Clear Sky Science · en
High-speed multiplexed DNA-PAINT imaging of nuclear organization using an expanded sequence repertoire
Seeing the Cell Nucleus in New Detail
Inside each of our cells, the genome is folded and arranged in an intricate way that helps control which genes are turned on or off. Until recently, much of this inner architecture was too small and crowded to see clearly. This paper describes a faster, more powerful imaging method that lets researchers map many different nuclear components at once with near-molecular precision, revealing how the genome and its surrounding structures reorganize when gene activity is shut down.
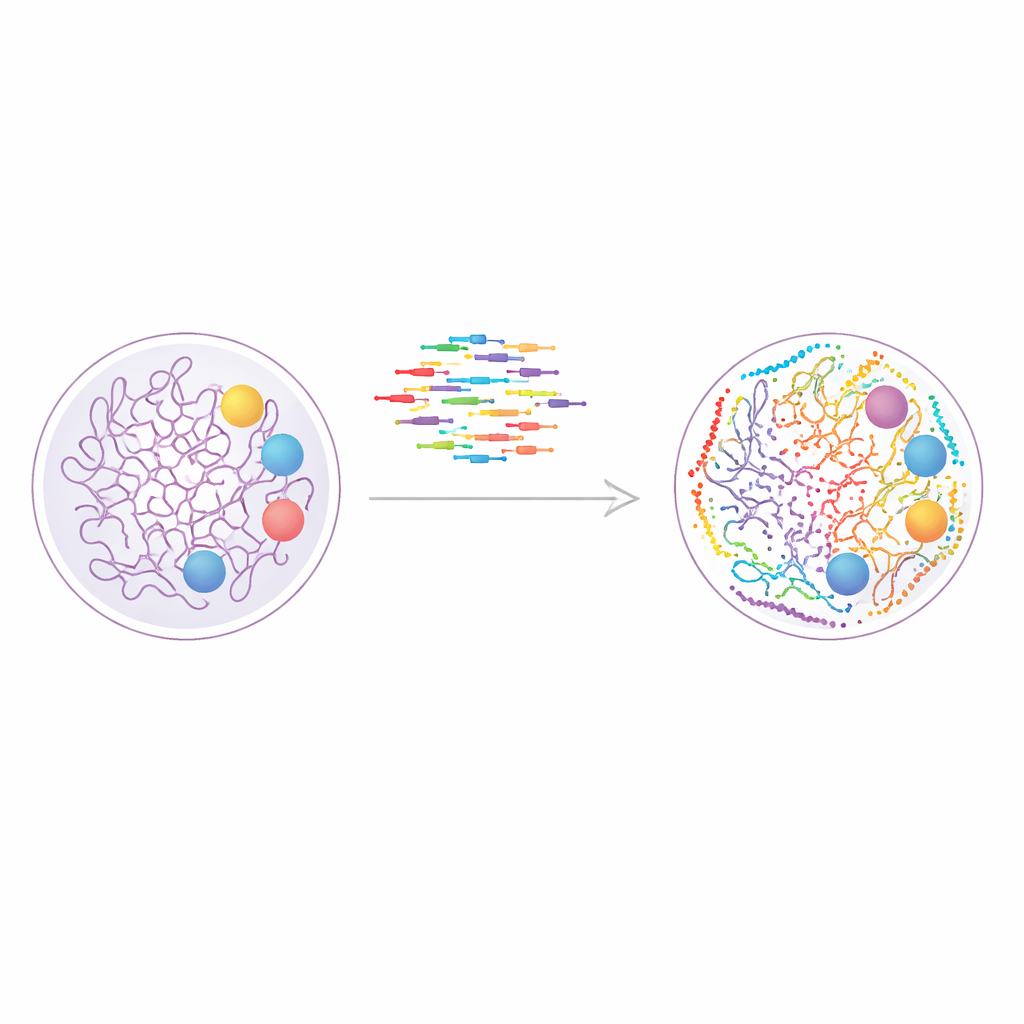
A New Way to Paint the Genome
The study builds on a technique called DNA-PAINT, which uses short fluorescent DNA strands that repeatedly bind and unbind to matching DNA tags on cellular structures. Each binding event produces a tiny burst of light, and many such events can be combined to reconstruct a sharp picture far beyond the limits of ordinary microscopes. DNA-PAINT is naturally well suited for looking at many targets in the same cell, because each protein or chromatin mark can be tagged with a distinct DNA sequence. The problem has been speed: traditional sequence designs bind too slowly, turning each imaging round into an hours-long experiment.
Expanding the Color Palette
To tackle this bottleneck, the authors systematically designed a broader set of “speed-optimized” DNA sequences that bind more frequently while still staying specific. They built designer DNA nanostructures, known as DNA origami, to test how well each imager strand recognized its matching docking strand and how often it flashed. By assembling origami tiles that carried docking sites arranged in a precise 4-by-3 grid just 20 nanometers apart, they could measure both resolution and unwanted cross-talk. This work yielded twelve fast sequences in total, of which ten could be used together without interfering, enabling direct imaging of up to ten targets in one experiment.
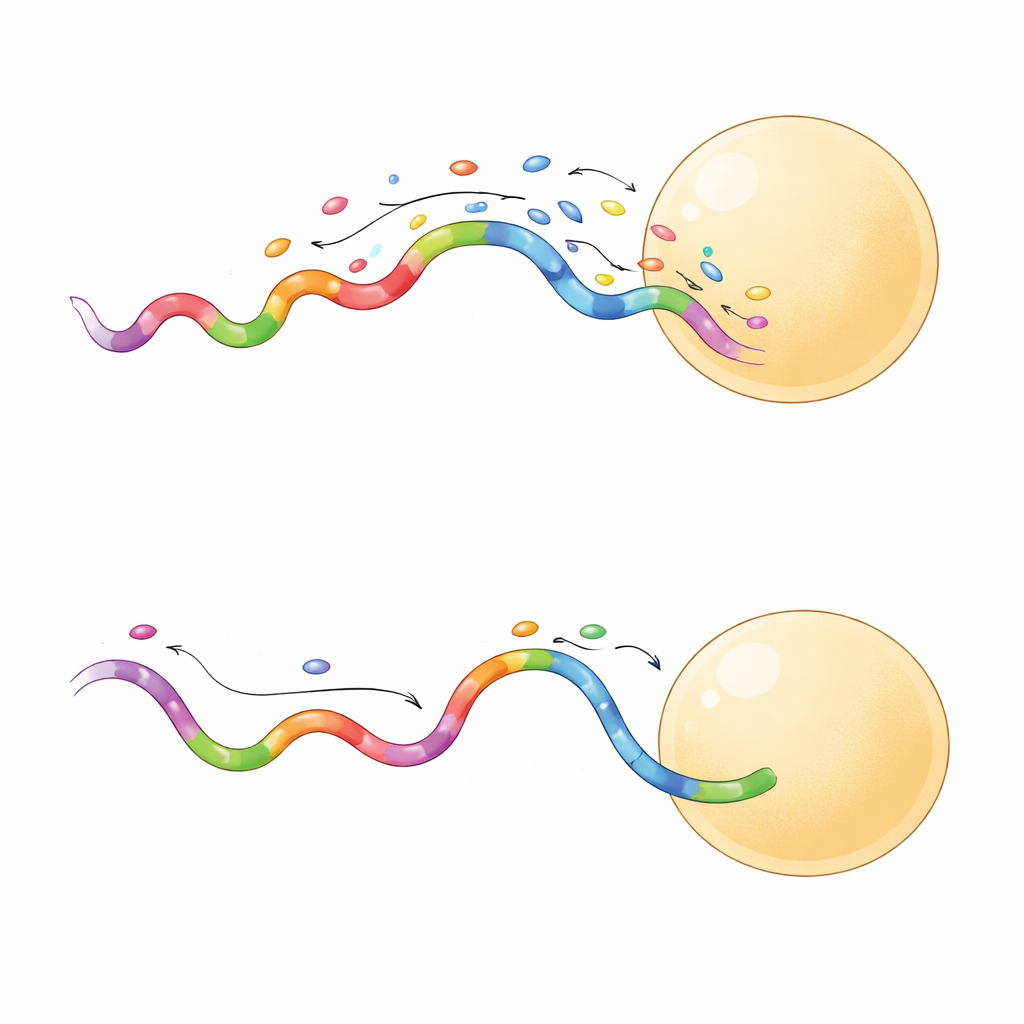
Sharper Images in Less Time
Not all fast sequences behaved the same way. A subset produced longer-lasting binding events, which turned out to be especially useful. When the team compared a short-binding and a long-binding sequence on the same origami grid, the long-binding probe produced crisper separation between neighboring sites and did so even at lower laser power. Longer events allowed the camera to collect more photons from each binding, boosting localization precision to just a few nanometers. These probes also sampled target sites more efficiently, reaching reliable coverage in fewer frames. In practice, this means researchers can acquire high-quality super-resolved images in minutes instead of hours, with less risk of damaging the sample.
Mapping the Nuclear Landscape
Armed with their expanded sequence set, the researchers moved from artificial DNA structures to living cell biology. They used DNA-PAINT to image nine different nuclear features in human cells, including active and paused forms of RNA polymerase II, several chemical tags on histone proteins that mark active or silent chromatin, the nuclear lamina at the periphery, and nuclear speckles—droplet-like hubs rich in splicing factors. By coupling conventional antibodies to DNA docking strands and imaging the targets in nine sequential rounds, they assembled a super-resolved, multiplexed map of the nucleus, where each dot corresponds to a molecular neighborhood only a few nanometers across.
Watching Chromatin Respond to Stress
The team then asked how this nuclear map changes when transcription—the copying of DNA into RNA—is blocked. Treating cells with the drug actinomycin D, which stalls RNA polymerases on DNA, dramatically reshaped the spatial relationships inside the nucleus. Nuclear speckles, which normally have irregular outlines and sit amid active chromatin, became more rounded, and their close contacts with transcription sites and active histone marks largely vanished. At the same time, repressive chromatin marks showed strengthened association with the nuclear lamina, consistent with a more compact, silenced genome. By combining several layers of statistical analysis, the authors could distinguish global trends from local pockets of enrichment or exclusion between different nuclear components.
Why This Matters for Biology
In essence, this work turns DNA-PAINT into a faster, more versatile tool for charting the cell’s inner geography. With a larger library of speed-optimized DNA sequences, scientists can now track many nuclear landmarks in the same cell at nanometer resolution and on practical time scales. The authors show that this approach is sensitive enough to capture how chromatin detaches from nuclear speckles and consolidates at the periphery when transcription is halted. As the method is extended to more proteins and modifications, it promises detailed “atlases” of nuclear organization in health and disease, helping to reveal how gene regulation is wired in three dimensions.
Citation: Banerjee, A., Anand, M., Srivastava, M. et al. High-speed multiplexed DNA-PAINT imaging of nuclear organization using an expanded sequence repertoire. Nat Commun 17, 3655 (2026). https://doi.org/10.1038/s41467-026-72206-0
Keywords: super-resolution imaging, DNA-PAINT, chromatin organization, nuclear speckles, transcription inhibition