Clear Sky Science · en
A double-staining automated flow cytometry method for real-time monitoring of bacteria in continuous bioreactors
Watching Microbes Work in Real Time
Modern biotechnology relies on huge cultures of bacteria to make everything from medicines to crop protectants. Yet, inside those foaming tanks, it is surprisingly hard to know what the microbes are doing at any given moment. This study introduces an automated microscope-like system that can watch individual bacterial cells every hour, revealing not just how many are present but whether they are actively growing and copying their DNA. Such real-time insight could make industrial fermentation more efficient, reliable, and easier to control.

Why Counting Cells Is Not Enough
In industrial fermenters, cell density helps determine when to feed more nutrients, when to harvest, and how productive a process can be. Traditional tools—like optical density readings, dry weight measurements, or electrical sensors—measure the bulk cloudiness or mass of a culture. They are useful but crude: they cannot distinguish living from inactive cells, overlook differences in size and shape, and say nothing about whether cells are preparing to divide. As a result, operators often fly partly blind, adjusting processes based on delayed or indirect signals.
A New Automated Window Into Bacterial Life
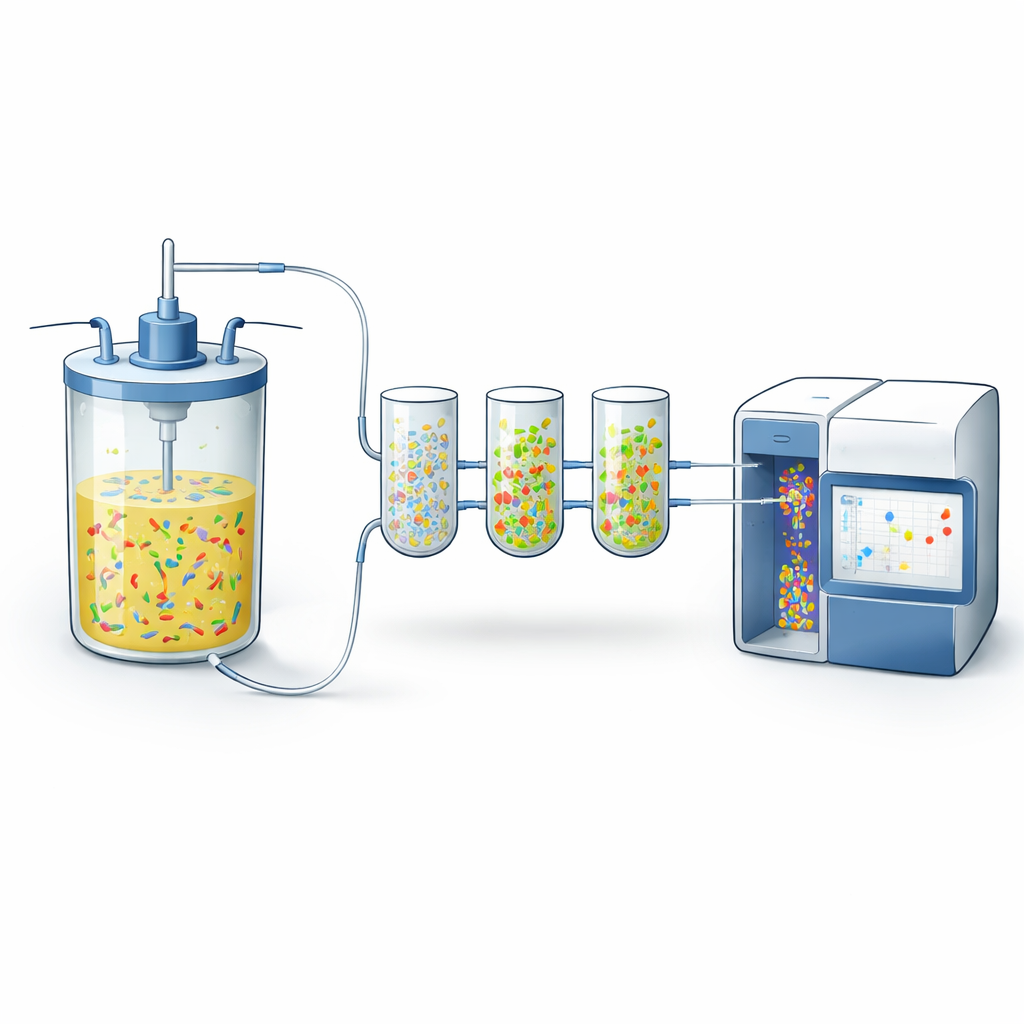
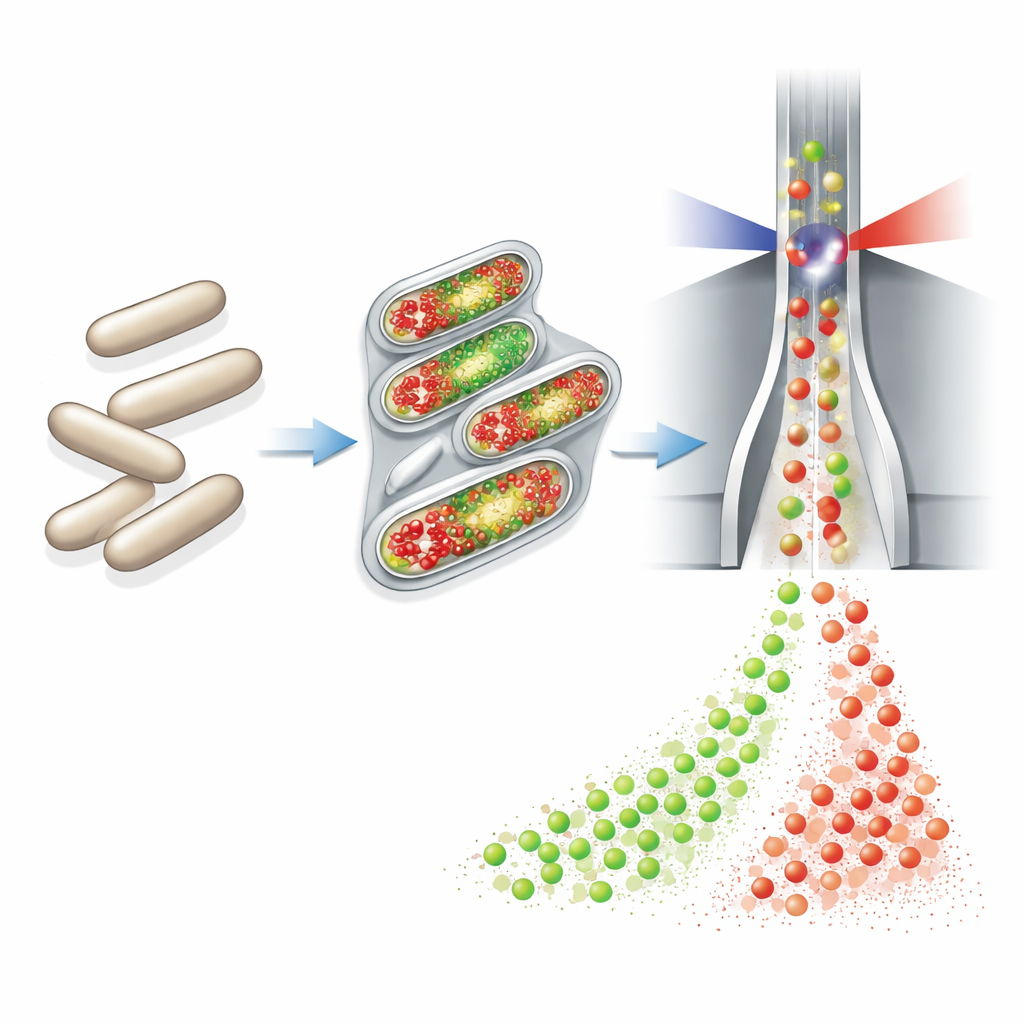
The authors built on a technology called flow cytometry, which rapidly measures properties of thousands of individual cells as they pass a laser beam. They connected a fully automated sampling and staining device to small continuous bioreactors. Every hour, this device drew a tiny volume of culture, chemically fixed and permeabilized the cells, added two fluorescent dyes, diluted the sample, and sent it directly to a cytometer. One dye (DAPI) glows in proportion to how much DNA each cell contains, serving as a sensitive indicator of cell growth and chromosome copy number. The second dye, attached through a “click” reaction to a DNA building block called EdU, specifically marks cells that are in the act of copying their DNA—those on the verge of division.
Following Bacterial Growth, One Cell at a Time
The team tested this setup on three non-pathogenic, Gram-negative bacteria: a soil-dwelling Bradyrhizobium strain, the model species Escherichia coli, and the plant-associated Stenotrophomonas rhizophila. In continuous cultures, they varied how fast fresh medium flowed through the reactors (the dilution rate) and let the automated system run. The cytometer reported both cell counts and the average DAPI signal per cell, which reflects DNA content. Across conditions, a rise in DAPI signal reliably appeared before visible increases in cell number or changes in oxygen use. This early warning showed that cells were ramping up DNA replication and growth long before traditional indicators would flag a shift, allowing, in principle, earlier adjustments to feeding or operating conditions. The method also captured how each species entered and maintained steady growth at different flow rates, and how, at very high dilution, cultures thinned out but were enriched in faster-growing cells with higher DNA content.
What Worked—and What Did Not
In batch experiments (closed flasks or plates), the EdU-based dye successfully highlighted bursts of DNA replication, peaking soon after inoculation and then declining as cultures slowed. It tracked the appearance and disappearance of subgroups of cells with many chromosome copies, confirming that it reliably marks actively replicating cells. However, when EdU was continuously supplied in the medium of the small reactors, it often slowed or disturbed growth. For S. rhizophila, it almost halted the culture; for Bradyrhizobium, it strongly reduced cell numbers; and even E. coli grew more slowly and showed replication signals only in the early hours. Increasing the amount of fluorescent dye did not fix this problem. The authors conclude that, under continuous exposure, EdU is too growth-inhibiting and technically fussy to serve as a routine replication marker in industrial monitoring.
Implications for Smarter Bioreactors
Despite the EdU limitations, the double-staining workflow proved that automated online flow cytometry can deliver rich, time-resolved pictures of bacterial physiology. Counting cells and tracking their DAPI signal gave a robust, direct measure of biomass and an early indicator of growth changes at different dilution rates. The integrated sampler–stainer–cytometer chain operated hourly with minimal human intervention, showing that such systems can be woven into future production lines. While the specific replication dye tested here is not yet ready for day-to-day control of bacterial factories, the same automated framework can be paired with other fluorescent markers. In the long run, this kind of real-time, single-cell monitoring could help keep bioreactors in their optimal “sweet spot,” improving yields, stability, and product quality for a wide range of microbial processes.
Citation: López-Gálvez, J., Schönfelder, E., Mayer, H. et al. A double-staining automated flow cytometry method for real-time monitoring of bacteria in continuous bioreactors. npj Syst Biol Appl 12, 47 (2026). https://doi.org/10.1038/s41540-026-00694-3
Keywords: automated flow cytometry, bacterial bioreactors, real-time bioprocess monitoring, cell cycle and DNA replication, industrial biotechnology