Clear Sky Science · en
Mitotic microhomology-mediated break-induced replication promotes chromoanasynthesis
When Chromosomes Go Haywire
Cancer and some birth defects often arise when our chromosomes are scrambled in extreme ways: stretches of DNA are duplicated, flipped, and stitched together into bizarre patchworks. This paper asks a basic question with big medical implications: how does such chaos happen in a single leap, rather than bit by bit over many years? By peering closely at damaged chromosome ends, the authors uncover a highly error‑prone repair process that can rewrite long stretches of the genome in one go.
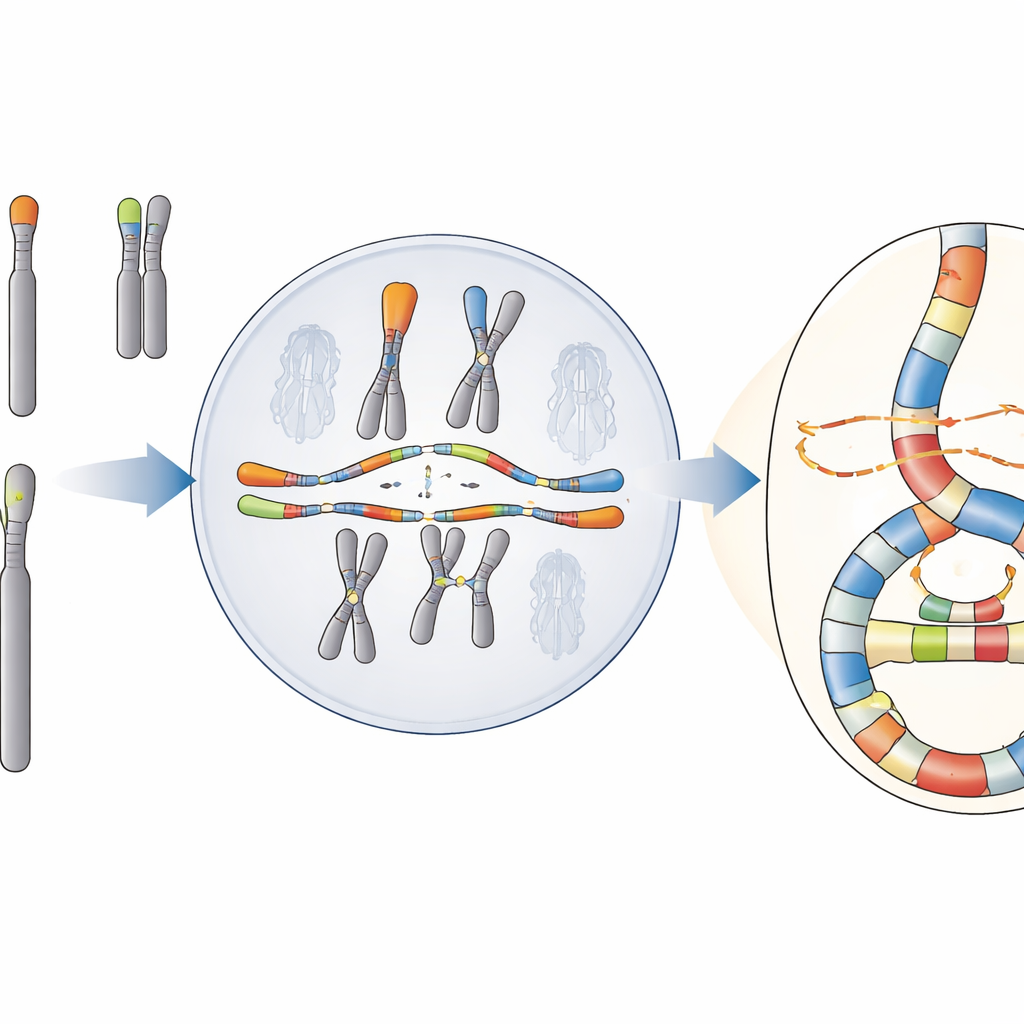
A Catastrophe with a Hidden Pattern
The team focuses on a phenomenon called chromoanasynthesis, in which pieces of DNA are copied and reinserted multiple times, creating dense clusters of rearrangements and extra copies of genes. Unlike another well‑known catastrophe, chromothripsis, which looks like random re‑gluing of shattered DNA, chromoanasynthesis carries the fingerprint of repeated DNA copying and switching between templates. These events are common in cancers and in some rare developmental disorders, but their origins in human cells have been hard to pin down because standard sequencing often misses the full complexity of the reshuffled regions.
Reading Ultra‑Long DNA Molecules
To tackle this, the authors built a single‑molecule long‑read sequencing method, Fusion‑seq Long‑Read (FSLR), aimed specifically at chromosome ends, or telomeres, that have fused together. They studied human fibroblasts pushed into “telomere crisis,” a state where chromosome ends become dangerously short, and also used engineered molecular scissors to cut DNA just inside telomeres in cancer cell lines. Long‑read sequencing allowed them to follow each fused DNA molecule from end to end, revealing not only where the two telomeres met but also every extra fragment wedged in between and where in the genome those fragments came from.
A Mutagenic Copy‑and‑Paste Repair Pathway
The fused molecules they uncovered were astonishingly complex. Some contained dozens or even more than a hundred inserted fragments drawn from many chromosomes, with frequent “hotspots” that had been copied repeatedly. The junctions between fragments often shared a few matching bases, a hallmark of “microhomology”‑based pairing rather than clean, textbook repair. The patterns strongly pointed to a copy‑and‑paste mechanism known as microhomology‑mediated break‑induced replication (MM‑BIR). In this process, a broken DNA end briefly pairs with a short matching sequence on another DNA molecule, starts copying, then lets go and latches onto yet another template, leaving behind a trail of duplicated and rearranged segments. Genetic and drug‑based experiments showed that this pathway depends on a collaboration between enzymes normally associated with small‑patch end joining and a more powerful replication machine that can copy long stretches of DNA.
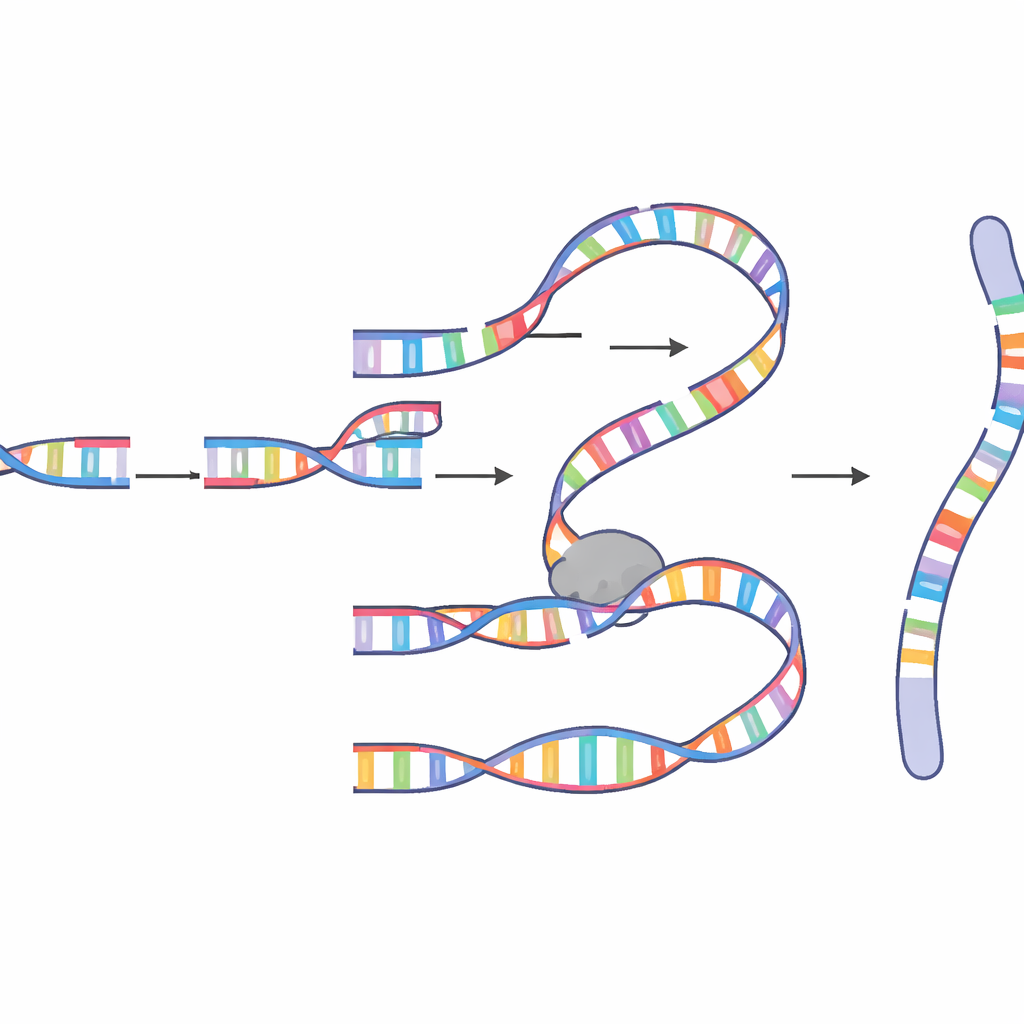
Why Mitosis Is a Dangerous Moment
A key surprise was timing. Chromoanasynthesis in this system arose specifically during mitosis, the brief window when cells divide and chromosomes are highly condensed. The authors showed that when the cell’s G2/M damage checkpoint—its main safeguard that blocks entry into mitosis with broken DNA—was weakened, complex events became much more frequent. They also demonstrated that an enzyme called polymerase theta, known as a “last‑ditch” repair factor, plays a central role, helping broken ends find tiny matching sequences and initiate synthesis. More processive enzymes such as polymerase delta then extend the new DNA, aided by factors like PIF1, POLD3, and PCNA that keep the replication machinery moving. When these helpers were altered, the length and pattern of the inserted DNA changed, confirming that chromoanasynthesis is driven by a specialized, mitotic version of MM‑BIR rather than by simple ligation of broken fragments.
New Clues to Cancer and Birth Defects
Altogether, the work reveals a highly mutagenic “emergency repair” pathway that kicks in when broken chromosome ends encounter mitosis. Instead of gently restoring the original sequence, mitotic MM‑BIR can rapidly amplify and remix genomic regions, including gene‑rich areas and potential oncogenes, in a single event. For a healthy cell this is a risky but sometimes unavoidable way to finish replication and division. For a precancerous cell, however, it may provide exactly the kind of genomic upheaval that drives aggressive tumor growth or contributes to complex chromosomal changes seen in congenital disorders. By mapping this process in molecular detail, the study points to new ways of understanding—and perhaps one day targeting—the catastrophic genome reshuffling that underlies some of the most difficult human diseases.
Citation: Ngo, G.H.P., Cleal, K., Seifan, S. et al. Mitotic microhomology-mediated break-induced replication promotes chromoanasynthesis. Nat Commun 17, 3375 (2026). https://doi.org/10.1038/s41467-026-70086-y
Keywords: chromosomal rearrangements, telomere crisis, DNA repair, genome instability, cancer genetics