Clear Sky Science · en
An orthogonal RNA aptamer for targeted RNA regulation in phase-separated subcellular compartments
Watching and Controlling RNA Inside Tiny Cell Droplets
Inside every cell, strands of RNA are constantly being made, moved, and destroyed. Many of these RNAs gather in tiny liquid-like droplets that float in the cell’s interior. These droplets help organize life’s chemistry, but they are hard to study in real time. This paper describes a new molecular toolkit that lets scientists both light up specific RNAs and deliberately erase them inside these droplets, revealing what those RNAs actually do in living cells.
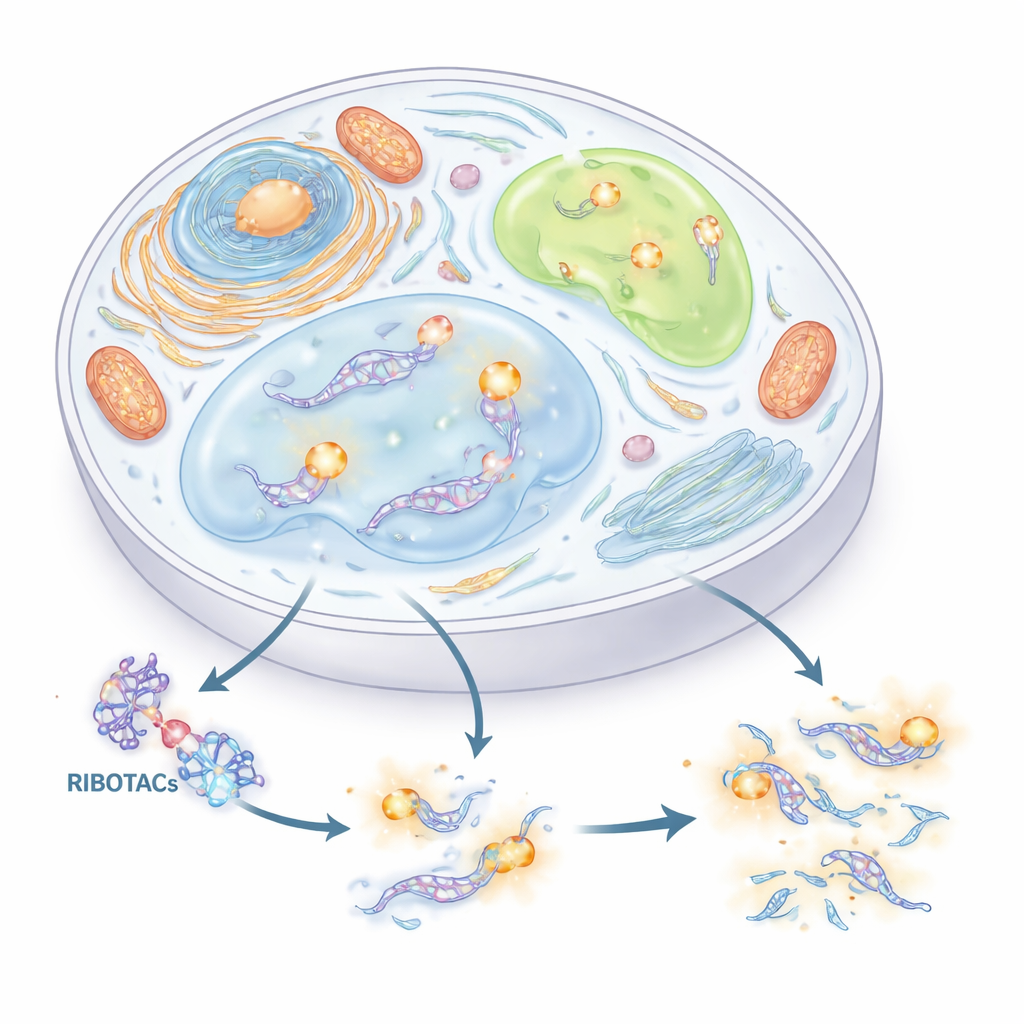
A New Handle for Finding and Cutting RNA
The authors set out to build a precise, two-in-one system for tracking and controlling RNA. They combined two ideas. The first is a short RNA “tag” (an aptamer) that can fold into a shape and grab a matching small molecule. The second is a class of designer molecules called RIBOTACs, which latch onto a chosen RNA and then recruit a natural RNA-cutting enzyme to destroy it. By carefully screening thousands of small molecules and hundreds of RNA shapes, the team identified a compact RNA tag, derived from a viral sequence called HIV-TAR, and optimized it into an aptamer they call HT. They then built RIBOTACs that recognize HT and drag in the cell’s RNase L enzyme, turning any RNA that carries HT into a selectively destroyable target.
Adding a Bright Color to Follow RNA in Living Cells
To make these tagged RNAs visible, the researchers fused HT to a separate fluorescent aptamer called Clivia, which glows orange-red when it binds a harmless dye (NBSI). The resulting hybrid tag, Clivia-HT, gives each labeled RNA two distinct abilities: the Clivia side attracts the glowing dye for imaging, while the HT side is recognized by the RIBOTAC for controlled cutting. The team showed that these two functions operate independently: the dye lights up Clivia without affecting degradation, and the RIBOTAC binds HT and triggers cutting without dimming the fluorescent signal until the RNA itself is removed. By repeating Clivia-HT several times on one RNA, they boosted brightness enough to follow tagged molecules in live cells.
Probing Hidden RNA Roles in Stress Droplets
Armed with this dual tag, the authors explored RNAs inside phase-separated subcellular compartments—droplet-like structures that form without membranes. They first inserted Clivia-HT into U1 small nuclear RNA, which marks U bodies, a type of RNA-rich granule. When cells were stressed, these U bodies lit up; adding the HT-targeting RIBOTAC then gradually erased the signal and reduced the number of U bodies, but only when the cutting enzyme RNase L was present. Next, they labeled ATF4 messenger RNA inside stress granules, droplets that appear during cellular hardship. Although ATF4 RNA clearly accumulated in these granules and could be degraded on demand, removing it did not change how the granules formed or dissolved. This indicates that, despite its presence there, ATF4 RNA is not a key driver of stress granule dynamics.
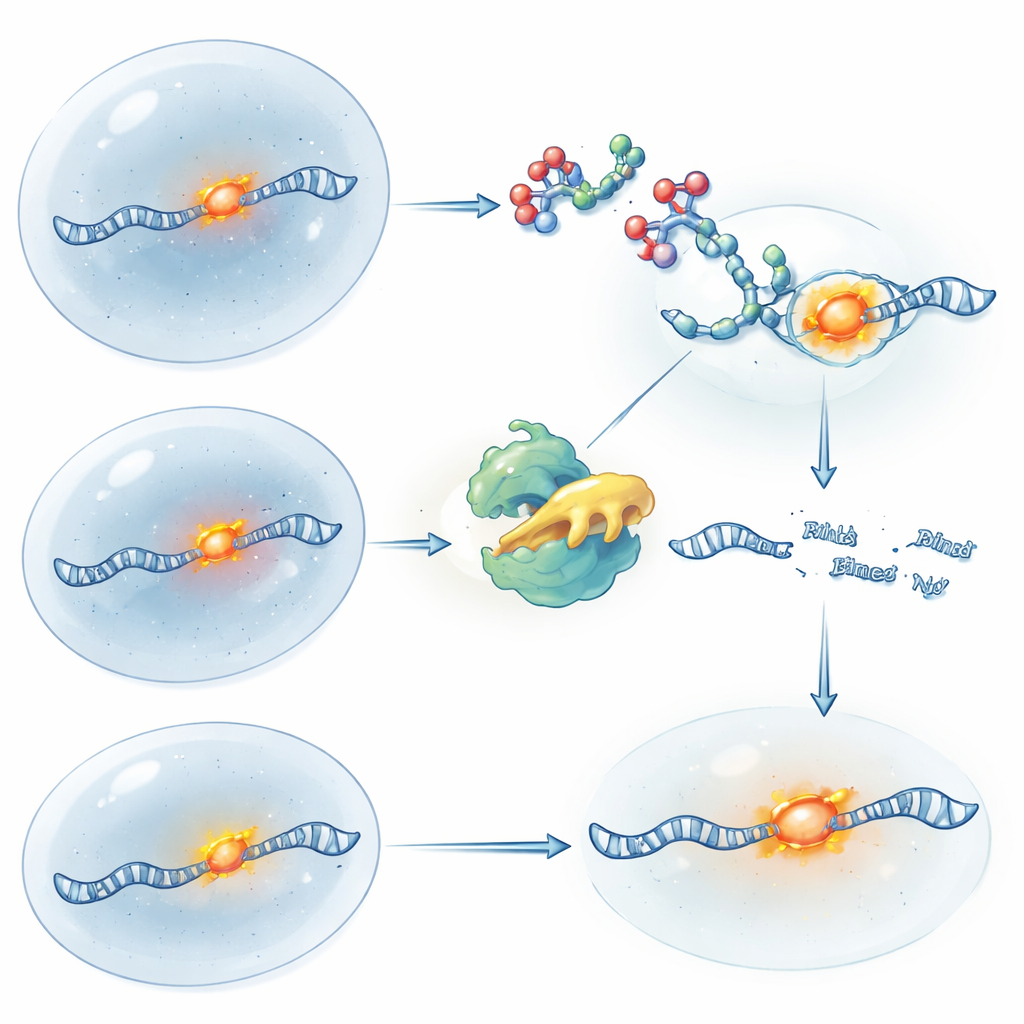
Turning RNA Control On and Off with Light
To gain even finer control over timing, the team made light-sensitive versions of their RIBOTACs. In one design, the molecule starts active and is “switched off” when a brief pulse of ultraviolet light cleaves a built-in safety link, stopping further cutting and allowing fresh RNA to build back up. In the other design, the RIBOTAC is caged and inactive until light removes the blocking group, suddenly unleashing its RNA-cleaving power. Using these tools on ATF4 RNA, the researchers could mimic the logic of gene knockout and gene restoration purely with chemistry and light, all while the RNA’s location was tracked through fluorescence. This approach allows scientists to ask not just where an RNA is, but what happens to the cell when that RNA is removed or restored at precise moments.
Revealing How a Long RNA Protects the Genome
The authors then turned to a long RNA called NORAD, known to help preserve the integrity of chromosomes. NORAD gathers with Pumilio proteins into distinct droplets called NP bodies in the cytoplasm. By tagging NORAD and a minimal NORAD fragment (circPRE8) with Clivia-HT, they could visualize these assemblies and selectively erase the RNA. When NORAD or circPRE8 was present, Pumilio proteins formed bright puncta and cells had fewer chromosome segregation errors. When the tagged RNA was degraded by RIBOTAC, the droplets disappeared, Pumilio proteins dispersed, and chromosome errors increased. These experiments show that specific NORAD regions are essential for forming protective protein–RNA droplets that safeguard the genome.
What This Means for Future Cell Biology
In summary, this work introduces a versatile “plug-in” RNA tag that allows any chosen RNA to be both seen and precisely erased inside the cell, even within delicate liquid-like compartments. By marrying bright imaging with targeted destruction, and adding light-based on–off switches, the Clivia-HT system makes it possible to test whether an RNA’s mere presence in a structure actually matters for cell behavior. This toolkit will help researchers move beyond static pictures of RNA location toward causal experiments that reveal how specific RNAs shape stress responses, droplet formation, and genome stability.
Citation: Wang, J., Ma, K., Cao, X. et al. An orthogonal RNA aptamer for targeted RNA regulation in phase-separated subcellular compartments. Nat Commun 17, 4140 (2026). https://doi.org/10.1038/s41467-026-70638-2
Keywords: RNA imaging, phase-separated condensates, RNA degradation, light-controlled chemistry, long noncoding RNA