Clear Sky Science · en
Multi-species integration, alignment and annotation of single-cell RNA-seq data with CAMEX
Why this research matters
Every animal body is built from a rich cast of cell types, yet we still lack a clear map of how these cells compare across species or change over evolution. This study introduces CAMEX, a computational tool that stitches together single-cell gene activity data from many different animals into a shared picture. For a lay reader, this is exciting because it brings us closer to answering questions such as which cell types are truly universal, which are unique to humans, and how organs like the brain, liver, and testis took shape over evolutionary time.
Looking at cells one by one
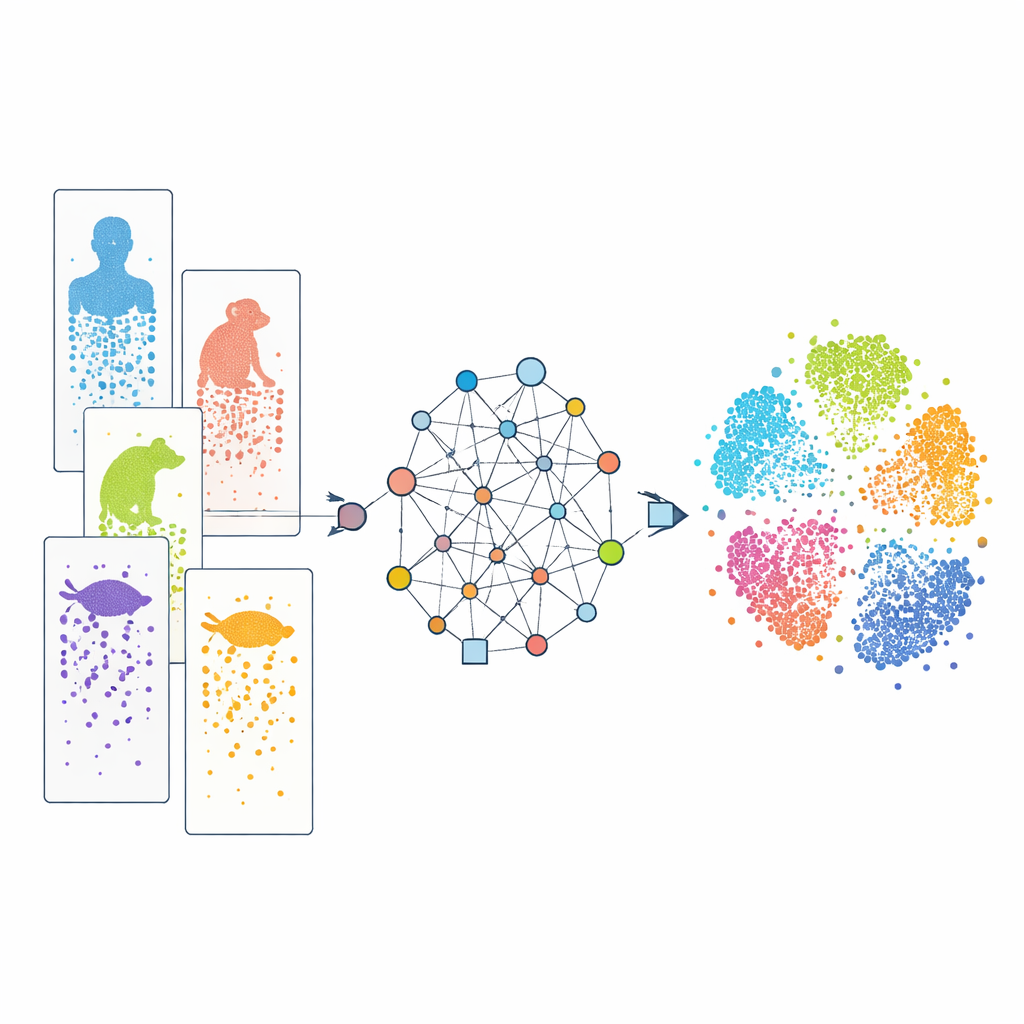
Modern single-cell RNA sequencing allows scientists to read out which genes are active in thousands to millions of individual cells in a single experiment. By comparing these patterns, researchers can sort cells into types and track how they develop. Many such datasets now exist for humans, monkeys, mice, fish, reptiles and more. However, each study often uses different experimental technologies, and species vary in their sets of genes. On top of that, our knowledge of genes is uneven: well-studied laboratory animals are much better annotated than obscure species. These differences act like “batch effects” and incomplete dictionaries, making it difficult to line up similar cells across species and to see which features are truly shared versus species-specific.
A graph-based way to connect species
CAMEX tackles these obstacles by turning all of the data into a single large network that includes both cells and genes. In this network, cells connect to the genes they express, to their most similar neighboring cells, and genes are linked across species if they are judged to be related by evolution, even when the relationship is many-to-many rather than a simple one-to-one match. A specialized type of machine learning model, a heterogeneous graph neural network, then passes information along these connections and learns a compact “embedding” for every cell and gene in a shared low-dimensional space. For data integration, the model is trained to reconstruct both the network structure and the original gene activity patterns without ever being told the cell types in advance. For cell annotation, the same encoder feeds into an attention-based classifier that can transfer known labels from a reference species to less studied ones.
Revealing shared cell types and development
The authors show that CAMEX outperforms a suite of popular tools when challenged with demanding, real-world datasets. In liver, ovary, and pancreas data spanning up to four species and multiple experimental platforms, CAMEX best balanced two competing goals: removing artificial batch differences while keeping true biological distinctions between cell types. It accurately aligned common cell populations such as hepatocytes and immune cells, and, importantly, preserved rare cell types that other methods tended to blur. In a dramatic test, CAMEX integrated testis data from 11 species, from primates to platypus and chicken. It recovered the continuous path by which germ cells mature into sperm and showed that using many-to-many gene relationships is crucial for maintaining performance as species become more distant. The model also successfully aligned organ development stages across seven species, extending the idea of classical Carnegie developmental stages beyond the small set of model organisms for which they were originally defined.
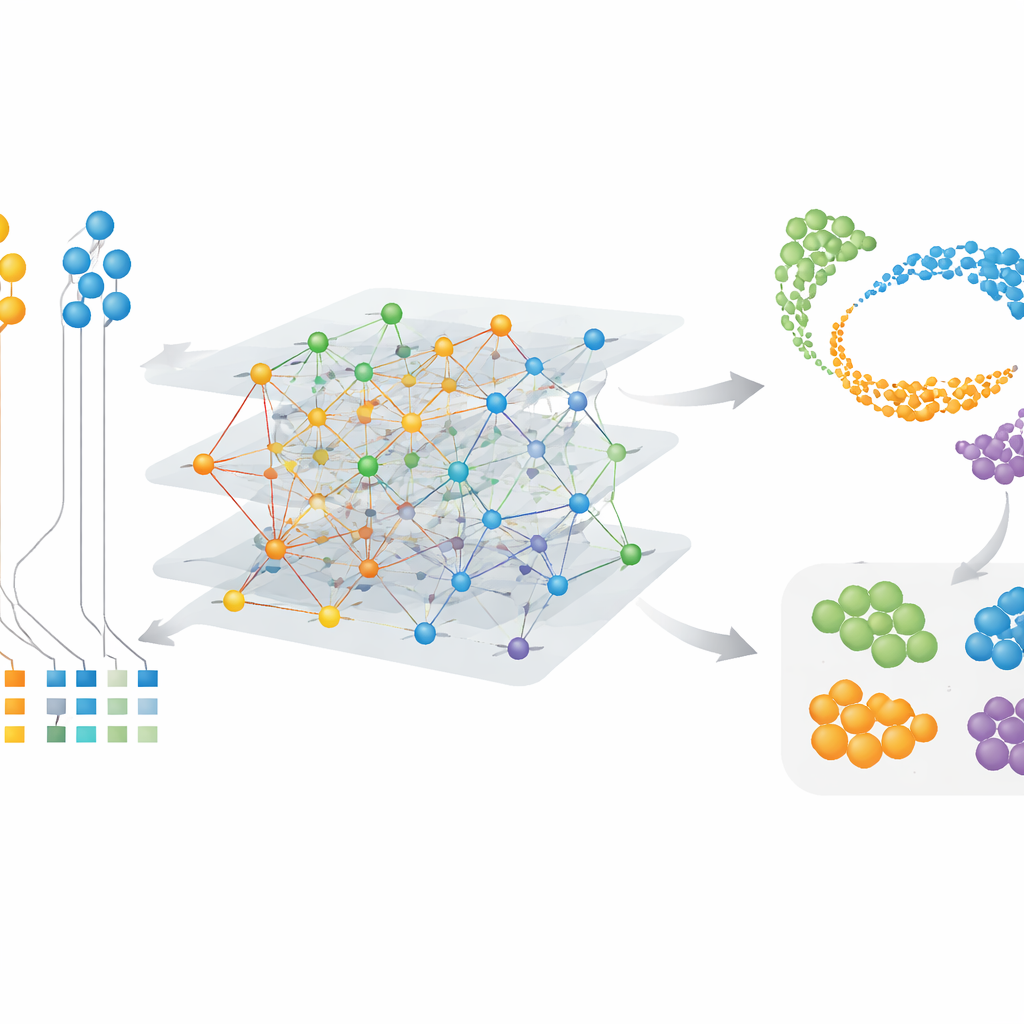
Finding species-specific cells and gene modules
Because CAMEX learns embeddings for both cells and genes, it can highlight special features rather than only shared ones. In brain datasets that included human, mouse, lizard, and turtle, CAMEX integrated the data and, when given human labels as a guide, accurately annotated cell types in the other species, even small subgroups such as brain pericytes in turtle. Applying the method to a detailed map of the primate dorsolateral prefrontal cortex, the authors were able to isolate specific subtypes of microglia—brain immune cells—that are present only in humans or shared with chimpanzees. By clustering the gene embeddings, they also found groups of genes linked to key functions: for example, modules active in somatic support cells in the testis, and others tied to meiosis, the cell division process that produces sperm. These results point to both conserved programs and species-specific tweaks in cell behavior.
What this means for the bigger picture
In plain terms, CAMEX is a powerful new “translation engine” for single-cell data across the tree of life. It helps scientists see when cells from different animals are doing essentially the same job, when they have diverged, and how developmental timelines compare across species. While the method still has limitations—such as relying on existing homology maps and the general challenges of interpreting graph-based models—it already enables richer evolutionary comparisons than were previously possible. Over time, tools like CAMEX could help build a genuine cell type tree of life, sharpen our models of organ development, and guide the search for disease-relevant cell types and drug targets in both humans and animal models.
Citation: Guo, ZH., Huang, DS. & Zhang, S. Multi-species integration, alignment and annotation of single-cell RNA-seq data with CAMEX. Nat Commun 17, 3017 (2026). https://doi.org/10.1038/s41467-026-69696-3
Keywords: single-cell RNA sequencing, cross-species integration, graph neural networks, cell type evolution, comparative genomics