Clear Sky Science · en
Critical evaluation of drug response prediction models with DrEval
Why smarter cancer drug tests matter
When doctors choose cancer drugs, they often rely on broad categories like tumor type and a handful of genetic markers. In the last decade, scientists have begun testing hundreds of drugs on hundreds of lab-grown cancer cell lines, while also measuring each cell line’s genes and other molecular features. Many computer models claim to predict which drugs will work based on these data. This article takes a hard look at those claims and introduces DrEval, a new evaluation toolkit that tests how well such models really perform in situations that resemble real medical decisions. 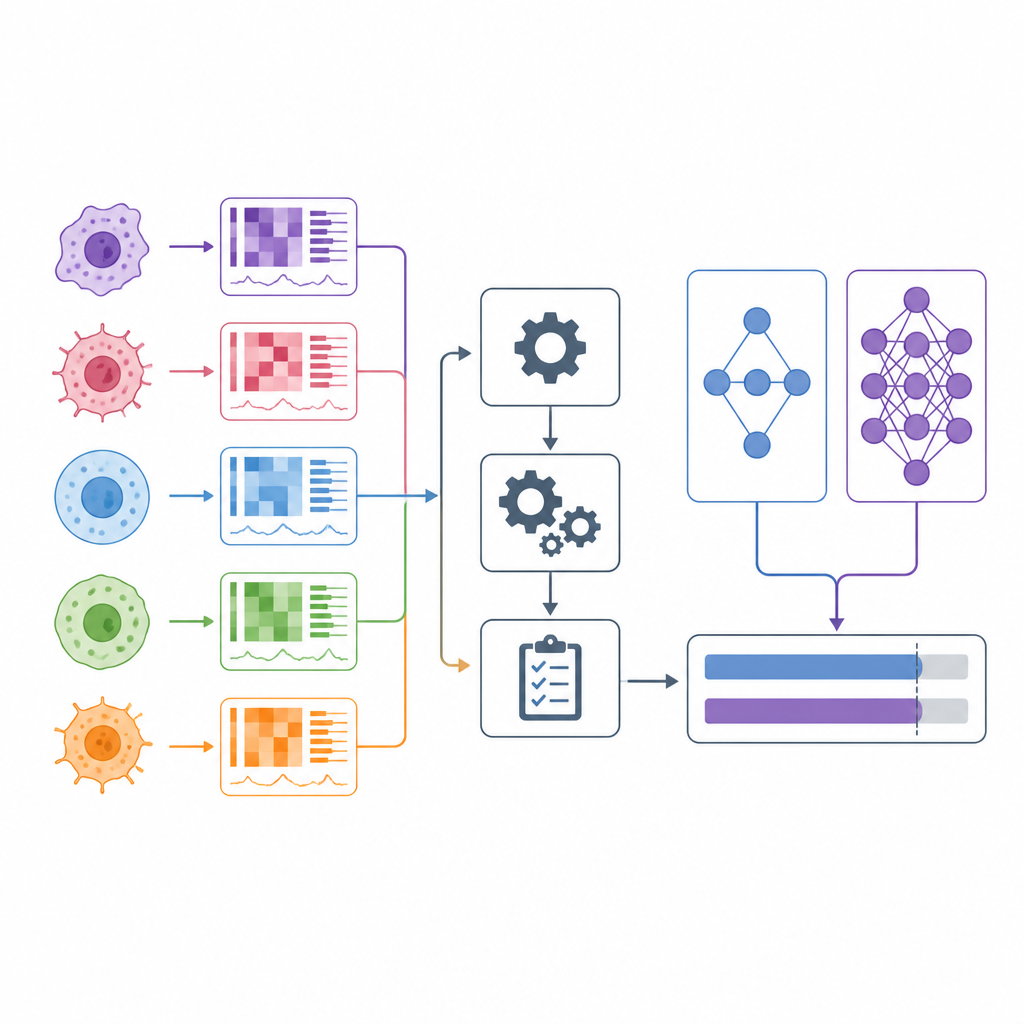
How computers try to guess drug success
Drug response prediction models learn from large screening projects where cancer cell lines are exposed to many drugs at different doses and their survival is measured. Researchers feed each model detailed molecular profiles of the cell lines and various descriptions of the drugs, then ask it to predict standard summary measures of how strongly each drug slows cell growth. Some approaches train one model per drug, while others train a single model across many drugs in hopes it can even estimate responses for new medicines. On paper, these methods often report impressive accuracy numbers, suggesting that personalized cancer treatment might be within reach.
Hidden pitfalls in current evaluations
The authors show that these optimistic results often come from how the data are split and scored, not from genuine insight into cancer biology. If the same cell line or drug appears in both training and test sets, the model may simply memorize typical behavior instead of learning deeper patterns. Because different drugs work at very different dose ranges, a model that only memorizes average response per drug can already explain much of the variation in the data. This creates a statistical illusion where global accuracy looks high, even though the model fails to distinguish which specific cell lines are more or less sensitive to a given drug.
What DrEval does differently
DrEval is a open benchmarking framework that standardizes how drug response models are tested. It provides harmonized datasets, careful data cleaning, and several realistic testing scenarios that mirror common goals: predicting responses for new patients (new cell lines), for new cancer types (new tissues), or for entirely new drugs. It also includes simple baseline methods, such as a predictor that uses only the average effects of each drug and cell line, and tree-based models that are much easier to train than deep neural networks. DrEval runs all models under the same cross-validation and tuning scheme, then reports results with metrics that remove the misleading influence of average drug and cell line effects. 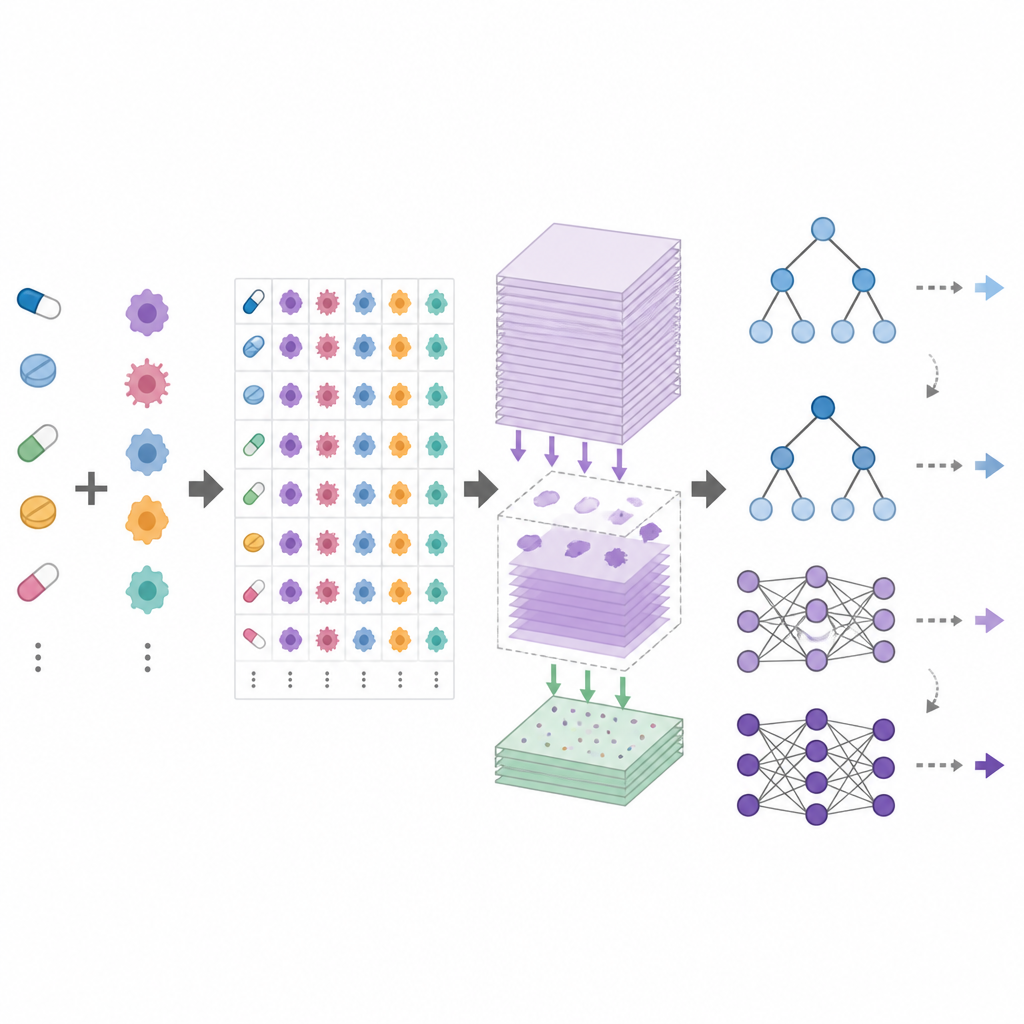
What the tests reveal about today’s models
When the authors ran a wide range of modern models through DrEval, they found that many barely beat the naive predictor that only uses mean drug and cell line behavior. In settings most relevant to patients, where models must handle cell lines they have never seen before, tuned tree-based ensembles such as random forests performed as well as or better than complex deep networks. All methods failed when asked to predict the effect of drugs that were not part of their training data, and performance dropped sharply when moving from one screening study to another or from cell lines to more realistic samples from patients. Careful “ablation” experiments showed that most of the useful signal comes from basic gene activity measurements, while additional data layers and sophisticated drug encodings often add little.
Why this matters for future cancer treatment
The main message of the study is that reliable prediction of cancer drug response remains an unsolved problem. Many earlier success stories were inflated by biased evaluation rather than true predictive power. By making these issues visible and providing a shared, reproducible testbed, DrEval helps the field focus on methods that genuinely learn how cancers respond to treatment instead of exploiting shortcuts in the data. For patients, this means that computer models are not yet ready to guide routine drug choices, but the path toward trustworthy tools is clearer: better data, stricter testing, and fair comparisons between simple and complex models.
Citation: Bernett, J., Iversen, P., Picciani, M. et al. Critical evaluation of drug response prediction models with DrEval. Nat Commun 17, 4238 (2026). https://doi.org/10.1038/s41467-026-72903-w
Keywords: drug response prediction, cancer cell lines, machine learning, benchmarking, model evaluation