Clear Sky Science · en
Causes and consequences of experimental variation in Nicotiana benthamiana transient expression
Why tiny plant experiments matter
Many breakthroughs in plant science and bio-based manufacturing start with a simple trick: briefly turning plant leaves into miniature test factories. In the species Nicotiana benthamiana, scientists can quickly introduce new DNA and see what happens within days. This speed has made the plant a workhorse for testing genes, building metabolic pathways, and prototyping new biological designs. But like baking the same recipe in different ovens, the results can vary from run to run. This study asks a deceptively simple question with big consequences: how consistent are these fast plant tests, and what can researchers do to tame the noise so they can trust subtle differences in their data?
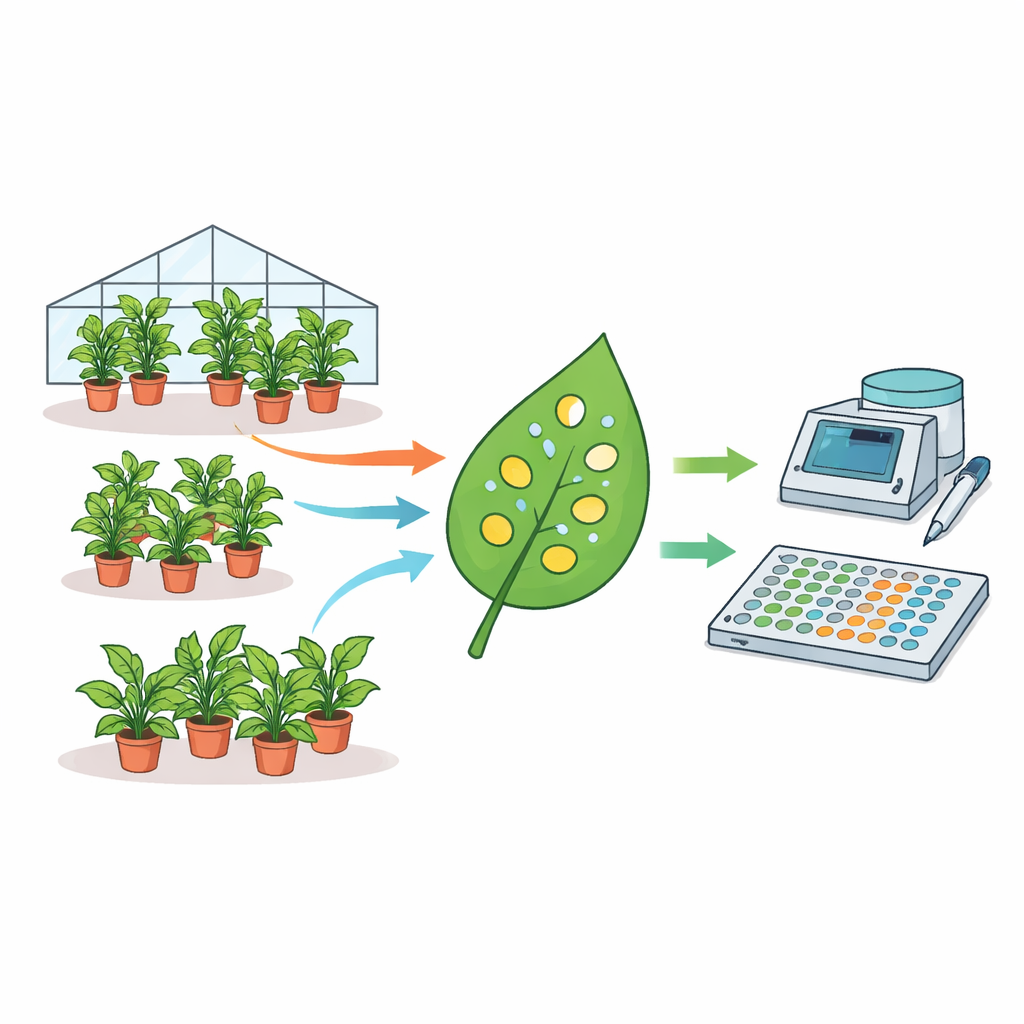
How a favorite test plant can give different answers
The authors assembled an unusually large dataset: measurements from 1915 individual N. benthamiana plants collected over nearly three years. Each plant was infiltrated with Agrobacterium bacteria carrying a fluorescent reporter gene, and tiny discs were punched from leaves and read out on a plate reader. Using statistical models, they broke down where variability comes from. They found that differences between plant batches, between plants within a batch, and even between discs taken from the same leaf all contributed substantially to the overall spread in fluorescence. In fact, mean expression levels could differ by up to fourfold between nominally identical experiments performed on different days. Factors such as plant age and watering volume also influenced signal strength, while time of day for sampling did not.
Testing tricks to even out the noise
To reduce this scatter, many labs use a “normalizer” gene: they deliver two fluorescent reporters at once and analyze the ratio of their signals, hoping that shared sources of noise cancel out. The team systematically compared 17 ways to co-deliver these dual reporters, including mixing separate bacterial strains, stacking both genes on the same DNA segment, and using single strains that carry two plasmids. They showed that all of these strategies can change baseline expression levels, sometimes dramatically, depending on gene orientation and plasmid copy number. Importantly, most—but not all—normalization schemes did reduce variation compared with measuring a single reporter alone, with a simple co-infiltration of two similar plasmids emerging as one of the most effective options.
When helpful corrections backfire
The story turned out to be more nuanced when the team examined promoters—the DNA switches that control how strongly each gene is turned on. They tried combinations of weak, medium, and strong promoters driving the two fluorescent proteins. Normalization worked best when both reporters used the exact same promoter; under those conditions, the ratio showed much smaller scatter. But when promoters differed, the ratio sometimes became more erratic than the raw signal from a single gene, actually worsening statistical power. The authors also found that adjusting the density of the bacterial strains mainly changed how bright the signals were, not how variable they were. Overall, the work shows that normalization is not a one-size-fits-all fix: its success depends heavily on the details of construct design.
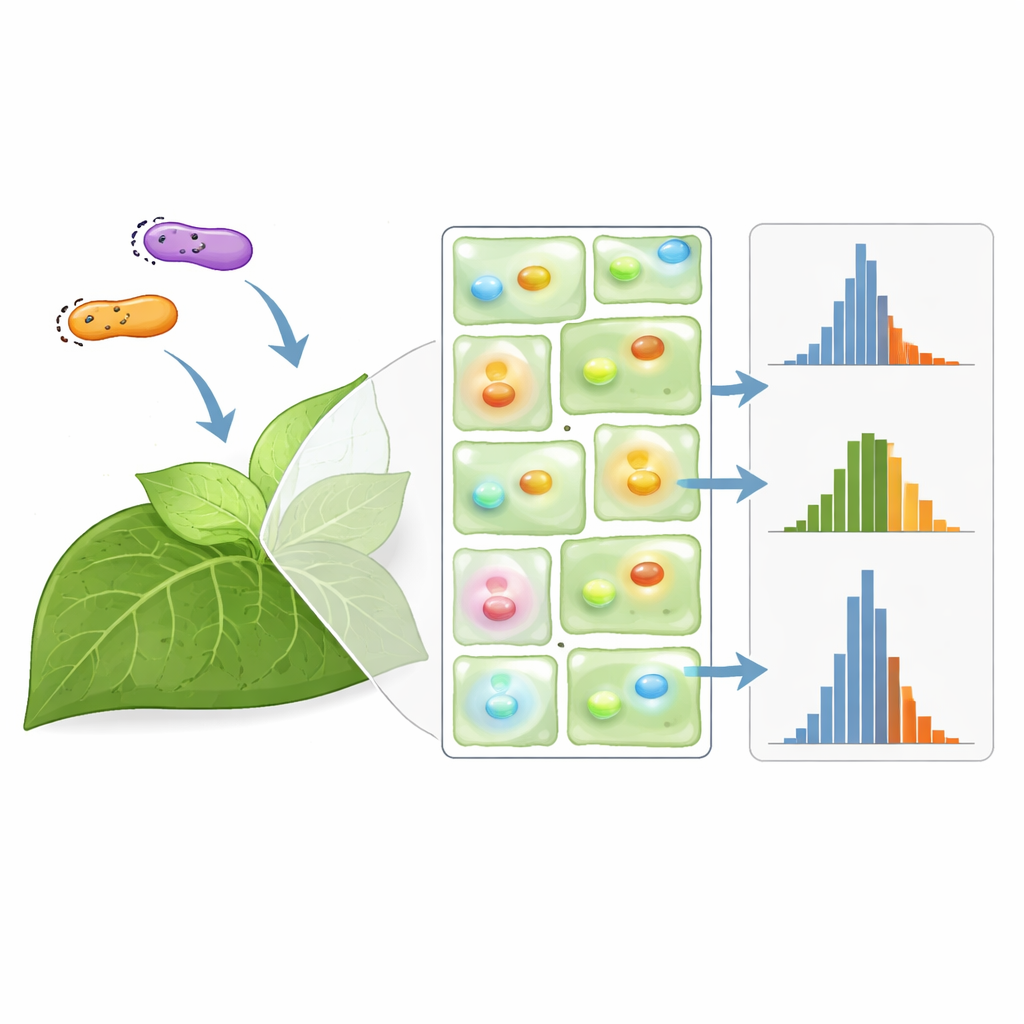
How many plants are enough?
Armed with their large dataset, the researchers built a Monte Carlo simulation to model how variability propagates through typical experiments. They used it to ask a question every experimentalist faces: given a certain level of noise, how many plants do you need to reliably detect a real difference between two constructs? For common Agrobacterium strains, they found that effects larger than about 50 percent can be detected with just a few plants, while teasing apart differences smaller than 20 percent may require dozens, sometimes more than is practical. Carefully chosen normalization can shave these numbers down slightly, particularly when identical promoters are used and only a few plants are available, but in some designs it offers little advantage.
What this means for future plant engineering
For non-specialists, the key message is that even in a well-loved, highly convenient plant system, experimental noise is both substantial and manageable. The study maps out where variation comes from, shows that popular correction methods can help or hurt depending on how they are built, and provides simple power-analysis guidelines for planning experiments. In everyday terms, it tells plant biologists how to size their studies so they do not mistake random fluctuations for real biological effects, and how to design reporter constructs that give more trustworthy comparisons. These insights should make N. benthamiana an even more reliable platform for rapidly testing genes, assembling new metabolic pathways, and advancing synthetic biology toward more predictable engineering of living systems.
Citation: Tang, S.N., Szarzanowicz, M.J., Lanctot, A. et al. Causes and consequences of experimental variation in Nicotiana benthamiana transient expression. Nat Commun 17, 2772 (2026). https://doi.org/10.1038/s41467-026-69458-1
Keywords: Nicotiana benthamiana, transient expression, synthetic biology, experimental variability, Agrobacterium