Clear Sky Science · en
Chromatin accessibility landscapes define stromal cell identities across tissues
Why the hidden code of our tissues matters
Every organ in the body knows what to do because its cells read the genome in slightly different ways. This study asks a simple but powerful question: can we tell what kind of cell we are looking at—and which organ it came from—just by seeing which parts of its DNA are open and ready to be read? The authors build a detailed map of this “open DNA” landscape across many mouse organs and show that it quietly records both cell identity and tissue of origin, with potential uses in diagnosis, cancer tracing, and regenerative medicine.
Peeking into open DNA across the body
To explore this hidden code, the researchers used a technique called single-cell ATAC-seq, which marks the stretches of DNA that are physically accessible inside each nucleus. Working with frozen samples from nine mouse organs, including brain, heart, lung, liver, kidney, intestine, pancreas, colon, and spleen, they isolated the nuclei and profiled more than 51,000 individual cells. Careful fine-tuning was needed for each tissue so that fragile nuclei from soft organs like lung and brain, as well as dense tissues like heart, could all be captured with high quality. After sequencing, advanced computational tools grouped cells by how similar their accessible DNA regions were, then projected these groups into two-dimensional “maps” where related cells cluster together. 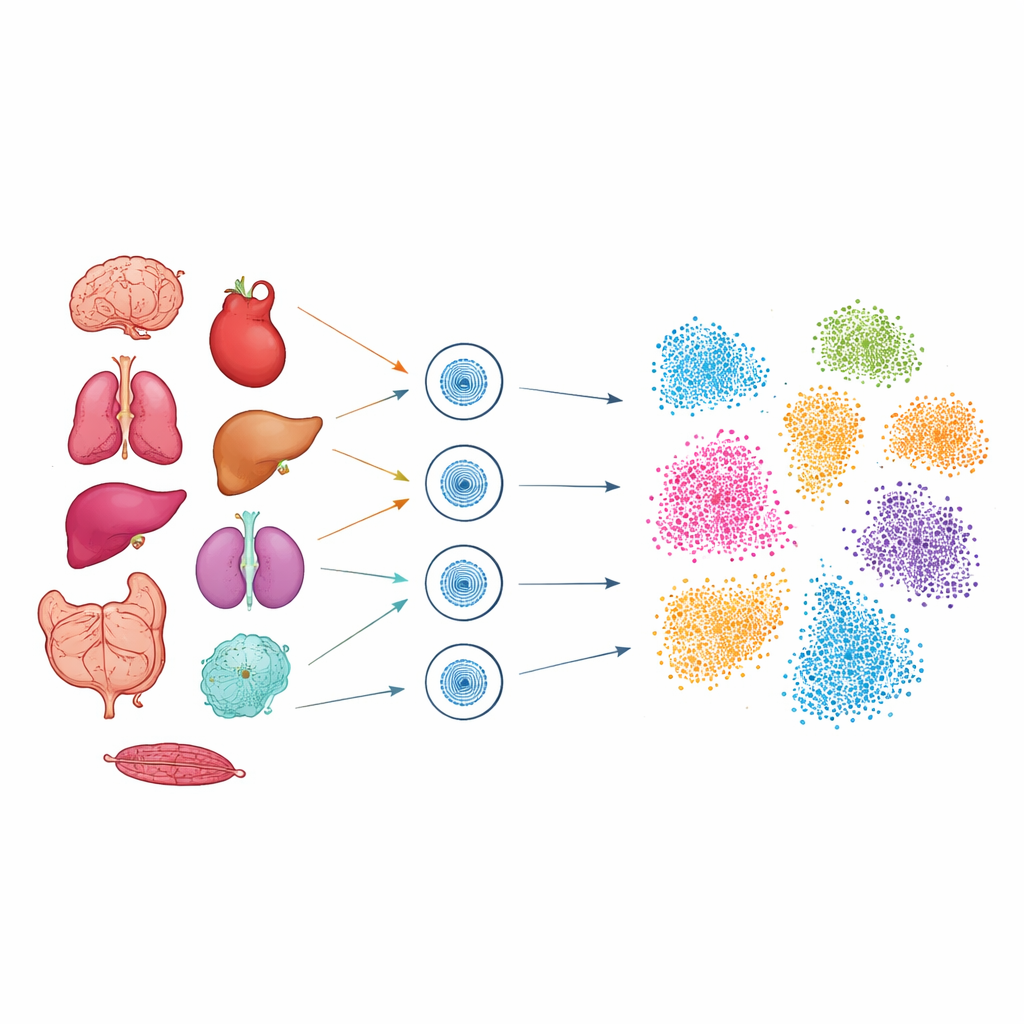
Recognizing cell types from their open DNA
From these maps, the team identified 28 major cell types spread across the nine organs, such as neurons, cardiomyocytes, immune cells, and various gut and liver cells. Even without measuring RNA directly, each cell type carried a characteristic pattern of open DNA peaks around genes known to be important for its function. For example, T cells showed open sites near genes involved in immune signaling, neurons near genes needed for synapses and brain activity, and heart muscle cells near genes tied to contraction and calcium handling. By comparing their results to large public reference atlases, the authors confirmed that the cell types they recovered matched well with previously described populations, but now with an added layer of regulatory detail: where in the genome the control switches for those cells lie.
Tissue fingerprints written in chromatin
Beyond distinct cell types, the study uncovered an extra layer of organization: cells from the same organ tended to share broader patterns of accessibility, even when they belonged to different types. This was especially striking in “stromal” cells—supporting cells such as endothelial cells, fibroblasts, and macrophages that exist in many organs. Although an endothelial cell in the heart and one in the liver share a core identity, their open DNA regions diverge in ways that reflect their local environment and jobs. In the liver, accessible regions near genes involved in detoxification and metabolism were prominent; in lung endothelium, open regions sat near genes tied to gas exchange and surfactant; in kidney, accessibility highlighted genes important for filtration and ion balance. Statistical comparisons confirmed that overall similarity in chromatin patterns was higher within organs than across them.
Zooming in on the control networks
The open DNA regions are not random—they serve as docking sites for transcription factors, the proteins that turn genes on or off. By searching for recurring sequence motifs within accessible regions, the authors could infer which transcription factor families are most active in different settings. Lung cells showed strong signatures of Nkx factors that help define airway identity, neurons were enriched for Rfx factors involved in brain development, and immune cells carried motifs for regulators such as Runx1. Footprinting analyses, which look at how these proteins shield DNA from the ATAC-seq enzyme, reinforced that these motifs correspond to real binding events. Together, the patterns of open chromatin and transcription factor activity outline cell type–specific regulatory programs that help lock in each cell’s role.
Tracing stromal cells back to their home organ
To test whether chromatin patterns alone could act as a “return address” for shared stromal cells, the researchers grouped similar single cells into compact units called metacells. Within macrophages and fibroblasts found across multiple organs, they discovered modules of accessible DNA that aligned closely with tissue of origin: spleen macrophages differed from liver macrophages, and heart fibroblasts from lung fibroblasts, even though all belonged to the same broad cell class. Endothelial cells showed the same principle. By combining a general endothelial marker region with a liver-specific accessible region, the team could cleanly pick out liver sinusoidal endothelial cells from the mix. 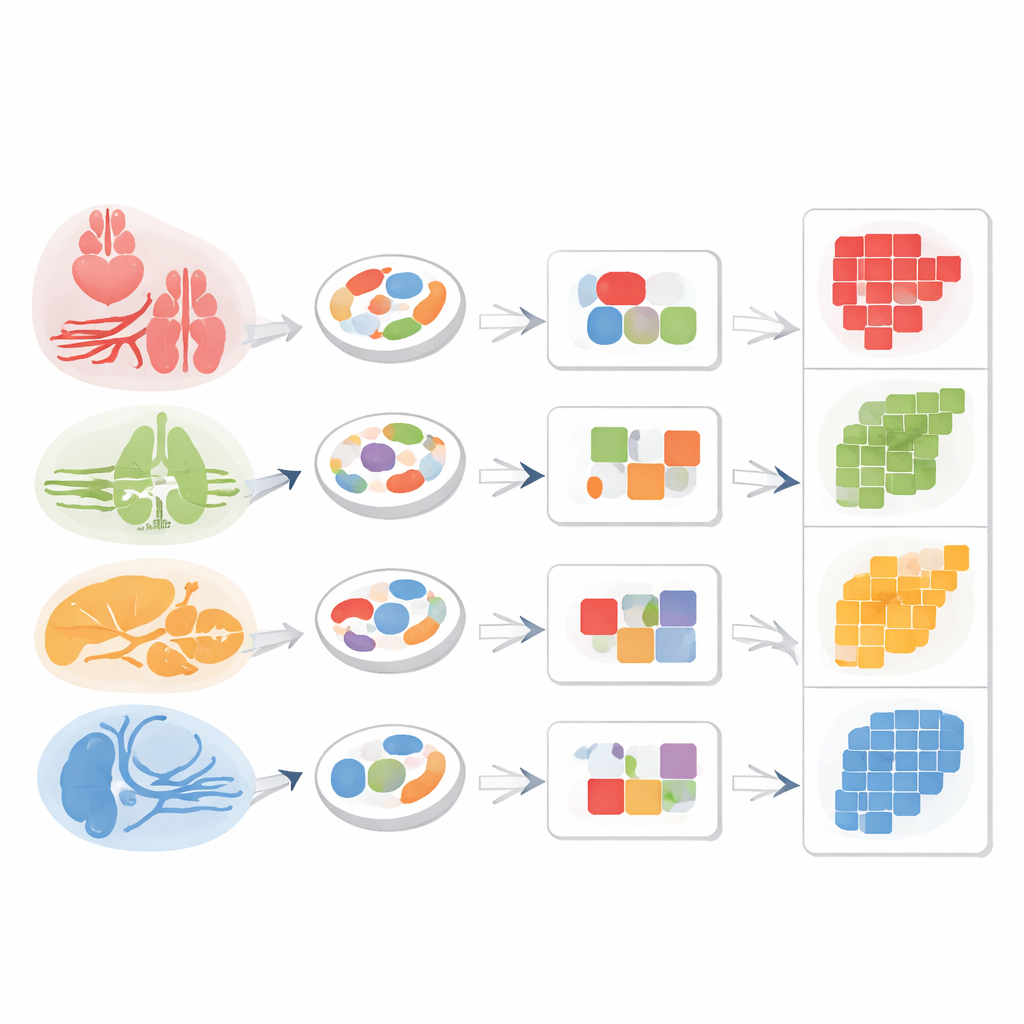
What this atlas means for health and disease
In everyday terms, this work shows that cells do not just carry a generic label like “blood vessel” or “support cell.” Their DNA is packaged in ways that remember which organ they serve and what tasks they perform there. Because chromatin states can persist as cells move or transform—for example, during cancer spread—such maps could eventually help trace where a tumor began or distinguish harmful from healthy cell states. The atlas the authors provide is a reference for normal mouse tissues, offering a baseline against which diseased or experimentally manipulated cells can be compared, and highlighting how a shared genome is sculpted into diverse organs through the opening and closing of regulatory DNA.
Citation: Nooranikhojasteh, A., Tavallaee, G., Khuu, N. et al. Chromatin accessibility landscapes define stromal cell identities across tissues. Commun Biol 9, 480 (2026). https://doi.org/10.1038/s42003-026-09720-w
Keywords: chromatin accessibility, stromal cells, single-cell epigenomics, tissue identity, gene regulation