Clear Sky Science · en
Multi-to-uni modal knowledge transfer pre-training for molecular representation learning
Why smarter drug computers matter
Designing new medicines increasingly depends on computers that can "understand" molecules. These programs turn a drug molecule into numbers a model can use to predict safety, potency, or harmful interactions. The paper introduces M2UMol, a new way to train such models so they work well even when only the most common kind of molecule data is available, promising faster and more reliable help for chemists and doctors.
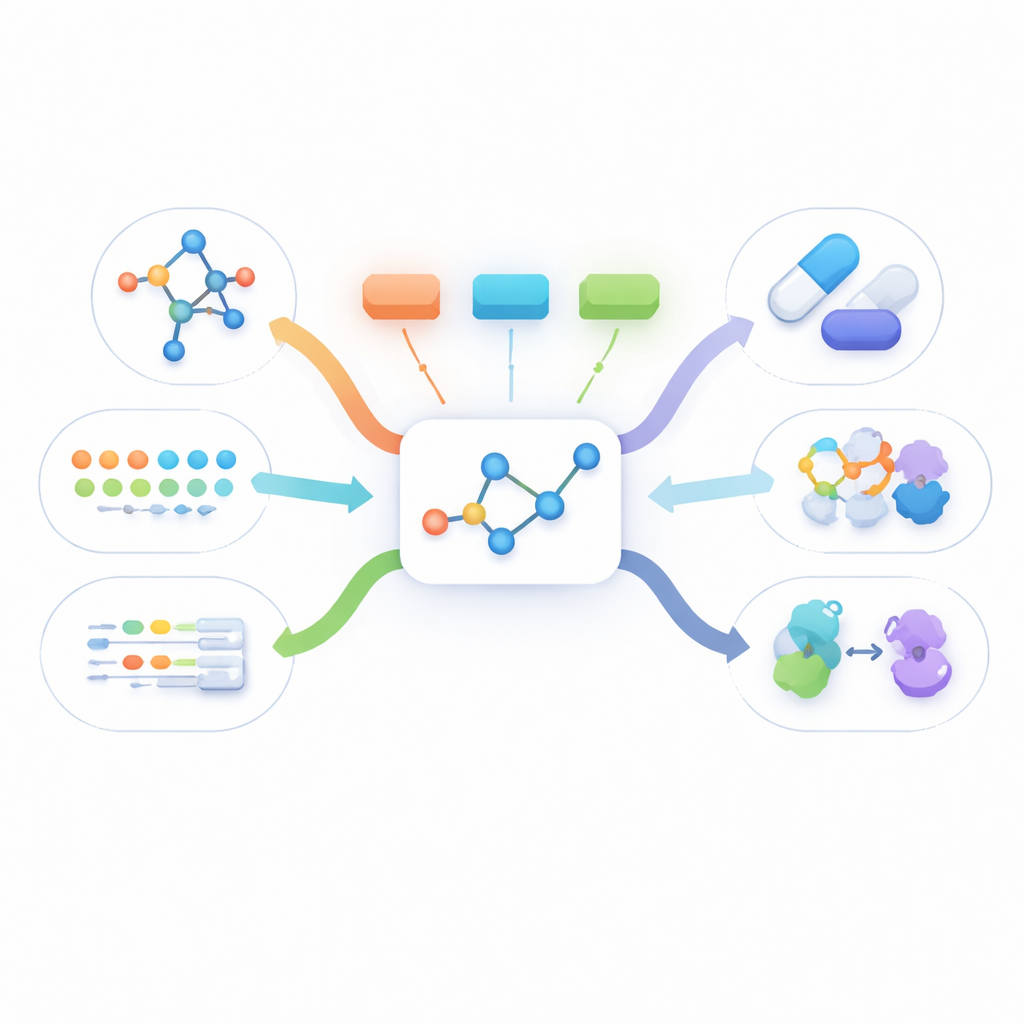
Seeing a molecule from many angles
Real drug information comes in many forms: a flat drawing of how atoms connect, a three-dimensional shape, written descriptions of how the drug works, and long lists of biological facts such as what proteins it binds to. Most existing methods either use just one of these views or require that all of them be present for every molecule during training. In practice, however, the only data that is almost always available is the simple two-dimensional structure. Other views are often missing, which prevents current multimodal methods from learning from large numbers of real-world molecules.
Teaching one view to think like many
M2UMol tackles this by treating the two-dimensional graph of a molecule as the hub and learning how that single view relates to the others. During pre-training, the system reads 2D graphs, 3D shapes, text summaries, and biological feature lists for more than eleven thousand drug-like molecules. It uses separate encoders for each type of data and then trains "adapters" that take only the 2D graph and generate what the 3D, text, or biological patterns should look like. A contrastive learning step pushes the generated patterns to closely match the real ones whenever they are available, even if some views are missing for some molecules. A second training task asks the system to guess which modality a generated pattern belongs to, helping it keep the different kinds of information distinct.
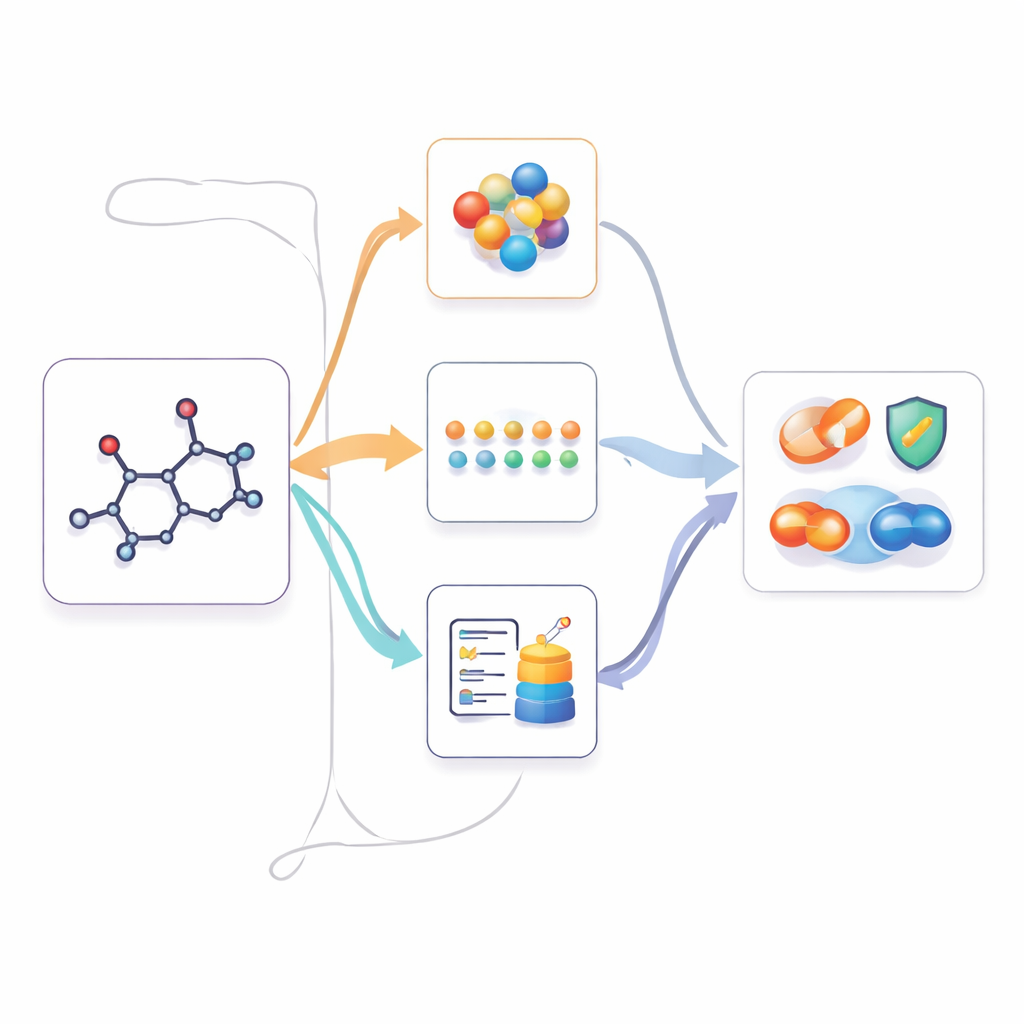
Using multimodal knowledge when only 2D is available
Once M2UMol has learned these relationships, it can be applied to practical tasks where only 2D structures are given, such as predicting toxicity, disease-related properties, or drug interactions. For each new molecule, the pre-trained 2D encoder produces a core representation, and the adapters produce three "imagined" versions: one resembling 3D geometry, one mimicking text-based knowledge, and one capturing biological context. A simple attention module then combines these four views into a final fingerprint used by task-specific predictors. This effectively lets the model act as if it had rich multimodal information, while users only need to supply a standard 2D structure.
Better predictions with less data and clearer reasoning
Across a wide panel of benchmarks, M2UMol outperforms both single-view and previous multimodal models on predicting molecular properties and on more complex tasks like drug–drug and drug–target interactions. It does so even though it is pre-trained on just over eleven thousand molecules, using modest computing resources, while many competing methods use millions. Visual analyses show that the learned representations are both well separated between different classes and evenly spread out, a sign of high information content. The model can also highlight specific atoms and bonds that drive its decisions, and these often match known functional groups responsible for toxicity, activity at a receptor, or problematic interactions between drugs.
What this means for future medicines
For non-specialists, the key idea is that M2UMol learns to think about a simple line drawing of a molecule as if it carried three extra layers of rich scientific context. That trick allows it to make stronger, more explainable predictions about how a drug will behave, even when detailed experimental data are unavailable. Because it is efficient, open-source, and packaged for easy use, this approach could help researchers screen compounds more wisely, understand why some drugs fail, and eventually guide the design of new medicines with fewer surprises.
Citation: Xiong, Z., Wang, Z., Huang, F. et al. Multi-to-uni modal knowledge transfer pre-training for molecular representation learning. Nat Commun 17, 3797 (2026). https://doi.org/10.1038/s41467-026-69302-6
Keywords: drug discovery, molecular representation learning, multimodal AI, drug interactions, computational chemistry