Clear Sky Science · en
Bioimage analysis for multiplexed FUCCI acquisitions powered by deep learning
Watching Cells Live Their Daily Lives
Every living cell constantly cycles through stages of growth, DNA copying, and division. Biologists now have fluorescent tools that make these cell-cycle stages glow in different colors, letting them watch this process unfold in real time. But when too many signals are recorded at once, the images become noisy and hard to interpret—even for computers. This paper introduces a deep-learning–based method that cleans up these complex movies, tracks individual cells, and reads out where each one is in its cycle, offering new power for cancer research, stem cell studies, and drug testing.
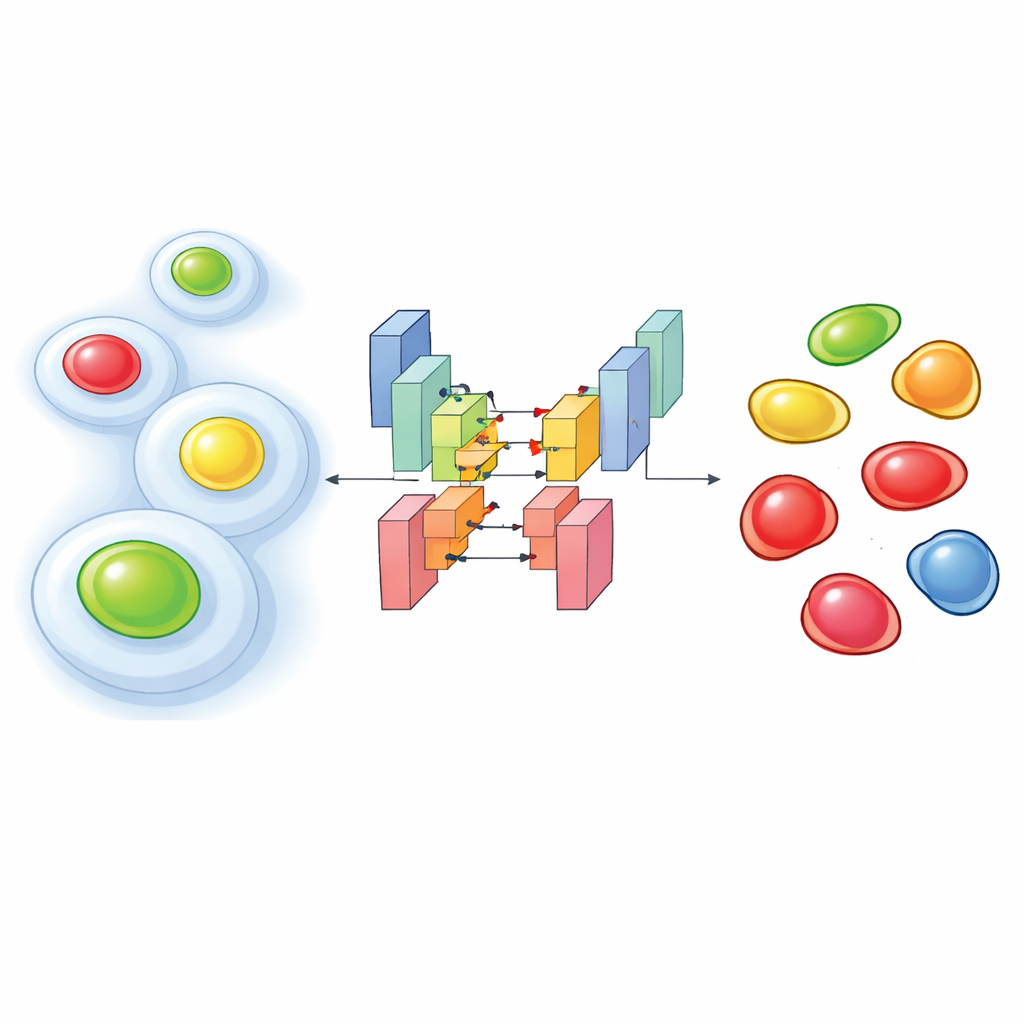
Glowing Timers Inside Living Cells
To see how cells progress through their life cycle, researchers often use a molecular tool called FUCCI, which makes the cell’s nucleus glow in one color during early stages and another color later on, with combinations in between. Newer experiments go a step further: they record FUCCI together with additional fluorescent markers that highlight the cell’s inner scaffolding or other structures. This “multiplexing” helps connect what a cell is doing structurally with where it is in its life cycle. The downside is that to avoid harming living cells, scientists must illuminate them gently, which produces dim, noisy images where colors bleed into each other. Under these conditions, standard image-analysis software often fails to correctly find each nucleus or assign the right cell-cycle stage.
Teaching an Algorithm to See Through the Noise
The authors tackled this problem by training custom deep-learning networks on thousands of carefully annotated images from two human cell lines. Their approach feeds multiple channels into a segmentation model based on the StarDist architecture, which is particularly good at outlining individual nuclei. They tested three input setups: one using only the cytoplasmic marker (a fluorescently tagged structural protein), one using just the two FUCCI colors, and one combining all three. Despite the noisy conditions, the two- and three-channel versions reached very high accuracy in finding nuclei, clearly outperforming popular pre-trained tools when the signal was weak. Surprisingly, even the model using only the cytoplasmic marker could often infer where the nuclei were, because the surrounding scaffold leaves a characteristic “hole” where the nucleus sits.
From Colored Dots to Cell-Cycle Labels
Finding nuclei is only half the battle; researchers also need to know which stage of the cell cycle each nucleus is in. Traditionally, this is done by setting intensity thresholds on the FUCCI colors, a simple method that can be easily fooled by bleed-through from other channels. The team extended their network so that it not only outlines each nucleus but also labels it as belonging to one of several cell-cycle phases based on the color patterns. When they compared this approach to the classic intensity-threshold method, the deep-learning classifier was more accurate, especially in noisy data. It even generalized to a different version of the FUCCI system from another lab, after the authors simply remapped which color combinations corresponded to which phases—no retraining required.
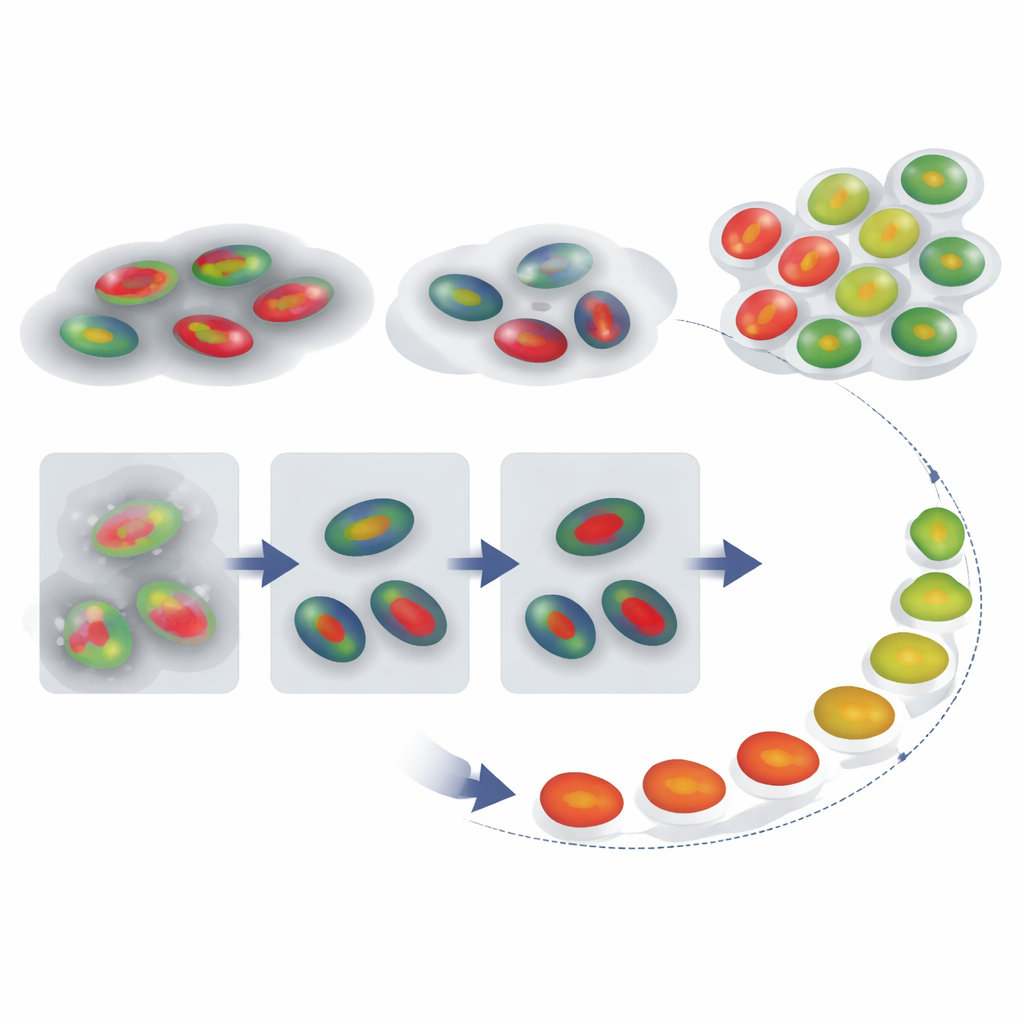
Reconstructing a Cell’s Journey Through Time
With reliable nuclear outlines and phase labels in hand, the authors next asked whether they could reconstruct how individual cells move through the cycle over time, even when a recording does not capture a full division-to-division sequence. They tracked FUCCI signals from single cells and compared each intensity trace to a reference pattern for a typical cycle using a technique called dynamic time warping, which flexibly stretches and compresses the time axis to find the best match. This gave each cell a “cell-cycle percentage” that acts like a pseudotime—an estimate of how far along it is. Just as important, the amount of time stretching needed serves as a red flag: cells whose traces must be heavily distorted to match the reference are likely behaving abnormally, such as cells stuck for a long time in one phase.
Why This Matters for Health and Disease
By combining multiplexed fluorescent imaging with tailored deep-learning tools, this work delivers an end-to-end pipeline: it segments dim, noisy images, labels each nucleus with a cell-cycle phase, tracks cells over time, and flags those that deviate from a standard pattern. For non-experts, the key message is that researchers can now follow thousands of living cells more reliably and with less manual effort, even under gentle imaging conditions that preserve cell health. This opens the door to more precise studies of how cancer drugs halt cell division, how stem cells decide their fate, and how mechanical forces in tissues influence when cells choose to divide or pause.
Citation: Zimmermann, J., Pezzotti, M., Torchia, E. et al. Bioimage analysis for multiplexed FUCCI acquisitions powered by deep learning. npj Imaging 4, 27 (2026). https://doi.org/10.1038/s44303-026-00159-6
Keywords: cell cycle imaging, deep learning microscopy, FUCCI sensor, cell tracking, bioimage analysis