Clear Sky Science · en
Low-resolution FAIMS for increased peptide coverage in low-load and single-cell proteomics
Seeing More from Less
Modern biology often works with vanishingly small samples—sometimes the contents of a single cell. To understand what is happening inside these tiny worlds, scientists rely on tools that can detect thousands of different proteins at once. This study shows that by gently changing how one such tool, called FAIMS, is operated, researchers can see many more protein pieces from the same tiny amount of material, without buying new instruments or rewriting software.
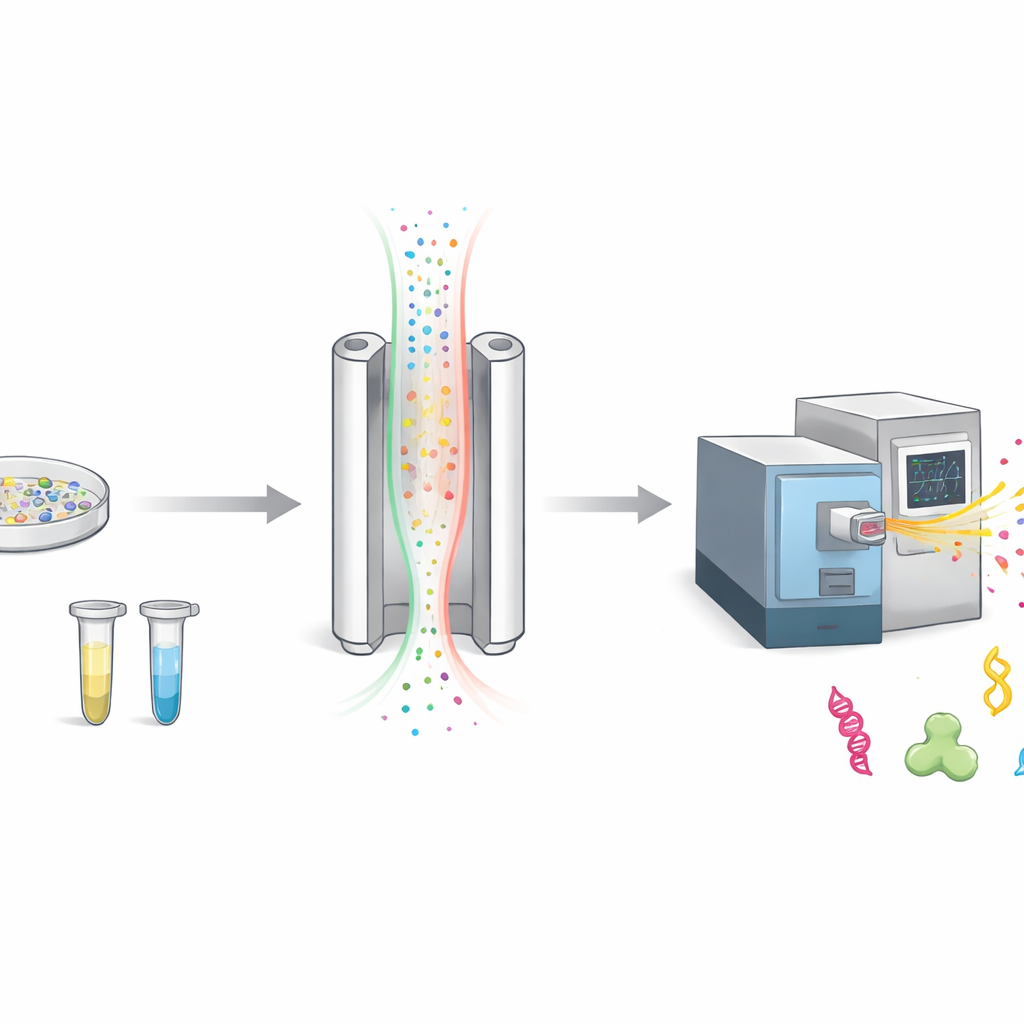
How We Sort Charged Molecules in the Air
When scientists study proteins, they usually chop them into smaller pieces called peptides and then weigh them with a mass spectrometer. Before weighing, FAIMS adds an extra step of sorting. It sends charged molecules through a narrow gap between two metal plates while rapidly switching an electric field on and off. Only molecules whose motion matches a chosen setting slip through to the detector; the rest are discarded as background. This makes the measurement cleaner, but it also means that many potentially interesting molecules never make it to the instrument.
The Trade-Off Between Sharpness and Brightness
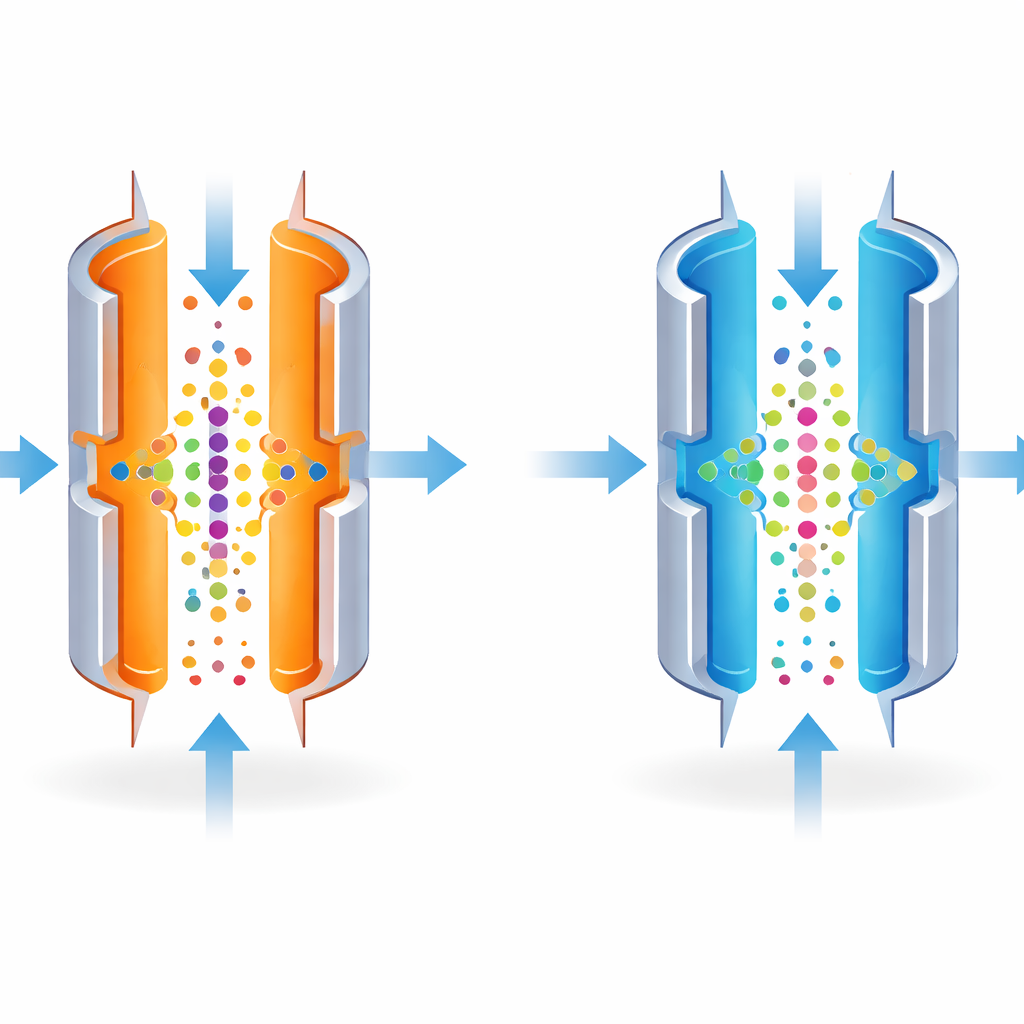
FAIMS can be tuned to separate molecules very precisely, like a camera lens focused on a single point. The sharpest settings, however, only let through a narrow band of molecules, dimming the overall signal. The authors realized that the temperature of the metal plates strongly affects how tightly this band is defined. By making the outer plate cooler than the inner one, they intentionally made the sorting less strict. This broadened the range of molecules that could pass through at a given setting, allowing more ions to reach the mass spectrometer in a single run.
Turning Down the Heat to Reveal Hidden Peptides
The team first tested the idea with standard mixtures of known molecules and with very small amounts of a well-studied cell line (HeLa). Lowering the temperature of the outer plate from 100 °C to 80 °C expanded the window of molecules that passed through FAIMS and increased the total number of detected ions. In practical terms, this meant that the instrument identified 25–34% more distinct peptides, and modestly more proteins, from the same tiny sample. These gains were largest at the lowest sample amounts, where every additional detected molecule matters most.
Better Views Inside Single Cells
The researchers then turned to true single-cell samples from a lung cancer cell line called H460. Using the gentler FAIMS setting again increased both the number of peptides and the number of proteins detected from each cell. The extra sensitivity improved the "fingerprints" that describe each cell’s protein makeup, making differences between cells clearer. Control wells that contained no cells also produced signals, but these mostly reflected background and handling steps. The key observation was that simply changing the plate temperature shifted how many meaningful proteins could be seen inside each individual cell.
A Simple Tweak with Big Payoff
Overall, the study shows that deliberately lowering the resolution of FAIMS—by cooling one of its plates—can substantially boost how many protein pieces are detected from ultra-small samples, including single cells. For non-specialists, the takeaway is that sometimes loosening a filter lets you see more of what matters: here, a slightly blurrier separation in FAIMS leads to a brighter overall picture of the cell’s protein landscape. Because this change uses existing hardware and straightforward settings, many laboratories can adopt it quickly to deepen their view into biology at the smallest scales.
Citation: Hoch, D.G., Belford, M., Heil, L.R. et al. Low-resolution FAIMS for increased peptide coverage in low-load and single-cell proteomics. Sci Rep 16, 14454 (2026). https://doi.org/10.1038/s41598-026-45228-3
Keywords: single-cell proteomics, FAIMS, mass spectrometry, protein analysis, ion mobility