Clear Sky Science · en
Unsupervised learning reveals novel disease-associated proteins in high-dimensional human proteomic data
Blood Clues Hidden in Plain Sight
Our blood is full of proteins that quietly reflect what is happening inside our bodies long before we feel sick. Doctors already use a handful of these proteins as tests, but modern technology can now measure thousands at once. This study asks a simple question with big implications: if we let a computer freely explore these massive blood protein maps without telling it what to look for, can it discover new links to diseases that we did not even think to test?
Letting the Data Speak for Itself
Most medical algorithms are trained with clear labels such as “healthy” or “has hypertension.” That approach is powerful but can miss unexpected patterns, especially when faced with thousands of measurements per person. Here, the researchers take the opposite route: they use “unsupervised” learning, which groups people purely by similarity in their blood protein patterns, without knowing who has which disease. The team worked with a huge resource, the UK Biobank, focusing on nearly 53,000 participants whose blood had been tested for 2,923 different proteins. Their goal was to see whether natural groupings in this ocean of numbers would line up with real-world illnesses and reveal new suspects among the proteins.
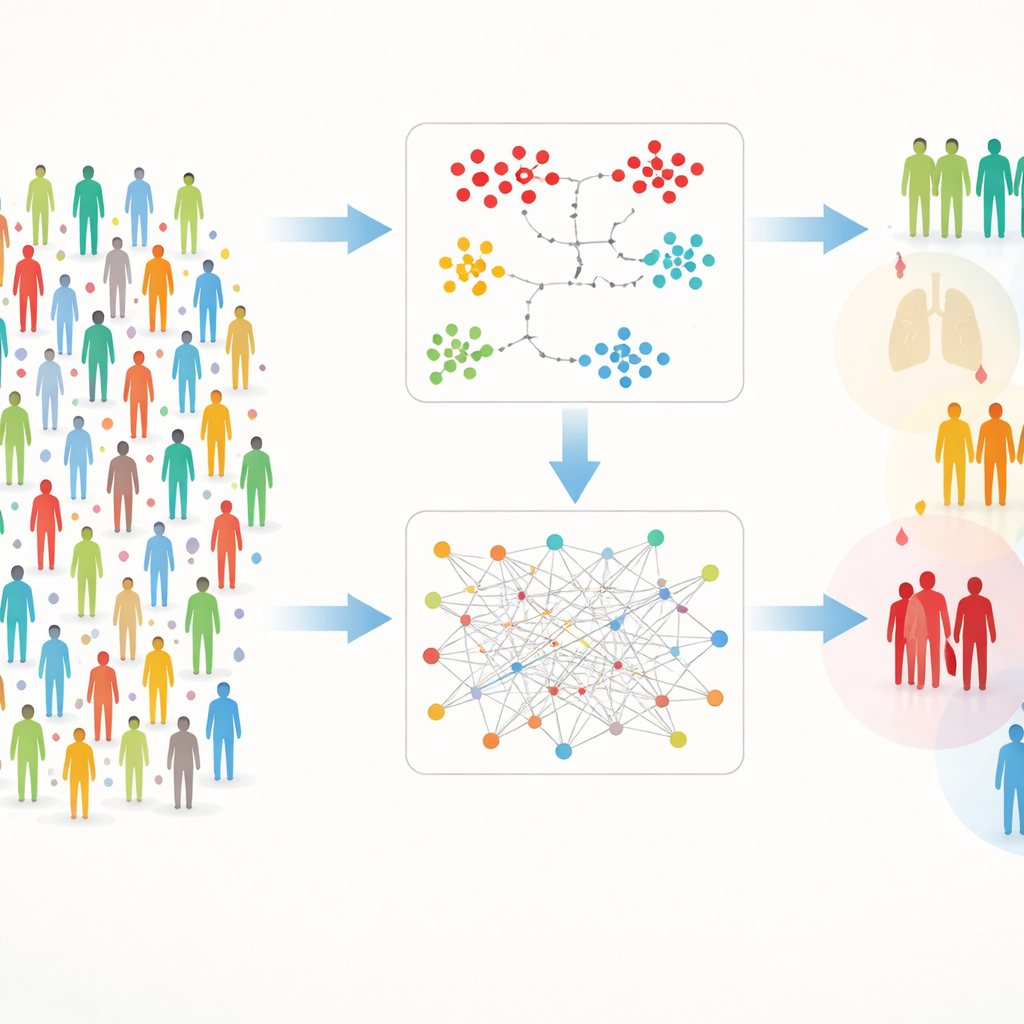
Two Paths to Finding Hidden Groups
Working with such rich data comes with practical hurdles: measurements are sometimes missing, and the sheer number of proteins can drown out signals. To tackle this, the authors built a twin-track framework they call DIRAM/COD. One track (DIRAM) slices the data to avoid missing values, reduces the complexity to two dimensions and then looks for dense “islands” of similar people. The other track (DIRCOD) first fills in missing values with careful estimates, then uses a community-finding method borrowed from network science to detect groups. Both tracks are repeatedly refined, eventually yielding 55 distinct clusters of participants whose blood protein fingerprints differ in meaningful ways.
Clusters that Mirror Real Diseases
Once the clusters were defined, the team checked how age, sex and, crucially, medical diagnoses were distributed across them. Certain clusters were enriched for serious conditions such as organ failure, transplants and cancer, suggesting that their protein signatures captured the biology of very ill patients. The authors then zoomed in on three conditions: celiac disease, hypertension and leukemia. They asked which proteins tended to be unusually high or low in clusters where these diseases were more common. By “recreating” disease-rich groups using only those proteins and simple thresholds, they showed that these protein patterns strongly tracked disease odds—even when mixing participants from across the whole study.
New Protein Suspects and Shifting Relationships
This approach did more than confirm familiar actors; it spotlighted new candidates. For hypertension, proteins such as UBE2L6, HNRNPUL1 and BECN1 stood out, all previously tied to blood vessel or heart problems in other work. For celiac disease, IGF2BP3 emerged as especially important, fitting with earlier hints that it helps maintain the intestine’s barrier, alongside other promising proteins like NRXN3 and CACNB1. In leukemia-related clusters, several proteins, including LRCH4, WDR46, SERPINB1 and NUB1, showed altered behavior. Not only were their levels different, but the way they rose and fell together changed, hinting at a rewiring of the body’s internal control systems in cancer and autoimmune disease.
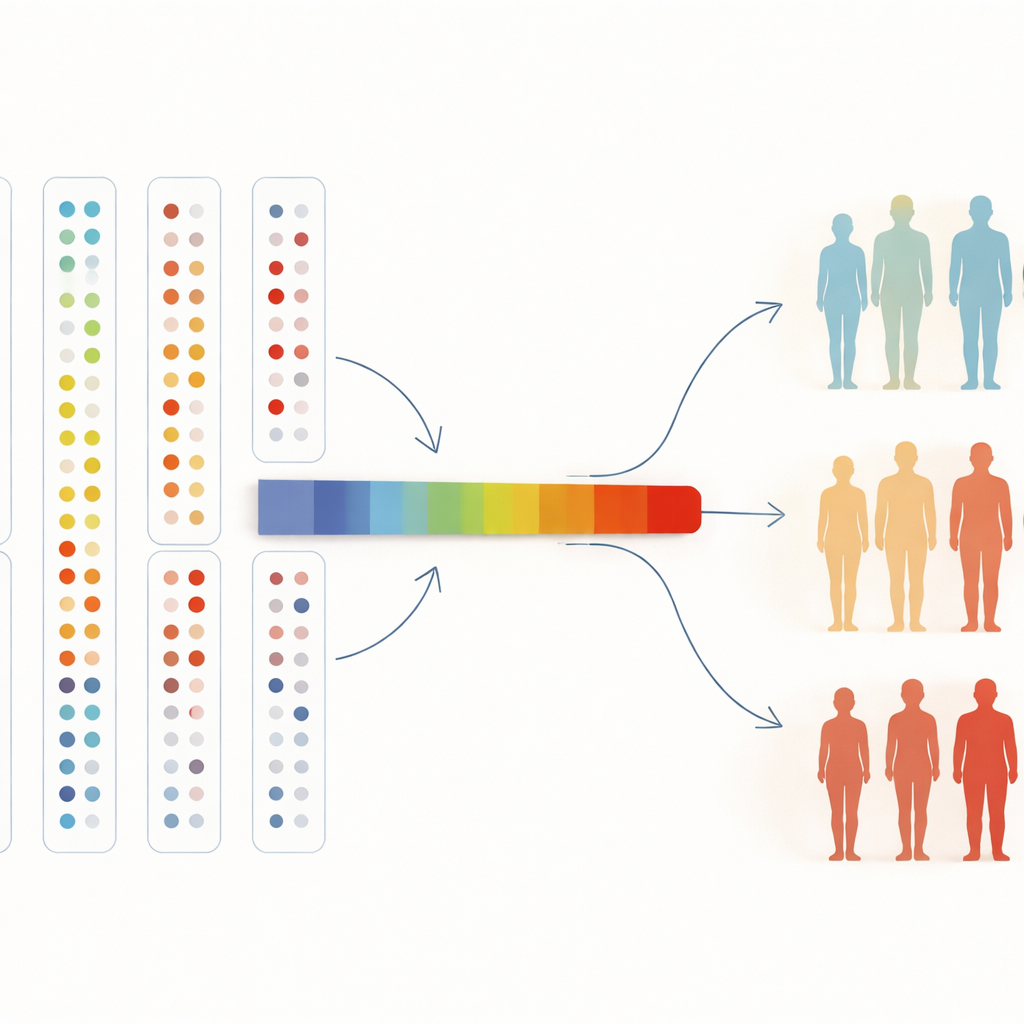
From Blood Maps to Future Medicine
To make sense of many proteins at once, the researchers also compressed them into a single “axis” summarizing overall pattern changes. Along this axis, the chances of having hypertension or celiac disease rose steadily, and this trend held even when they extended the analysis to everyone in the study. To a non-specialist, the takeaway is straightforward: by letting algorithms freely sort people based on thousands of blood proteins, we can find natural health-related groups, confirm known disease markers and uncover new ones. As biobanks grow to hundreds of thousands of people and even more proteins, this kind of unsupervised exploration could help doctors detect disease earlier, understand why some people get sick while others do not and point to fresh targets for future treatments.
Citation: Bernard, E., Wang, Y., Chen, M. et al. Unsupervised learning reveals novel disease-associated proteins in high-dimensional human proteomic data. Sci Rep 16, 10185 (2026). https://doi.org/10.1038/s41598-026-41385-7
Keywords: blood proteins, unsupervised learning, biomarkers, precision medicine, disease risk