Clear Sky Science · en
Context-aware synthetic promoter design using neural networks enables rewiring of eukaryotic transcriptional networks
Teaching Cells New Tricks
Modern biotechnology often hinges on a simple question: can we tell a cell exactly when to turn a gene on or off? This paper explores a new way to do just that in baker’s yeast, using artificial intelligence to redesign tiny DNA switches called promoters. By learning how these natural switches are wired, the authors show that computers can propose precise edits that let scientists plug new control knobs into the cell’s genetic circuitry.
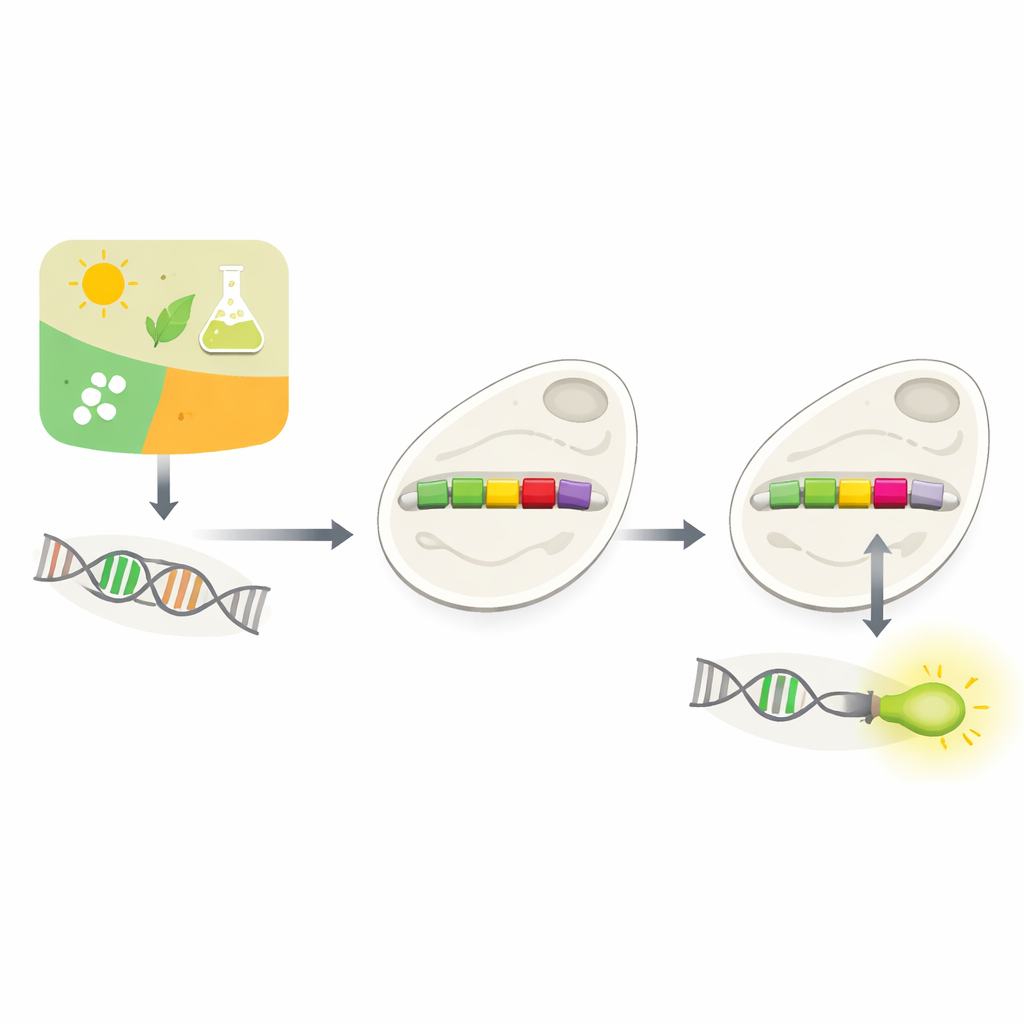
How Cells Decide What Genes To Use
Every cell must constantly decide which genes to use, and when. Much of this decision-making happens at promoters, short stretches of DNA that sit just before genes. Promoters act like dimmer switches, responding to signals from the environment or from within the cell. Biologists have long tried to build custom promoters so they can control genes at will—for example, to make yeast brew valuable chemicals or to sense toxins. But natural promoters are complicated and context-dependent, and simply dropping new control elements into them often breaks their function. Experimentally testing many possible designs is slow and expensive.
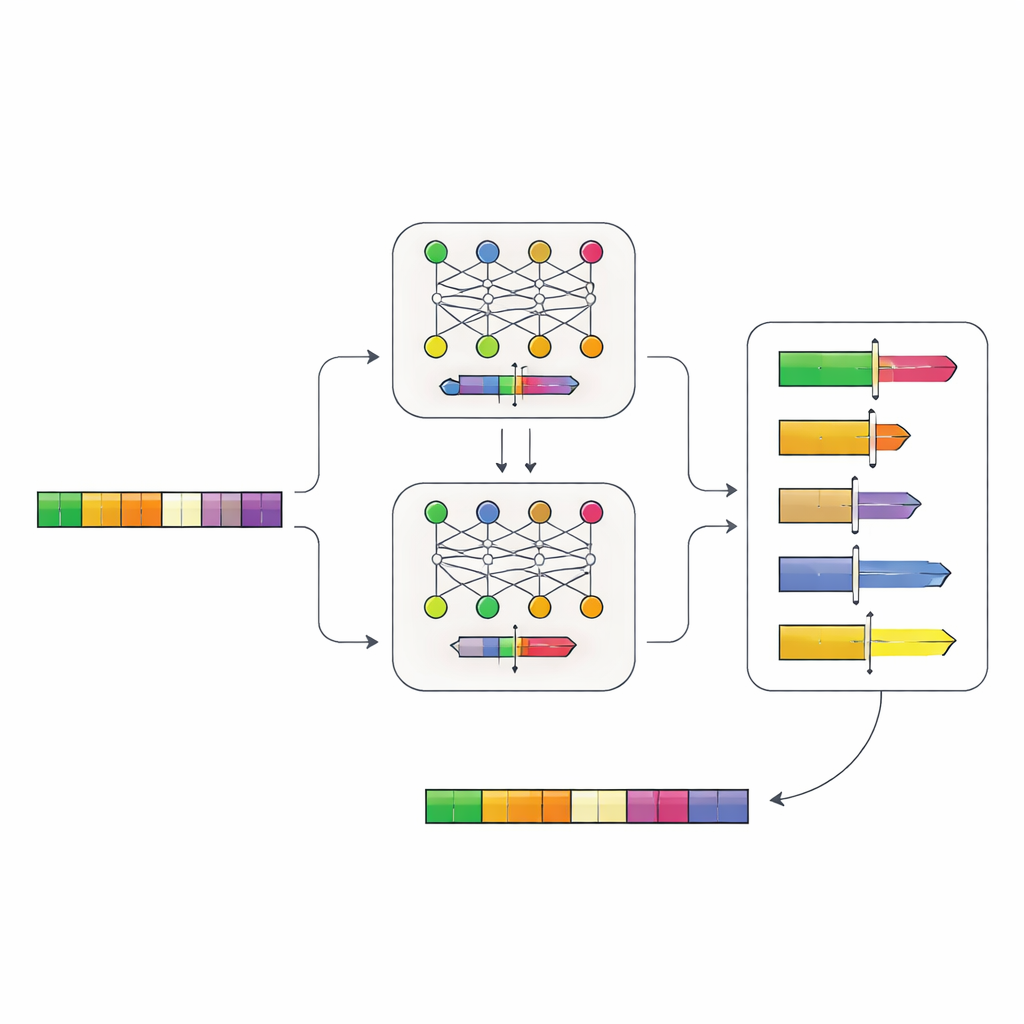
Letting Neural Networks Read DNA Logic
The authors built a two-step neural network system that learns promoter “grammar” directly from DNA. First, they collected promoter sequences from more than 100,000 genes in yeast relatives and trained a model, called Place-Back, to recognize where a short DNA segment originally came from after it had been scrambled and partially replaced. Because the model had to recover the right spot from context alone, it learned subtle patterns about which regions of a promoter can be changed without disturbing its core function. A second model, the Determiner, then looked at Place-Back’s predictions and decided both where a new regulatory element should go and how large a region of DNA should be rewritten.
Designing Switchable Promoters In Silico
Armed with this system, the team asked a practical question: where could they insert a well-known binding site for the TetR repressor protein into real yeast promoters, so that genes could be turned off on command? They virtually tested all 6,011 promoters in the yeast genome and produced a ranked list of thousands of candidates where the new element should “fit” without disrupting essential features, such as the core start region where transcription begins. The models often favored sites close to these core regions while still avoiding direct damage, suggesting they had learned biologically meaningful constraints.
Putting AI-Designed Switches To The Test
To see whether the computer’s suggestions worked in living cells, the researchers chose four native yeast promoters with different strengths and edited each one exactly as recommended, inserting the TetR binding site at the predicted location. They attached these promoters to a bright luciferase reporter and introduced TetR on a separate DNA piece, so the new system would not interfere with yeast’s own regulation. In the presence of TetR, three of the four designs showed strong repression, and one achieved nearly complete shutoff—about 98% reduction in activity—without any further tuning. Alternative insertion sites not suggested by the model often crippled the promoter or destroyed its ability to respond, underscoring that placement matters and that the neural network was identifying specific “sweet spots.”
Rewiring Yeast’s Native Decision-Making
The team then used the same approach to alter yeast’s own regulatory network, rather than an artificial add-on system. They edited the promoter of PCF11, a gene essential for survival, to insert a binding site for Mig1, a natural repressor that becomes active in the presence of glucose. In test assays, the redesigned promoter behaved as intended: when glucose was present, PCF11 activity dropped by more than half; when glucose was absent, activity was slightly higher than normal. After swapping this synthetic promoter into the yeast genome, cells grew almost normally in low sugar but reached a lower maximum density in high sugar, showing that the essential gene was now conditionally constrained by the new connection.
What This Means For Future Genetic Design
In simple terms, this work shows that neural networks can learn enough about the “language” of gene control to suggest smart, context-aware edits to DNA switches. Without needing experimental measurements as training labels, the models point to places where new control elements can be safely inserted so that genes become responsive to chosen signals. The successful tests in yeast, including rewiring control of an essential gene, suggest a path toward scalable, predictive design of regulatory DNA in many organisms. This could speed up the creation of bespoke genetic programs for medicine, agriculture, and industrial biotechnology, while revealing new rules about how cells naturally organize their own decision-making circuits.
Citation: Kuhajda, L., Honzik, T., Svec, J. et al. Context-aware synthetic promoter design using neural networks enables rewiring of eukaryotic transcriptional networks. npj Syst Biol Appl 12, 65 (2026). https://doi.org/10.1038/s41540-026-00684-5
Keywords: synthetic biology, gene regulation, neural networks, promoter engineering, yeast