Clear Sky Science · en
A light-weight, data-driven segmentation method for multi-state Brownian trajectories
Watching Molecules Wander
Inside living cells, countless molecules are constantly jostling, binding, and unbinding, and their motion holds clues to how life works. Modern microscopes can follow single molecules as they move, but turning these tangled paths into clear stories about how fast a molecule diffuses or when it binds to a partner is surprisingly hard. This paper introduces a simple, fast way to break such motion into distinct “states” of movement, making it easier for biologists to read what single-molecule movies are trying to tell us.
Why Tracking Molecules Is Tricky
Single-particle tracking techniques record the positions of individual molecules over time, building up a trajectory that reflects how they explore their surroundings. In many real systems, a molecule does not move in just one way: it might roam freely for a while and then slow down when it binds to a receptor or becomes trapped in a cluster. That means the trajectory is a mixture of several modes of motion, each with its own characteristic speed. Existing analysis tools can, in principle, separate these modes, but widely used methods based on hidden Markov models or deep learning are often computationally heavy, require expert tuning, or act as “black boxes” whose decisions are difficult to interpret.
A Lean Alternative to Heavyweight Algorithms
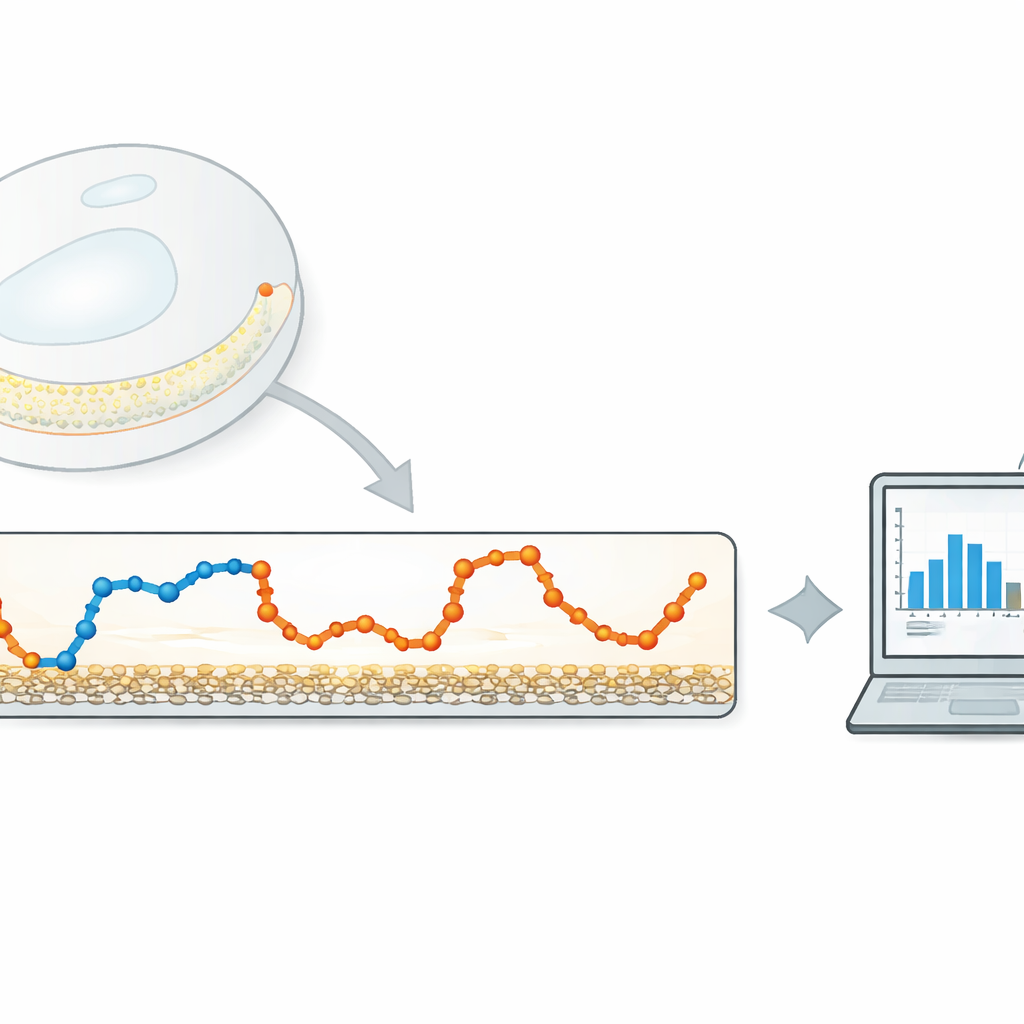
The authors propose a light-weight, data-driven method that focuses on a very simple quantity: how far the particle moves between consecutive camera frames. First, they compute this step-by-step displacement time series from a tracked trajectory. They then apply a one‑dimensional Gaussian filter—a type of sliding, smoothly weighted average—to this series. The filter width controls how strongly nearby steps are averaged together. If the filter is too narrow, little noise is removed; if it is too wide, the transitions between different movement states are washed out. The key idea is to automatically tune this width so that the filtered steps from each movement state overlap as little as possible.
Letting the Data Sort Itself
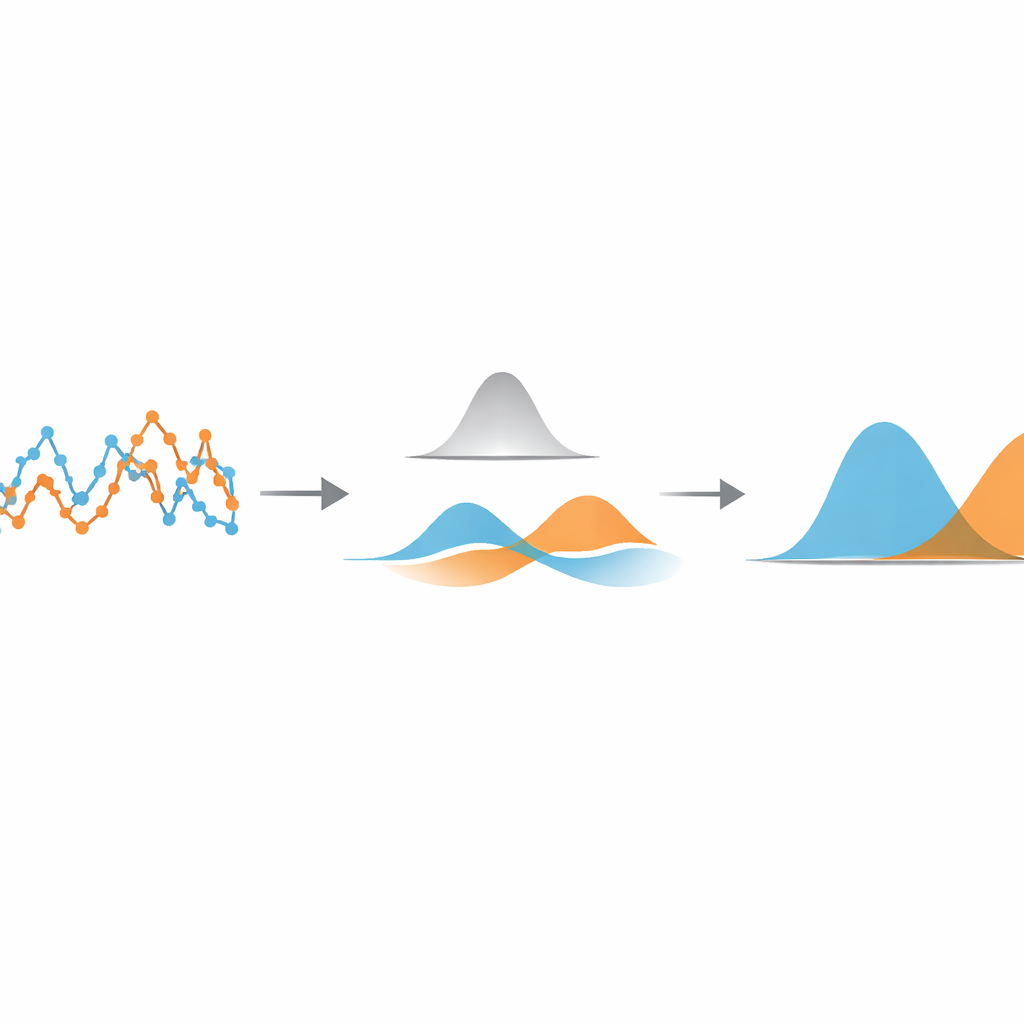
To find that sweet spot, the method treats the filtered displacements as coming from a mixture of simple bell-shaped curves. Using a standard statistical tool called a Gaussian mixture model, it fits two such curves to the filtered data and calculates how much they overlap. By scanning through different filter widths and picking the one that minimizes this overlap, the algorithm makes the different movement states as distinct as the data allow. Once this optimal setting is found, every displacement is assigned to the state whose bell curve it most likely belongs to, effectively segmenting the original trajectory into fast and slow stretches. Importantly, this segmentation happens before any attempt to estimate physical parameters like diffusion coefficients or state lifetimes, so those can later be measured with familiar, well-tested formulas.
Testing with Simulations and Real Molecules
The researchers stress‑tested their method on computer-generated trajectories where a particle switches between two diffusion speeds with known properties. They varied how different the two speeds were, how long the particle stayed in each state, how noisy the position measurements were, and how much blur the camera introduced. Over a wide range of realistic conditions—whenever the fast and slow states differed by at least about a factor of four and each state lasted more than several camera frames—the algorithm correctly labeled more than 90 percent of the time points. Crucially, the method also proved robust when applied to experimental data from fluorescently labeled proteins diffusing in a supported membrane, where it revealed two clearly separated populations: one mobile and one nearly immobile.
From Clean Segments to Biological Insight
Once the trajectories were segmented, the team showed that diffusion coefficients could be recovered with good accuracy, even in the presence of substantial measurement noise or motion blur. Estimating how long molecules typically remained in each state was more demanding and required longer trajectories, but still yielded reasonable lifetimes under suitable conditions. The overall message is that a relatively simple, transparent procedure—filtering steps in time and fitting a pair of bell curves—can rival more complex approaches while running quickly on ordinary computers. For experimentalists, this means they can process single‑particle data on the fly, tune their imaging conditions in real time, and obtain clearer pictures of how molecules bind, unbind, and move in crowded cellular environments.
Citation: El Korde, I., Lewis, J.M., Clarkson, E. et al. A light-weight, data-driven segmentation method for multi-state Brownian trajectories. npj Biol. Phys. Mech. 3, 6 (2026). https://doi.org/10.1038/s44341-026-00037-7
Keywords: single-particle tracking, molecular diffusion, trajectory segmentation, Brownian motion, membrane proteins