Clear Sky Science · en
DeepStackVEGF a stacking ensemble deep learning framework for vascular endothelial growth factor prediction
Why predicting a healing signal matters
Our bodies depend on a protein called vascular endothelial growth factor, or VEGF, to grow new blood vessels. This signal is essential for healing wounds, repairing bone, and supporting normal development—but cancers also hijack it to feed tumors and spread through the body. Measuring and characterizing VEGF in the lab is slow and expensive. This study introduces DeepStack-VEGF, a powerful computer model that can quickly predict whether a given protein behaves like VEGF, potentially speeding up drug discovery and precision medicine.
From lab bench to laptop
Traditionally, researchers use sophisticated techniques such as crystallography, NMR, and tissue staining to study VEGF. These methods reveal the molecule’s structure and location, but they demand specialized equipment and time. At the same time, vast public databases now contain millions of protein sequences whose functions remain only partly known. The authors saw an opportunity: instead of first growing crystals or running complex experiments, why not let computers sift through protein sequences and flag those that are likely to act like VEGF? DeepStack-VEGF was designed as a fast, scalable tool to do exactly that—turn raw protein letters into meaningful predictions.
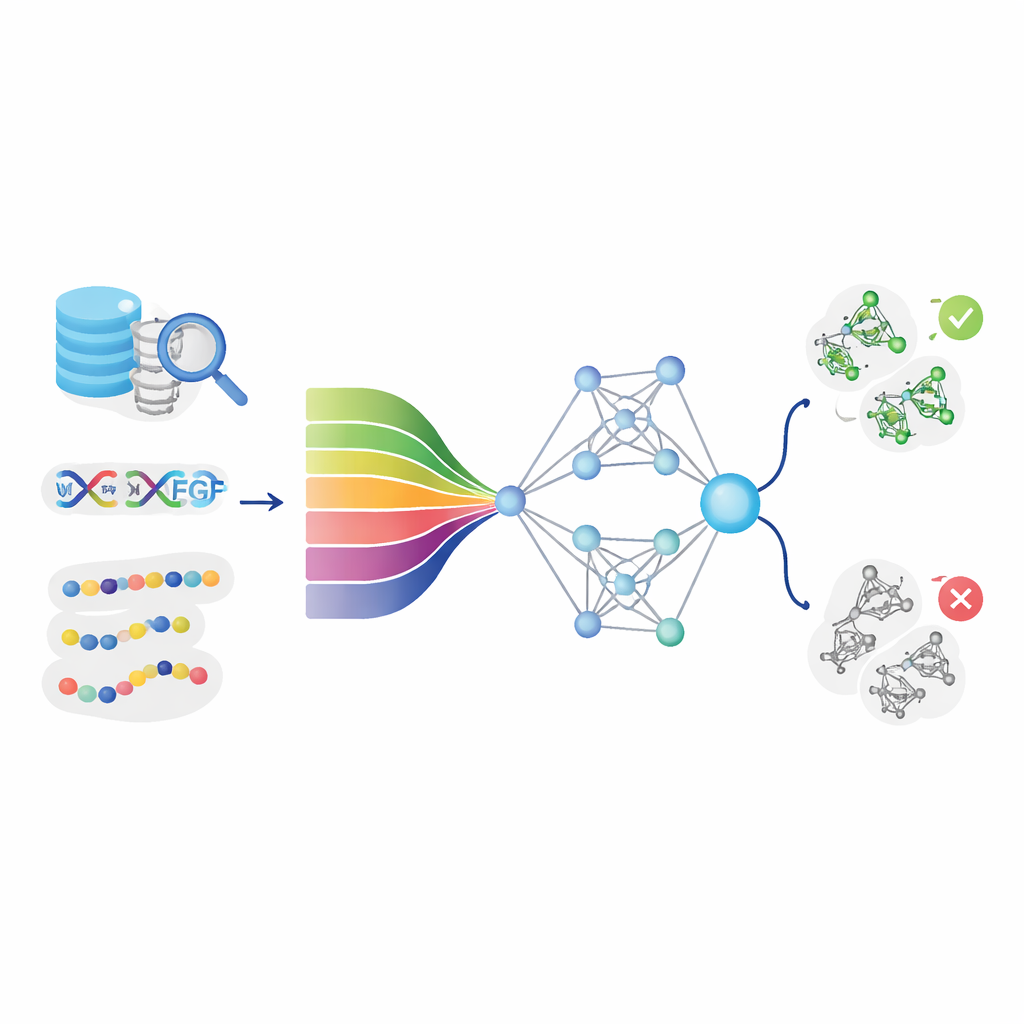
Teaching computers to read protein "language"
The core idea behind DeepStack-VEGF is that a protein’s sequence contains hidden patterns that hint at its behavior. The team gathered thousands of VEGF and non-VEGF proteins from major databases and carefully cleaned the data to avoid near-duplicates. They then described each protein from many angles. Some features captured basic chemistry, such as how oily or charged different positions are. Others summarized how often certain pairs or triplets of building blocks appear, or how the chain is likely to fold into helices and sheets. Crucially, the model also used modern "protein language models"—artificial intelligence systems that, like language tools for text, learn deep patterns from millions of natural protein sequences and turn each one into a rich numerical fingerprint.
Uniting many viewpoints into one decision
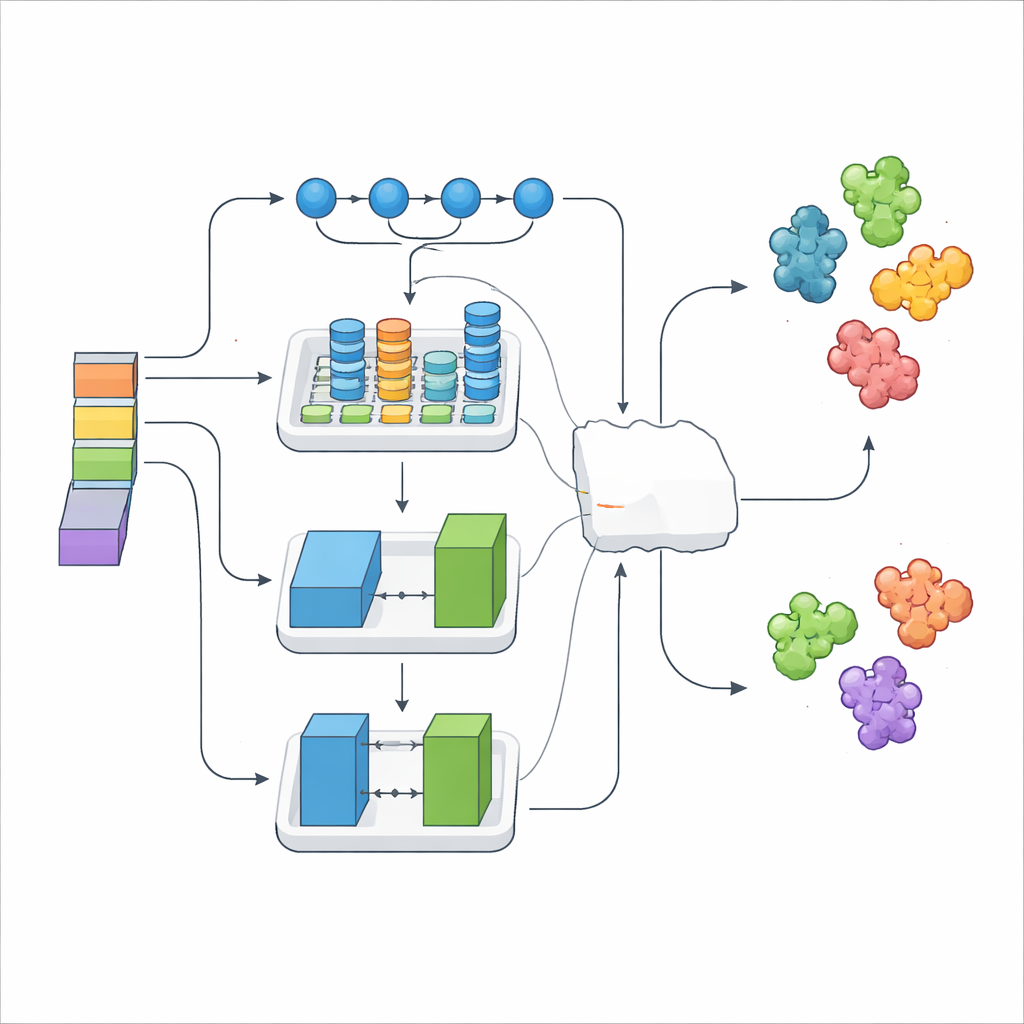
Simply stacking thousands of numerical features can introduce noise, so the researchers used a selection method that keeps only the most informative signals. These refined features were then fed into three different deep-learning modules, each with a distinct specialty. One model excelled at tracing long-range patterns along the sequence, another captured local structural motifs and their relationships, and a third used a game-like generator–critic setup to enrich and regularize the data. On top of these, a "meta" layer learned how to best combine their outputs, forming the DeepStack-VEGF ensemble. This layered strategy mirrors how a panel of experts, each with different training, might weigh in before reaching a joint conclusion.
Checking accuracy and opening the black box
To test their system, the authors used rigorous cross-validation and an independent test set. Across multiple accuracy measures, DeepStack-VEGF outperformed each of its component models and two earlier state-of-the-art VEGF predictors. Its final version correctly classified VEGF-like proteins in well over nine out of ten cases, with fewer false alarms than competing approaches. The team also applied an explanation method that estimates how much each input feature pushes a decision toward "VEGF" or "not VEGF." This analysis showed that the learned protein language fingerprints supplied most of the predictive power, while traditional chemistry- and structure-based features added fine-grained detail and stability.
What this means for medicine and research
For non-specialists, DeepStack-VEGF can be viewed as a highly trained pattern recognizer for a key healing signal in the body. Instead of waiting for laborious experiments, scientists can now feed protein sequences into the model to quickly estimate whether they behave like VEGF. This capability may help narrow down candidates for new cancer or eye-disease treatments, guide the design of anti-angiogenic drugs, and support broader protein research. While any promising prediction still needs laboratory confirmation, tools like DeepStack-VEGF move some of the discovery work from the bench to the computer, potentially making future therapies faster and cheaper to develop.
Citation: Ali, F., Khalid, M., Algarni, A. et al. DeepStackVEGF a stacking ensemble deep learning framework for vascular endothelial growth factor prediction. Sci Rep 16, 13035 (2026). https://doi.org/10.1038/s41598-026-40134-0
Keywords: VEGF prediction, angiogenesis, deep learning in biology, protein language models, drug discovery