Clear Sky Science · en
4polar3D single molecule imaging of 3D orientation in dense actin networks using ratiometric polarization splitting
A New Way to See the Hidden Order Inside Cells
Inside every living cell, long, thin protein filaments form a scaffold that gives the cell its shape and helps it move. These fibers are crowded and tangled, so even advanced microscopes often see only a blur of dots. This study introduces a new imaging method that can reveal not just where these tiny molecules are, but how they are oriented in three dimensions, letting scientists map the hidden architecture of this inner scaffold in intact cells.
Why Looking at Direction Matters
Many proteins in cells act like tiny arrows: their direction in space affects how they interact, assemble, and generate forces. Traditional super-resolution microscopes can pinpoint the position of single fluorescent molecules but usually ignore how they are tilted or rotated. For densely packed structures such as actin filaments at the cell edge, knowing positions alone is not enough to understand how fibers are arranged or how they push and pull on the cell membrane. A technique that can capture both position and orientation at once, over large areas, can bridge the gap between detailed structural methods like electron microscopy and faster optical methods used in living cells.
A Simpler Optical Trick for 3D Orientation
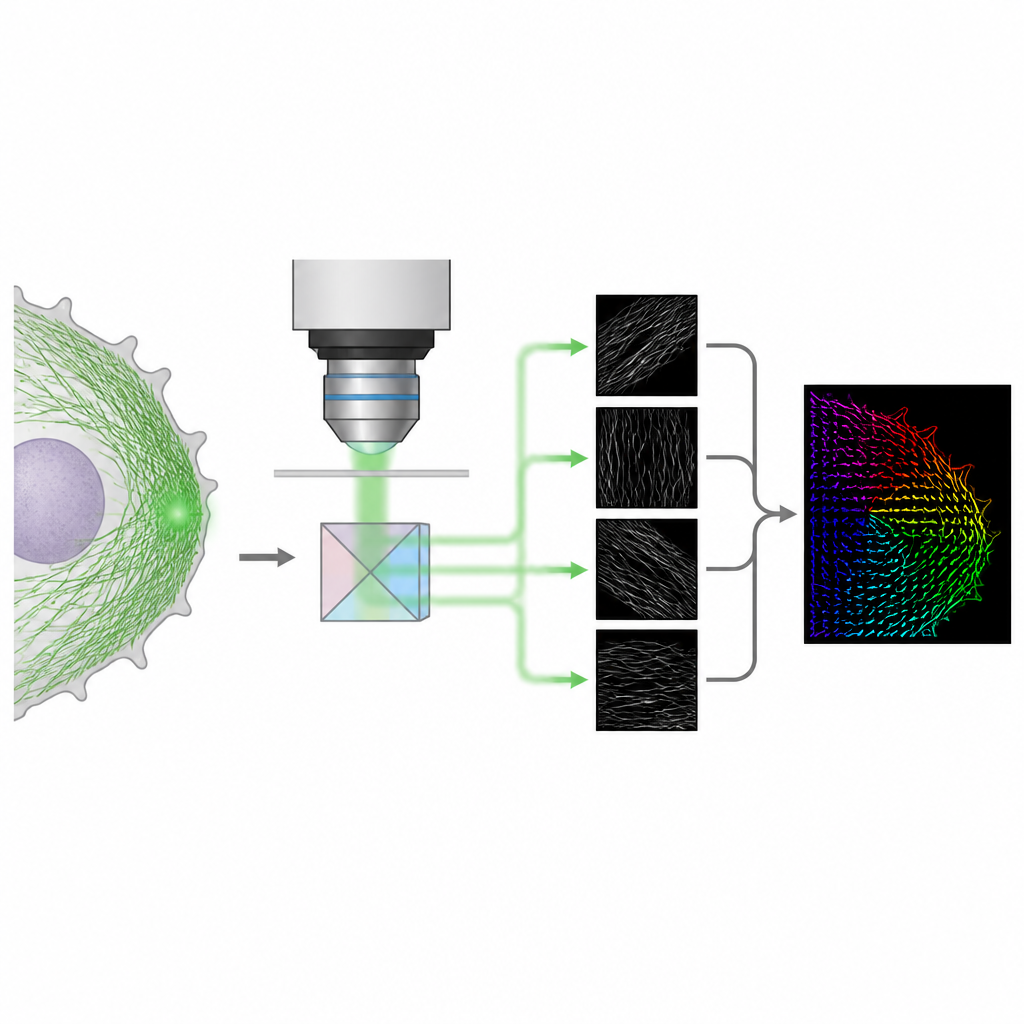
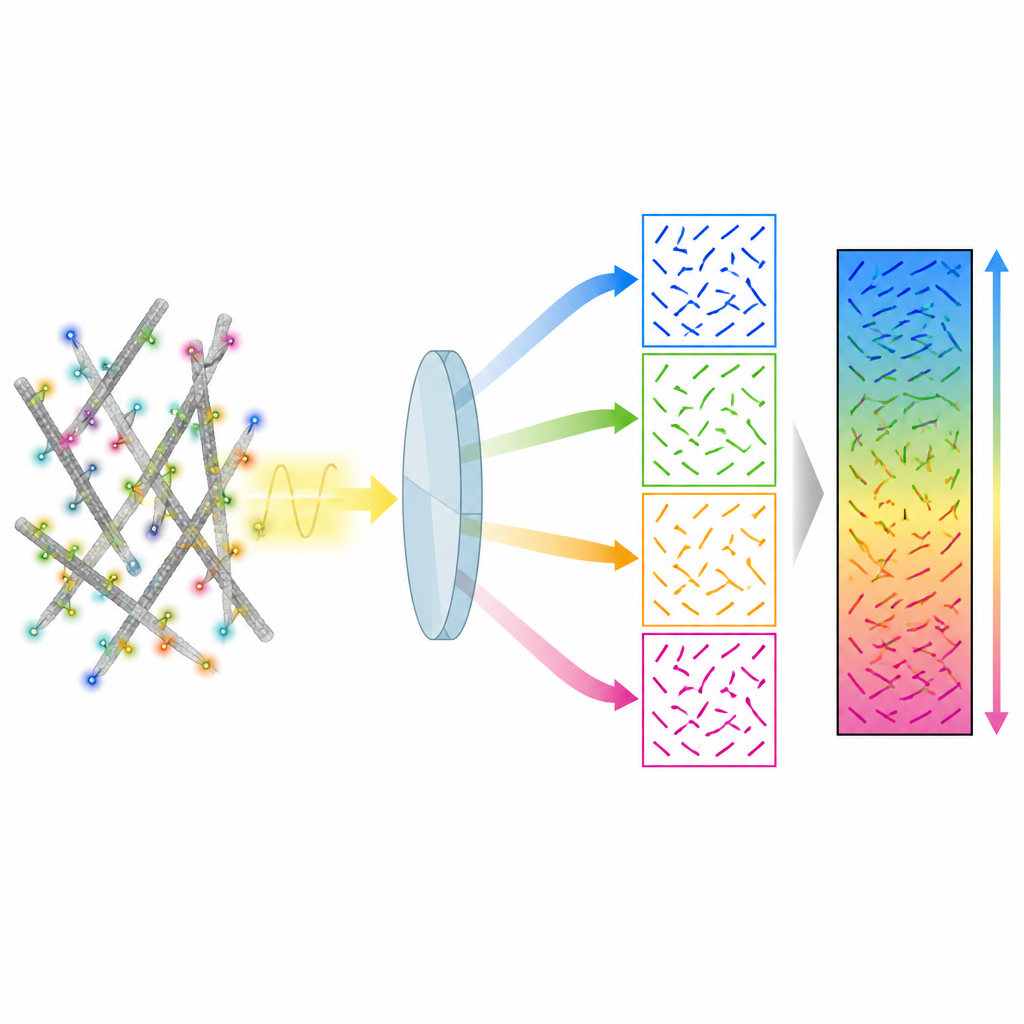
The authors present a method called 4polar3D that measures how single fluorescent molecules are oriented and how much they wobble, while also locating them in the cell. Instead of reshaping the light pattern from each molecule with complex optics, the system splits the emitted light into four beams, each filtered by a different polarization and numerical aperture. By comparing how bright the same molecule appears in these four channels, the method extracts three key properties: the direction of the molecule in the plane of the microscope slide, how much it is tilted out of that plane, and how freely it can rotate. Because it relies only on integrated brightness rather than detailed pattern fitting, the data analysis is fast and robust, even when many molecules emit light at the same time.
Testing the Method on Model Membranes
To check that 4polar3D works as expected, the team first tested it on simple lipid membranes labeled with a fluorescent dye that tends to align with the fatty acid chains. On flat supported membranes, the method measured a preferred tilt of the dye molecules and a broad wobble, in agreement with earlier, more complex approaches. When the same membrane was wrapped around tiny silica beads, the measured orientations followed the curvature of the sphere: molecules near the bottom pointed more toward the microscope, while those near the sides became more parallel to the surface. These tests showed that 4polar3D can accurately recover a wide range of tilt angles and distinguish between in-plane and off-plane orientations.
Revealing 3D Actin Architecture in Living-Like Cells
The researchers then turned to the dense actin networks that drive cell movement and adhesion. In fast-moving melanoma cells, the thin sheet at the front of the cell, called the lamellipodium, is packed with actin filaments. Using a fluorescent tag that binds along these filaments, 4polar3D recorded millions of single-molecule events and reconstructed a detailed map of filament directions. Near the very edge, the actin network showed a mix of in-plane branched filaments, with a characteristic pair of preferred directions, and a second population of filaments tilted out of the plane with more varied orientations. Further back, in the transition region and in thick stress fibers, filaments became mostly in-plane and aligned along clear directions.
Uncovering 3D Structure in Tiny Adhesion Organelles
The method also probed podosomes, tiny adhesion structures in immune cells that are smaller than the diffraction limit of light. By averaging data from hundreds of podosomes, the authors found a clear pattern: at the center, many actin filaments were tilted out of the plane and likely extended upward, while around the edges, in-plane filaments formed a radial ring that pointed outward from each core. Between neighboring podosomes, this ordered radial pattern was largely lost. Measurements of the apparent focus of the fluorescent spots suggested that off-plane filaments in the core sit higher above the glass surface than the surrounding in-plane ring, supporting a picture of a layered, three-dimensional actin scaffold.
What This Means for Cell Biology
In summary, 4polar3D offers a practical way to map how molecules are pointed in three dimensions across large areas of a cell, using a relatively simple optical setup and fast computations. While it is less precise at extreme tilt angles than some more elaborate methods, it works well for most orientations and, crucially, can handle crowded, biologically relevant samples. By revealing how actin filaments weave in and out of the cell plane in lamellipodia and podosomes, this approach opens the door to routine studies that link nanoscale fiber organization to cell movement, force generation, and signaling.
Citation: Senthil Kumar, C.S., Valades Cruz, C.A., Sison, M. et al. 4polar3D single molecule imaging of 3D orientation in dense actin networks using ratiometric polarization splitting. Nat Commun 17, 4246 (2026). https://doi.org/10.1038/s41467-026-70852-y
Keywords: single molecule imaging, actin cytoskeleton, polarization microscopy, super resolution, cell mechanics