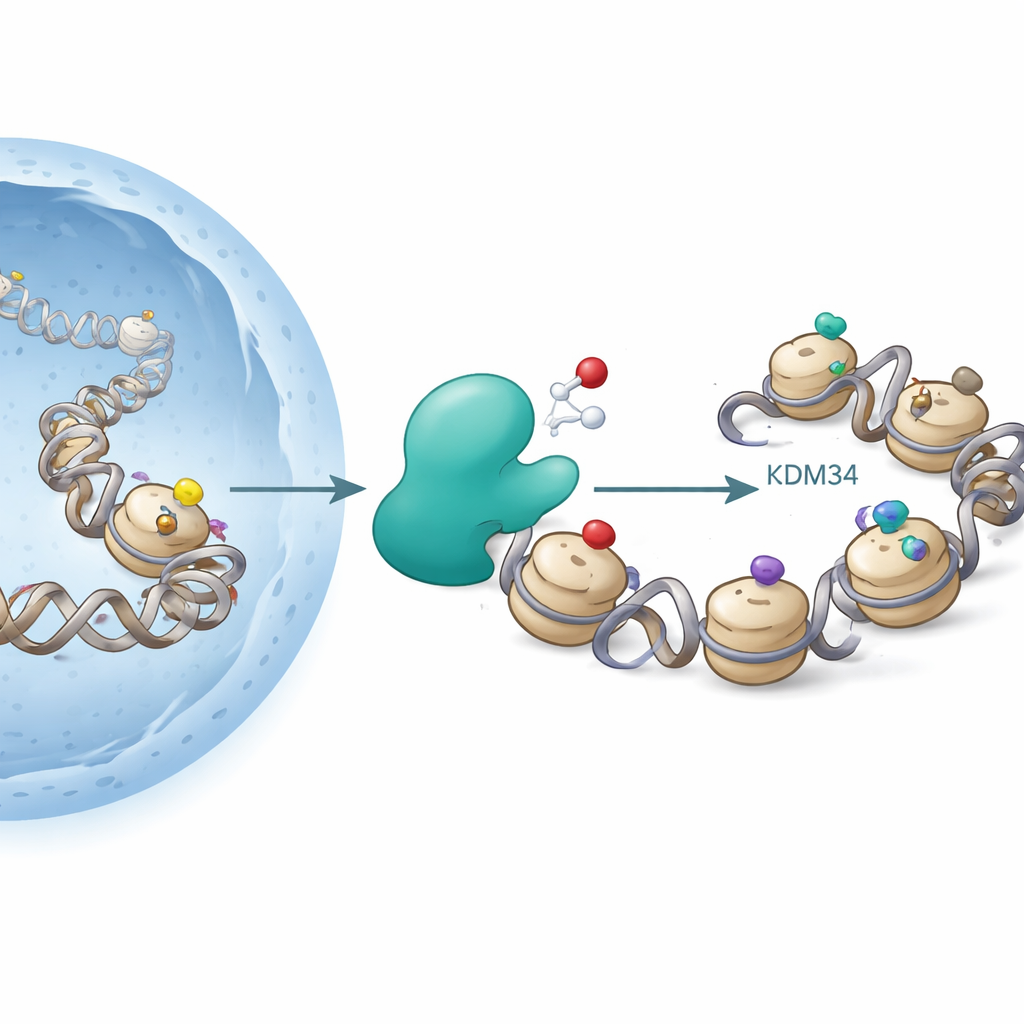
Discovering an Oxygen-Linked Histone Mark
The authors focused on KDM3A, an enzyme already known for removing methyl groups from H3K9. KDM3A belongs to a family of oxygen-dependent enzymes that require iron and a small molecule cofactor (2-oxoglutarate) to work, similar to the enzymes that sense low oxygen (hypoxia) in our tissues. When the researchers screened various modified histone fragments as possible KDM3A targets, they confirmed that KDM3A efficiently erased certain methyl marks on H3K9. Unexpectedly, they also found that purified KDM3A could oxidize the acetyl group on H3K9, converting it into a new form called hydroxyacetyl-lysine. This change corresponds to adding an oxygen atom to the acetyl group, and further experiments showed that under some conditions the enzyme can oxidize even further, hinting at a pathway to more exotic chemistries.
Proving the New Mark Exists in Cells
Finding a new chemical product in a test tube is only the first step; the team also needed to show that hydroxyacetyl-lysine appears on histones inside living cells. They generated a dedicated antibody that recognizes H3K9 carrying the hydroxyacetyl group but not the standard acetyl form. Using this tool, they showed that adding purified KDM3A to isolated histones increased the signal for the new mark, while known methyl marks on H3K9 decreased, as expected from KDM3A’s dual activity. Overexpressing KDM3A in human cells had similar effects: methyl marks dropped, and the hydroxyacetyl signal rose on bulk histones and at the single-cell level. Mass spectrometry, a highly sensitive technique for weighing molecules, independently confirmed that the precise H3K9 peptide carrying hydroxyacetyl-lysine is present in cells when KDM3A is active.
Where the New Mark Sits on the Genome
To see where this mark appears along DNA, the researchers performed ChIP-seq, a genome-wide mapping method. They found that H3K9 hydroxyacetylation clusters around gene start sites, especially at genes that are already highly expressed. Its pattern closely tracks that of classic “active” marks such as H3K9 acetylation and H3K4 trimethylation, suggesting that the new tag also belongs to the toolkit of active chromatin. Treating cells with a histone deacetylase inhibitor, a type of drug already used in cancer therapy, increased both standard acetylation and the new hydroxyacetyl mark, likely by boosting substrate availability for KDM3A and slowing removal of these acyl groups. 
How Other Proteins Read and Process the New Tag
The study also tested how existing chromatin “reader” and “eraser” proteins respond to the novel mark. A reader domain from the AF9 protein, which normally recognizes acetyl and crotonyl tags, bound the hydroxyacetyl version with only modestly reduced affinity, implying that known readers can sense this modification. Two deacetylating enzymes, SIRT1 and HDAC8, processed hydroxyacetyl-lysine differently: SIRT1 was much less efficient in removing the new tag than the standard acetyl group, whereas HDAC8 had little preference. These differences suggest that hydroxyacetylation can subtly alter how long a mark persists and which enzymes engage with it, fine-tuning the chromatin landscape.
Why This Oxygen-Dependent Switch Matters
Overall, the work uncovers H3K9 hydroxyacetylation as a previously unrecognized histone mark produced by KDM3A, and probably in some contexts by its close relative KDM3B. Because KDM3A is itself activated by low oxygen via the hypoxia-inducible factor pathway, this creates a direct chemical link between oxygen levels and histone acetylation states. In simple terms, the enzyme can both remove repressive methyl tags and convert activating acetyl tags into a slightly different oxygen-containing form at the same histone site, potentially reshaping gene programs during stress, development or cancer. The discovery broadens our view of what “demethylase” enzymes can do and opens the door to exploring similar hydroxyacetyl marks on other proteins, as well as their roles in health and disease.
Citation: Belle, R., Bukowski, JP., Schiller, R. et al. KDM3A catalyses the oxidation of acetyl-lysine to hydroxyacetyl-lysine on histone H3K9. Nat. Chem. 18, 823–834 (2026). https://doi.org/10.1038/s41557-026-02112-x
Keywords: histone modification, epigenetics, oxygen sensing, KDM3A enzyme, gene regulation