Clear Sky Science · en
Complete genome sequence of Sphingomonas sp. gentR, a high-level gentamicin-resistant bacterium
Why a lab microbe matters to everyday medicine
Antibiotics like gentamicin are workhorses in hospitals and farms, but bacteria are steadily learning how to survive them. This study zooms in on a single, unusually tough microbe called Sphingomonas sp. gentR that can shrug off massive doses of gentamicin. By decoding its entire genetic blueprint, the researchers reveal where its drug-resistance tricks are hidden and make those data freely available so others can track, understand, and perhaps disarm similar microbes before their defenses spread to disease-causing bacteria.
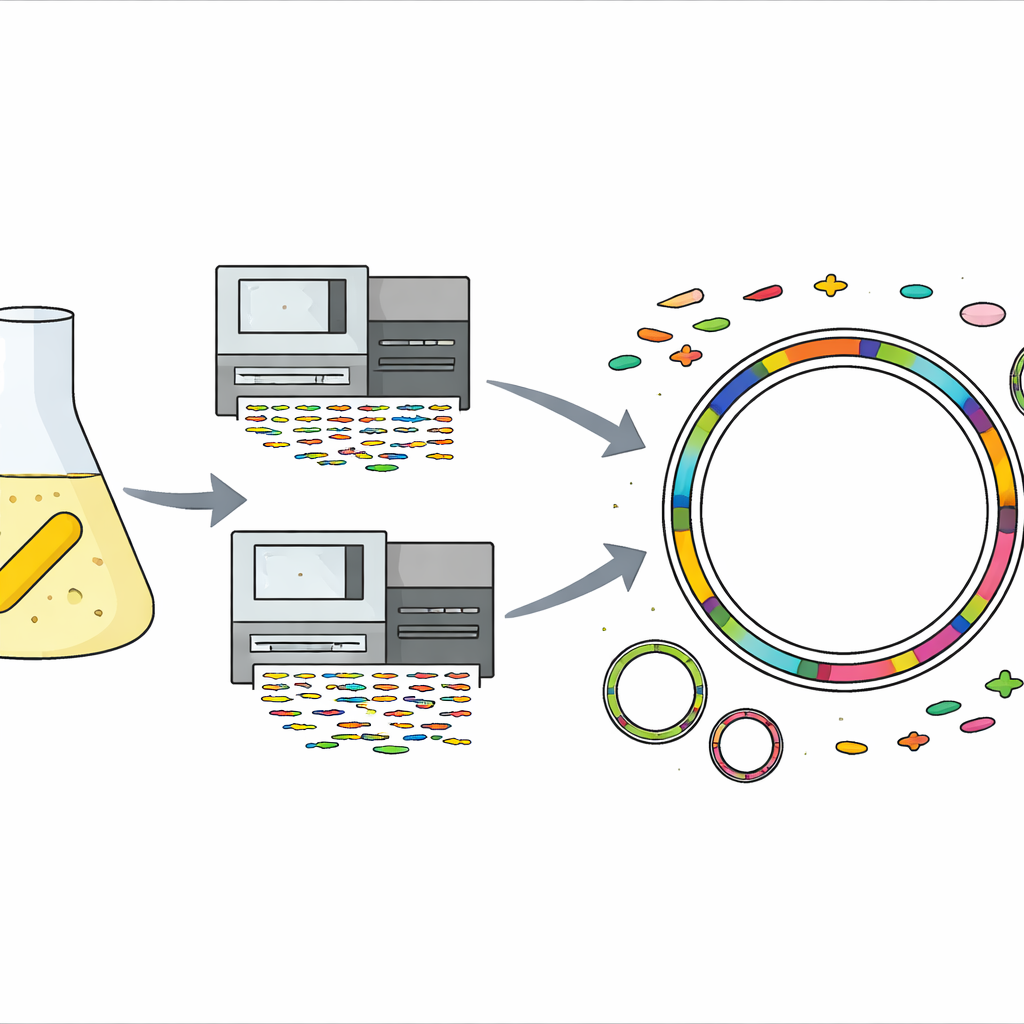
A hardy microbe from an antibiotic bottle
The story begins in an ordinary laboratory, where a working solution of gentamicin—an antibiotic used to treat serious infections—had been stored in the cold. From that very solution, the team unexpectedly isolated a yellow, rod-shaped bacterium belonging to the Sphingomonas group. When they tested how much gentamicin it could endure, the microbe grew even at concentrations about a thousand times higher than those that stop many bacteria, marking it as an extreme survivor. Sphingomonas species are already known for living in harsh places such as desert sands, glacial ice, deep underground rocks, and even spacecraft, but this level of gentamicin resistance had not been seen in the group before.
Reading the microbe’s entire instruction manual
To find out what made this strain so resilient, the scientists extracted its DNA and used two complementary sequencing technologies. One produced many short, accurate DNA fragments, while the other generated fewer but very long fragments. By combining these data, they assembled a complete, gap-free genome for the bacterium: one main circular chromosome and two smaller circular DNA pieces called plasmids. They then used a suite of specialized computer tools and reference databases to predict genes, locate mobile elements like viruses and genomic islands, and assign likely functions to almost every piece of the genome, from basic metabolism to stress responses.
Where the resistance is hiding
With the full genome in hand, the team searched it against curated collections of known antibiotic resistance genes. They found a small set of genes clearly tied to resistance against several drug classes. Most strikingly, three confirmed resistance genes—two that can inactivate aminoglycoside antibiotics such as gentamicin and streptomycin, and one that protects against sulfonamides—sit together on a distinct stretch of DNA embedded in one of the plasmids. This region, known as a genomic island, is the molecular equivalent of a plug-and-play module: a cluster of genes that can move between DNA molecules and, potentially, between bacteria. The genome also carries many genes that fine-tune how the cell senses its surroundings, pumps out toxic compounds, and maintains its membranes, all of which can bolster survival under antibiotic stress.
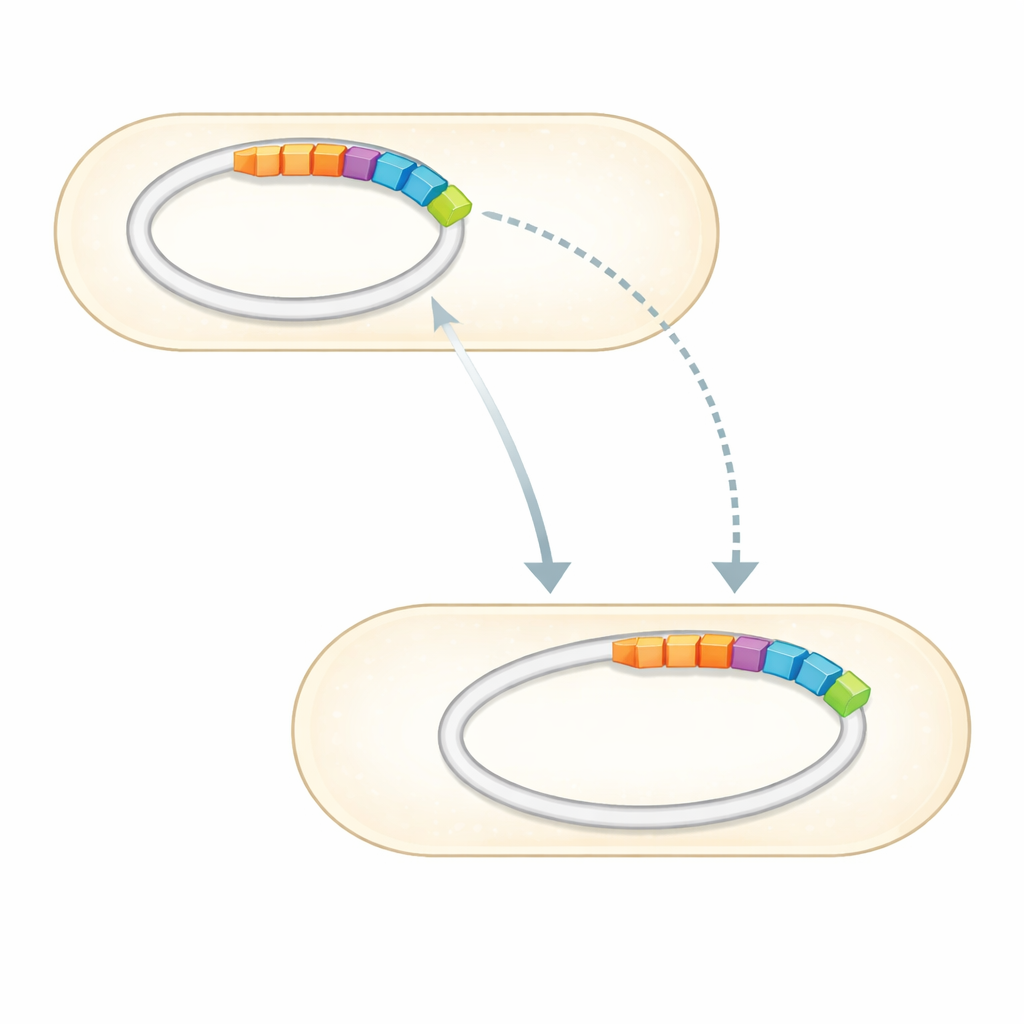
A versatile toolkit for life in tough places
Beyond resistance itself, the gene catalog paints Sphingomonas sp. gentR as a metabolic generalist well-suited to life in poor or polluted environments. Many of its genes support energy production, amino acid use, and the breakdown or transport of diverse molecules, suggesting it can tap into unusual food sources. Others hint at the ability to withstand different stresses and perhaps degrade foreign chemicals. This versatility helps explain why Sphingomonas species are being explored both as plant-friendly partners and as clean-up agents for industrial contaminants, and why an individual strain might flourish in an antibiotic-rich niche like a drug solution.
What this genome means for health and the environment
By fully charting the DNA of an exceptionally gentamicin-resistant Sphingomonas strain and mapping its key resistance genes to a mobile island on a plasmid, the study provides a detailed reference for scientists worldwide. For public health, it highlights a potential pathway by which powerful resistance traits could jump from harmless environmental bacteria into pathogens that infect people, animals, or crops. For environmental science and biotechnology, it offers a genetic toolkit that could be harnessed for cleaning up polluted sites where antibiotics or other chemicals accumulate. In simple terms, the work shows both how tough some background microbes have become and gives researchers the instructions they need to monitor and manage that toughness.
Citation: Liu, Y., Jiang, L., Zhang, J. et al. Complete genome sequence of Sphingomonas sp. gentR, a high-level gentamicin-resistant bacterium. Sci Data 13, 672 (2026). https://doi.org/10.1038/s41597-026-06723-4
Keywords: antibiotic resistance, gentamicin, Sphingomonas, bacterial genome, plasmid genes