Clear Sky Science · en
A high-affinity split-HaloTag for live-cell protein labeling
Seeing the hidden machinery inside living cells
Modern biology depends on being able to watch individual proteins at work inside living cells, but attaching bright molecular “flashlights” to those proteins without disturbing them is surprisingly hard. This study introduces a new labeling system that tucks a tiny tag into a protein and then lights it up with powerful chemical dyes, allowing researchers to follow even scarce, fragile targets in real time with cutting-edge microscopes.
A tiny tag meets a powerful light
The work builds on HaloTag, a widely used protein label that forms a permanent bond to special fluorescent dyes. HaloTag is bright and versatile but it is also bulky, which can interfere with the normal behavior of the protein it is fused to. The authors solve this by splitting HaloTag into two pieces: a very small peptide called Hpep, only 14 building blocks long, and a larger partner protein called cpHaloΔ3. The small Hpep can be wired directly into the protein of interest, while cpHaloΔ3 is supplied separately. When they meet inside a cell, the two pieces snap together with very high affinity and recreate an active HaloTag that can grab bright dyes.
Designing a snap-together light switch
To make this split system practical, the team had to balance several properties at once. Hpep and cpHaloΔ3 needed to recognize each other extremely tightly so that even a single copy on a low-abundance protein would be labeled, but cpHaloΔ3 on its own should remain almost inactive to avoid unwanted background glow. Using yeast display and high-throughput cell sorting, they screened libraries of slightly altered cpHalo fragments for variants that were quiet when alone but fast and efficient once bound to Hpep. The final version, cpHaloΔ3, combined nanomolar binding strength, low unintended labeling, and rapid reaction with dyes once the two pieces were reunited. Variants of Hpep were also tuned so that, when bound, they subtly changed how attached dyes behaved, altering both brightness and fluorescence lifetime.
Lighting up real cellular structures
The researchers then showed that their split-HaloTag works across a wide range of cellular settings. They attached Hpep to proteins that mark the nucleus, the mitochondrial surface, and the nuclear envelope, and co-expressed cpHaloΔ3 in human cells. After adding cell-permeable HaloTag dyes, the reconstituted tag produced sharp images of the expected structures with very little background from unbound cpHaloΔ3. Crucially, because Hpep is so small, it could be inserted directly into the cell’s own genes using CRISPR without the need for cloning large DNA constructs. The team demonstrated seamless “knock-ins” for many different endogenous proteins, from structural components like vimentin to membrane channels and clathrin, and showed that these could be enriched and analyzed by flow cytometry and microscopy.
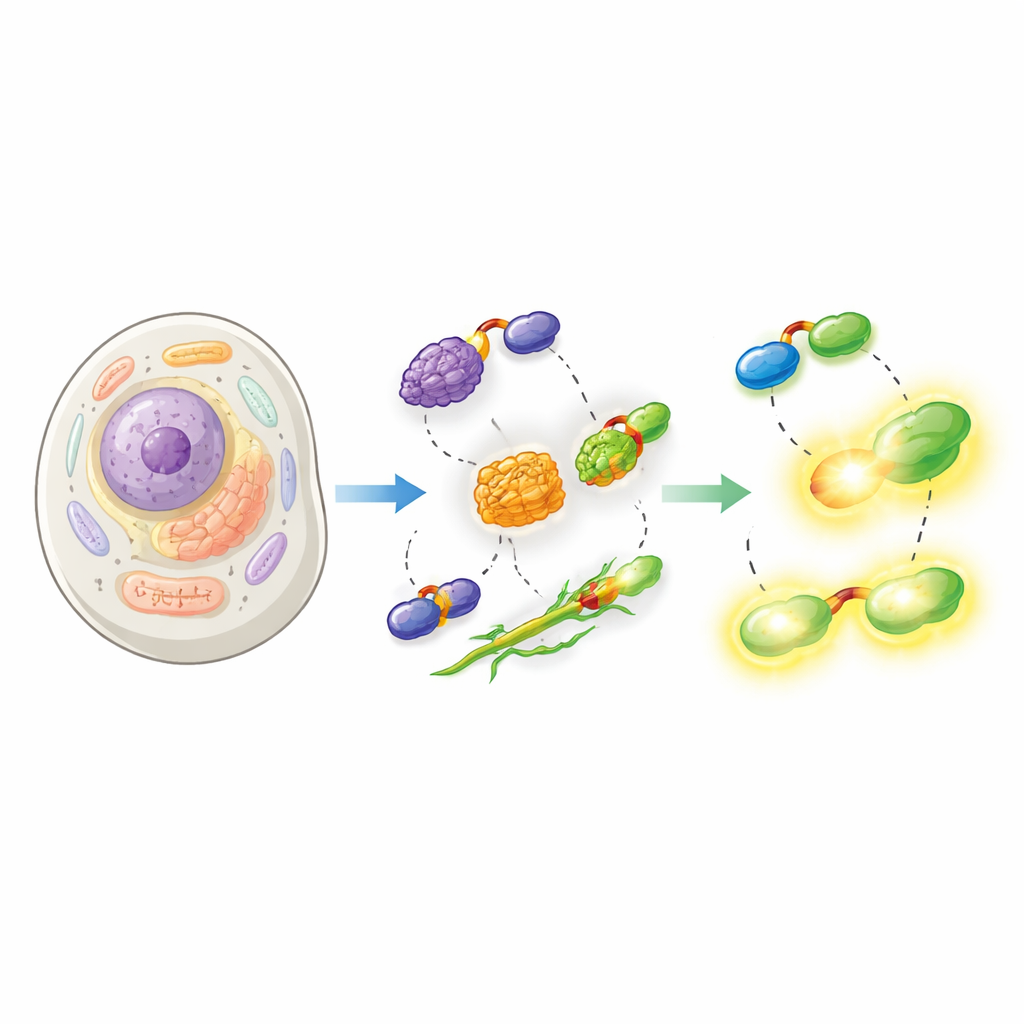
Pushing the limits of modern microscopy
Beyond standard fluorescence imaging, the new tag was tested with some of the most demanding optical techniques. Using special dyes compatible with super-resolution and depletion microscopy, the authors resolved mitochondrial membranes and microtubules with details well below the conventional diffraction limit, in some cases in living cells. They also combined the system with expansion microscopy, physically swelling preserved cells to reveal fine structure, and with fluorescence lifetime imaging. Different Hpep variants changed how long the dye stayed in its excited state, allowing two proteins labeled with the same color to be distinguished purely by their lifetimes and enabling multiplexed imaging with a single cpHaloΔ3 partner.
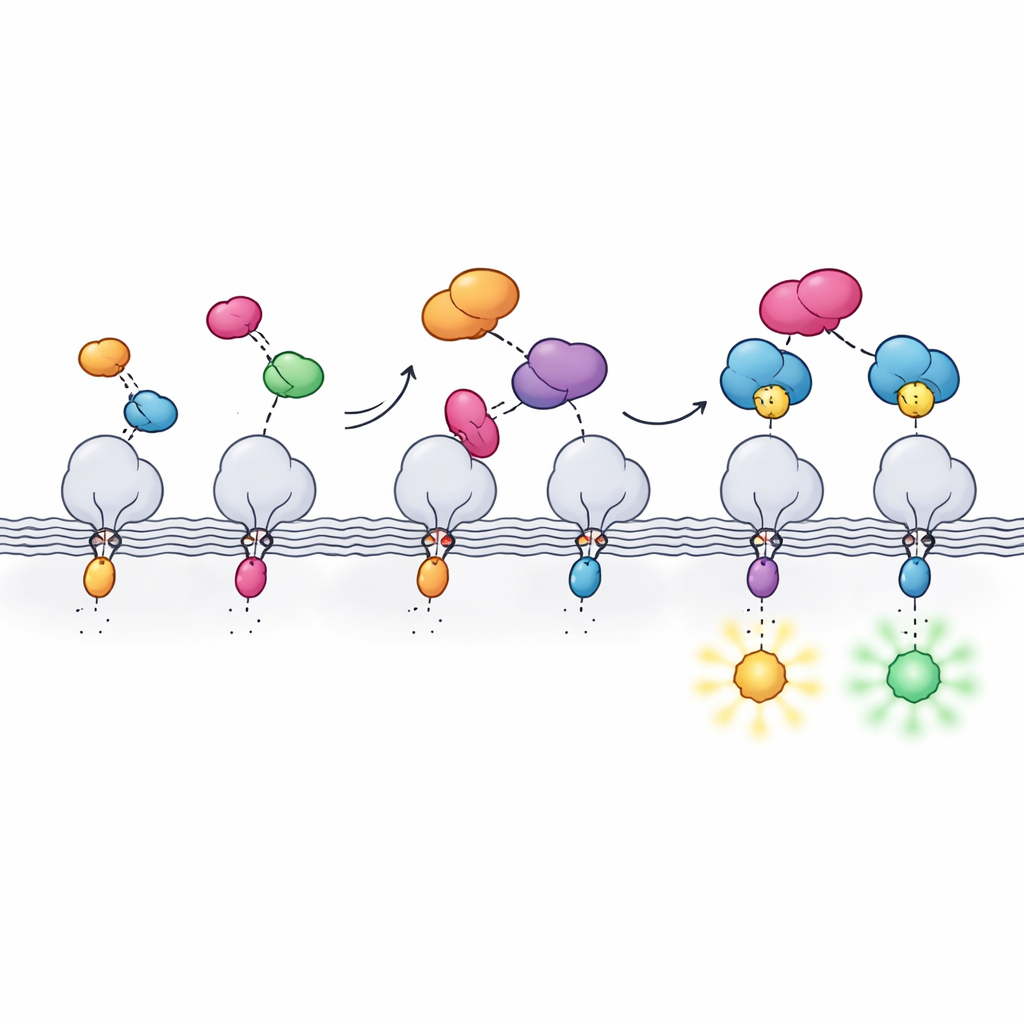
Why this new tag matters
Together, these advances create a compact, flexible labeling system that merges the genetic convenience of tiny peptide tags with the optical power of synthetic dyes. Hpep’s small size makes it easy to insert into native genes and less likely to disrupt protein function, while cpHaloΔ3 brings in bright, stable colors when and where they are needed. The result is a high-contrast, low-background way to watch proteins inside living cells, compatible with advanced microscopes and multi-target experiments. This split-HaloTag toolkit should help researchers chart how cellular components are organized and how they change over time in health and disease.
Citation: Lin, YH., Kompa, J., Sun, De. et al. A high-affinity split-HaloTag for live-cell protein labeling. Nat Commun 17, 2865 (2026). https://doi.org/10.1038/s41467-026-71032-8
Keywords: protein labeling, live-cell imaging, HaloTag, CRISPR tagging, super-resolution microscopy