Clear Sky Science · en
Calibrating tissue level PDE models of ligand dynamics using single cell and spatial transcriptomics data
How Cells Talk Across a Healing Wound
When our skin is injured, countless cells must coordinate to repair the damage without leaving too much scar tissue behind. They do this by sending and sensing chemical messages called ligands, which spread through the tissue and trigger responses in neighboring cells. This study shows how modern genetic measurements at single-cell and spatial resolution can be turned into a powerful new way to calibrate mathematical models of those chemical messages, helping researchers better understand normal healing and scarring-related diseases.
Why Chemical Messages Matter in Tissues
Cells in our tissues rarely act alone. They constantly exchange signals that tell them when to divide, move, or quiet down. Many of these signals are short-range chemicals released by one cell and captured by receptors on another. Such messages are central to wound healing, inflammation, and diseases like fibrosis and cancer. Researchers have long used mathematical equations, known as reaction–diffusion models, to describe how these chemicals are produced, spread, and broken down. Yet a long-standing problem has been how to choose realistic parameter values—such as production rates and binding strengths—so that the models truly reflect what happens inside living tissue rather than idealized lab conditions.
New Eyes on Living Tissue
Recently, two experimental technologies have transformed how scientists can observe tissues. Single-cell RNA sequencing measures which genes are active in thousands of individual cells, revealing their identities and signaling capabilities. Spatial transcriptomics, by contrast, measures gene activity across many tiny spots laid out like a grid on a tissue slice, preserving where different cells are located. In this work, the authors combine both data types from human skin samples taken 30 days after wounding, when the tissue is in the “remodeling” stage of healing. They focus on three versions of a key signaling molecule, transforming growth factor beta (TGFβ), known to influence scar formation and tissue repair.
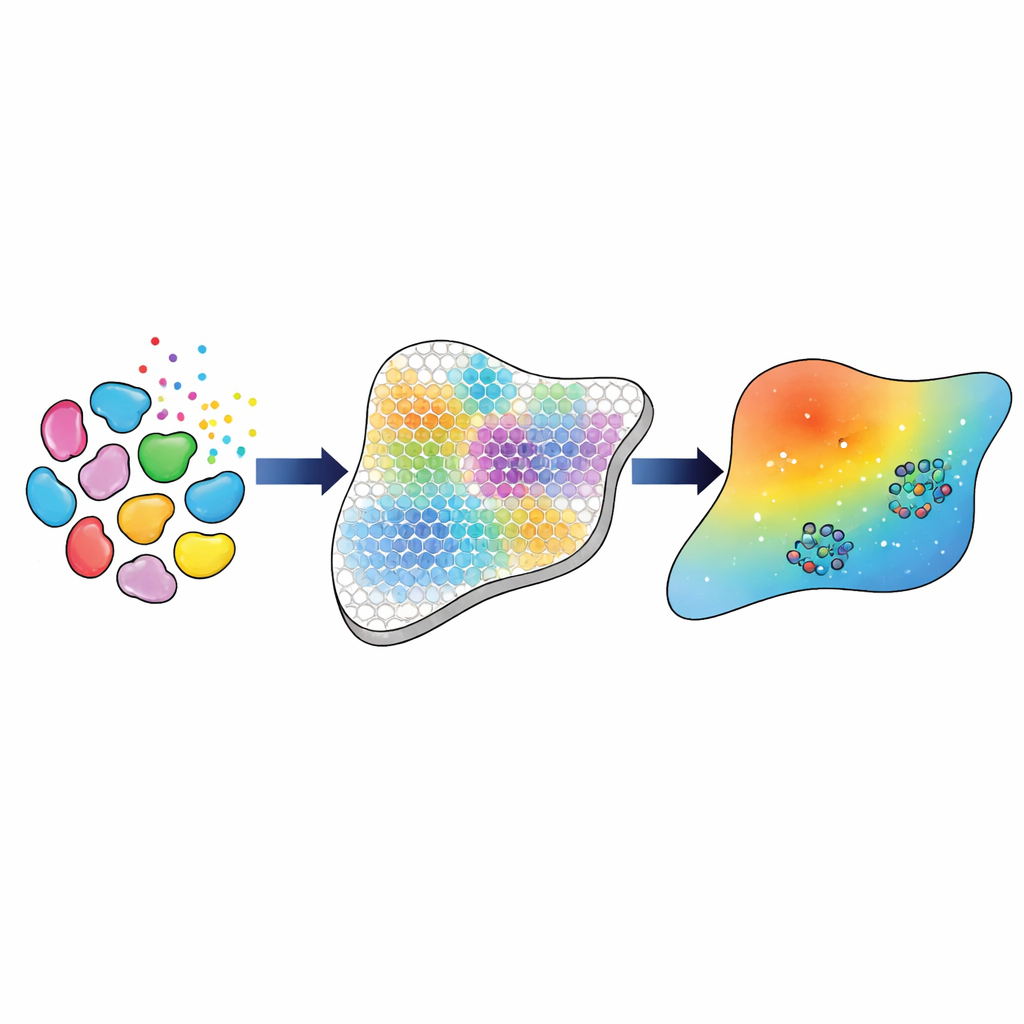
A Stepwise Pipeline from Genes to Model Parameters
The researchers build a computational pipeline that connects these rich genetic measurements to a tissue-scale model of TGFβ movement and action. First, they use established bioinformatics tools to identify the main cell types and subtypes—such as three distinct classes of fibroblasts, macrophages, and endothelial cells—and to estimate how many of each are present at every spot in the tissue grid. They then infer how strongly each type of cell is likely to talk to each other via TGFβ, based on the combined gene activity of ligands in sending cells and receptors in receiving cells. These inferred “interaction strengths” serve as the experimental target that the mathematical model must reproduce.
Teaching the Model to Match the Tissue
Next, the team represents TGFβ spreading through the tissue with a set of partial differential equations solved on a grid that mirrors the spatial transcriptomics layout. For any chosen set of parameter values—such as how fast TGFβ diffuses, how quickly it decays, and how much each cell type produces or absorbs—the model predicts concentration fields of the three TGFβ forms and the resulting interaction strengths between cell types. To tune those parameters, the authors use a three-stage calibration strategy. They first apply approximate Bayesian methods to scan broadly over plausible parameters informed by previous experiments, gradually narrowing in on combinations that correlate well with the interaction strengths derived from the genetic data. They then feed these promising candidates into a gradient-based optimization method that fine-tunes the parameters to maximize agreement. This hybrid approach achieves an almost perfect linear match (correlation of 0.99) between model-predicted and data-inferred interaction strengths.
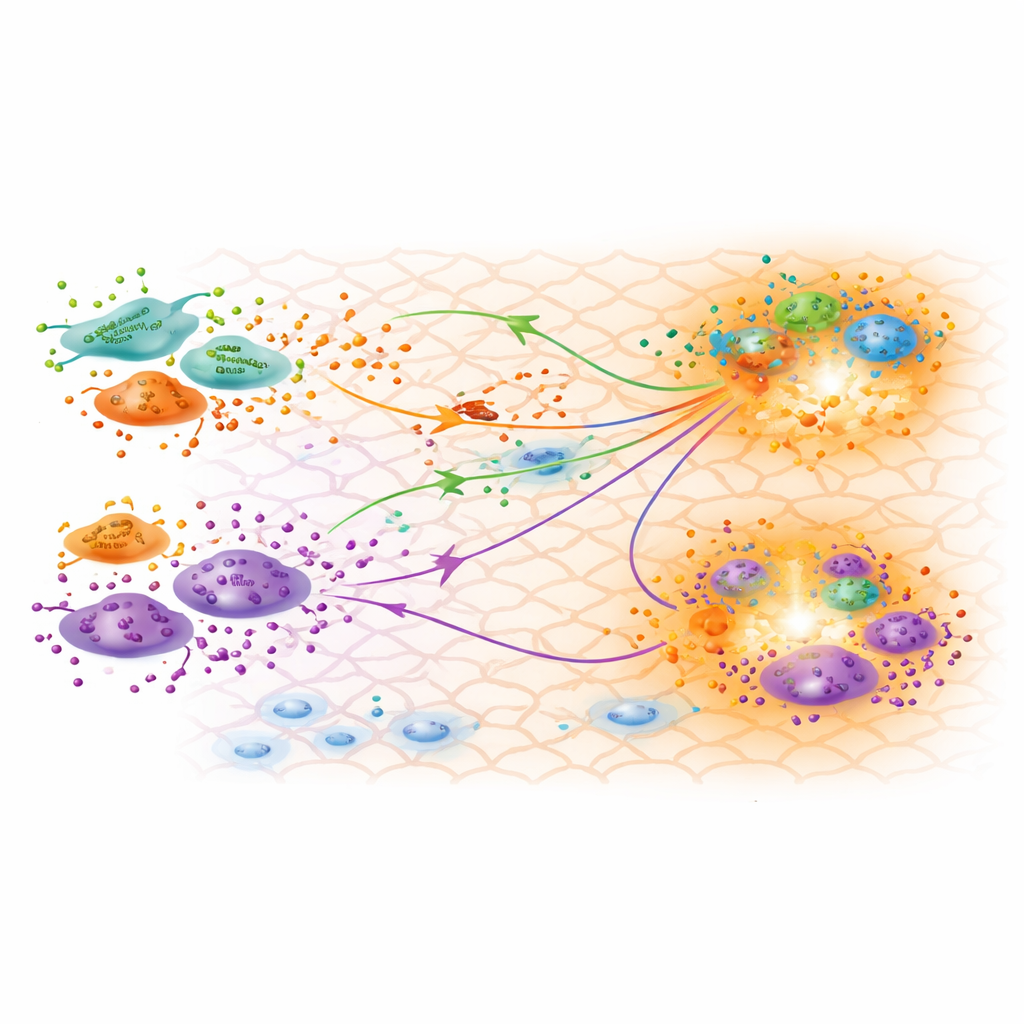
What the Calibrated Model Reveals About Healing
With a well-matched model in hand, the researchers explore what the inferred parameters say about biology. They confirm that the three fibroblast subtypes identified from gene expression data are functionally distinct, with one showing a strong inflammatory profile and another displaying features of myofibroblasts that help contract the wound. The calibrated production rates suggest that pro-inflammatory fibroblasts produce the least TGFβ during the remodeling stage, consistent with inflammation subsiding by day 30. Surprisingly, a subtype known as papillary fibroblasts appears to produce relatively high amounts of TGFβ3, a form often linked to reduced scarring and more regenerative outcomes. The model’s predicted TGFβ concentration maps align with regions where fibroblasts and macrophages cluster, hinting at hotspots of active cross-talk that may steer how the scar forms.
A Flexible Framework for Future Tissues and Diseases
Beyond this specific wound-healing case, the study’s main contribution is a general pipeline for turning modern genetic data into well-calibrated tissue-level models. By combining single-cell and spatial transcriptomics with advanced statistical and optimization tools, researchers can more rigorously estimate how fast ligands move, how strongly they bind, and which cells contribute most to signaling. This framework can be adapted to other tissues, other signaling molecules, and disease states such as fibrotic scarring or cancer, ultimately helping translate complex molecular snapshots into predictive models of how whole tissues behave.
Citation: Daher, A., Trucu, D. & Eftimie, R. Calibrating tissue level PDE models of ligand dynamics using single cell and spatial transcriptomics data. npj Syst Biol Appl 12, 44 (2026). https://doi.org/10.1038/s41540-026-00657-8
Keywords: wound healing, cell signaling, spatial transcriptomics, TGF beta, mathematical modeling