Clear Sky Science · en
Genetic diagnosis of CYP21A2-related CAH: adaptive sampling long-read sequencing is an accurate and scalable solution
Why this matters for families and doctors
Many babies are now screened at birth for congenital adrenal hyperplasia, a hormonal disorder that can be life‑threatening if missed. Yet confirming the diagnosis at the DNA level is surprisingly hard, because the key gene sits in a confusing stretch of our genome with a nearly identical twin “decoy” gene. This study shows that a new kind of DNA reading, called adaptive sampling long‑read sequencing, can cut through that confusion, giving faster and more reliable answers for patients and their families.
A hormone disorder with genetic twists
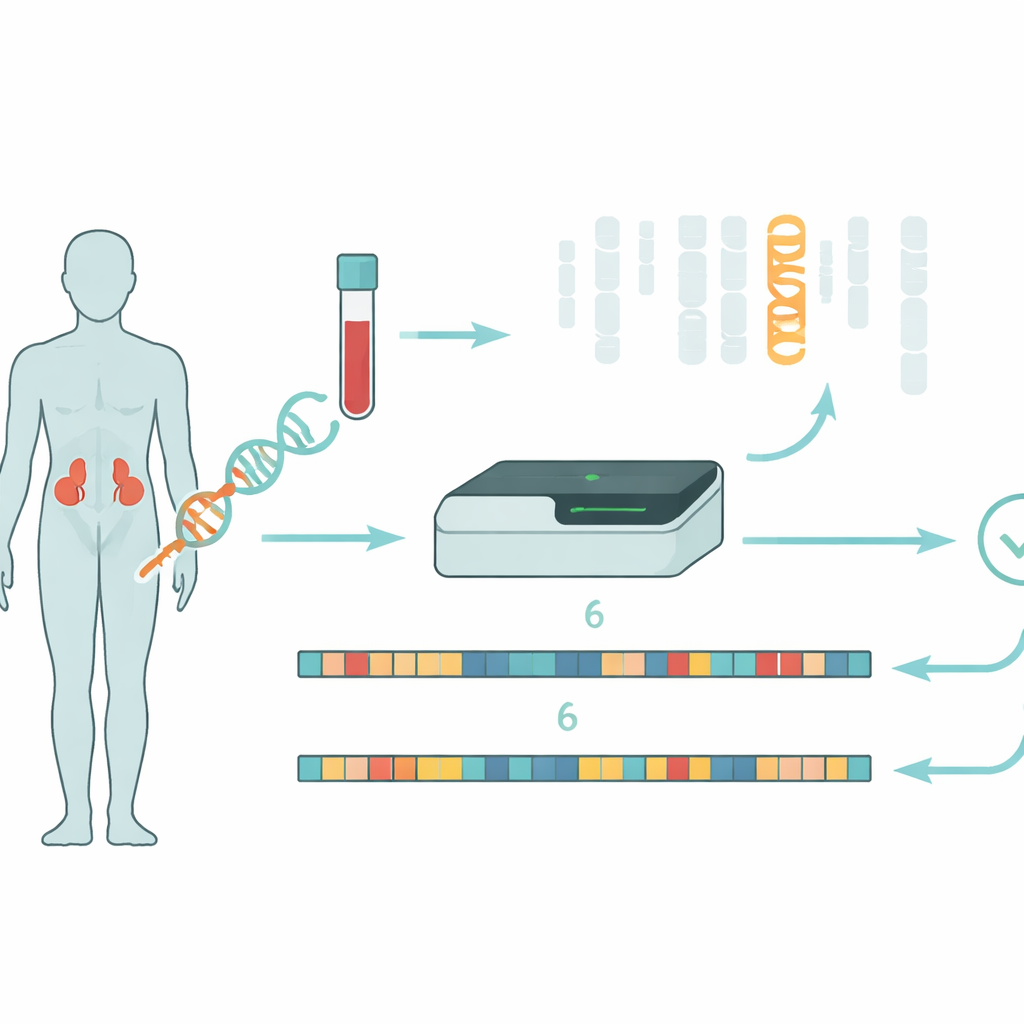
Congenital adrenal hyperplasia (CAH) affects how the adrenal glands make vital hormones that control salt balance, stress responses, and sex hormone levels. In about 95 percent of cases, the problem traces back to changes in a single gene, CYP21A2, which helps make the enzyme 21‑hydroxylase. If that enzyme is missing or weak, the body cannot produce enough cortisol and aldosterone, and instead overproduces androgens, leading to symptoms ranging from salt‑wasting crises in newborns to excess body hair and fertility problems later in life. Most patients carry different disease‑causing changes on each copy of the gene, and the mix of these changes strongly influences how severe their illness will be.
A gene hiding next to its look‑alike
The real challenge is that CYP21A2 lies in a complicated neighborhood on chromosome 6 beside a nearly identical “pseudogene,” CYP21A1P, that no longer works. Their DNA code is about 98 percent the same, and the region is prone to reshuffling events in which pieces of the working gene and the pseudogene swap places, fuse together, or get deleted. Traditional testing methods rely on amplifying specific stretches of DNA by PCR and then reading them with Sanger sequencing, sometimes combined with a technique called MLPA to count gene copies. These methods are slow, labor‑intensive, and can struggle to spot large rearrangements or to tell whether multiple changes sit on the same chromosome or on opposite ones—information that is crucial for understanding disease risk in patients and relatives.
A new way to read the tricky region
The researchers explored a different strategy using long‑read sequencing on the Oxford Nanopore platform. Instead of chopping DNA into very short fragments, this technology reads much longer stretches, which helps connect distant features along the chromosome. They used “adaptive sampling,” where the sequencer is instructed in real time to focus on a chosen region—in this case chromosome 6—boosting coverage without complex lab steps. From 34 patients with a clinical diagnosis of CAH, DNA was sequenced without PCR, avoiding biases that can cause one version of the gene to disappear from view. To handle the extreme similarity between CYP21A2 and its pseudogene, the team built a custom software tool called NanoCAH, which carefully examines how each long DNA read aligns, decides whether it truly comes from the working gene or the decoy, and reconstructs each patient’s two versions of the region.
Clearer answers in most patients
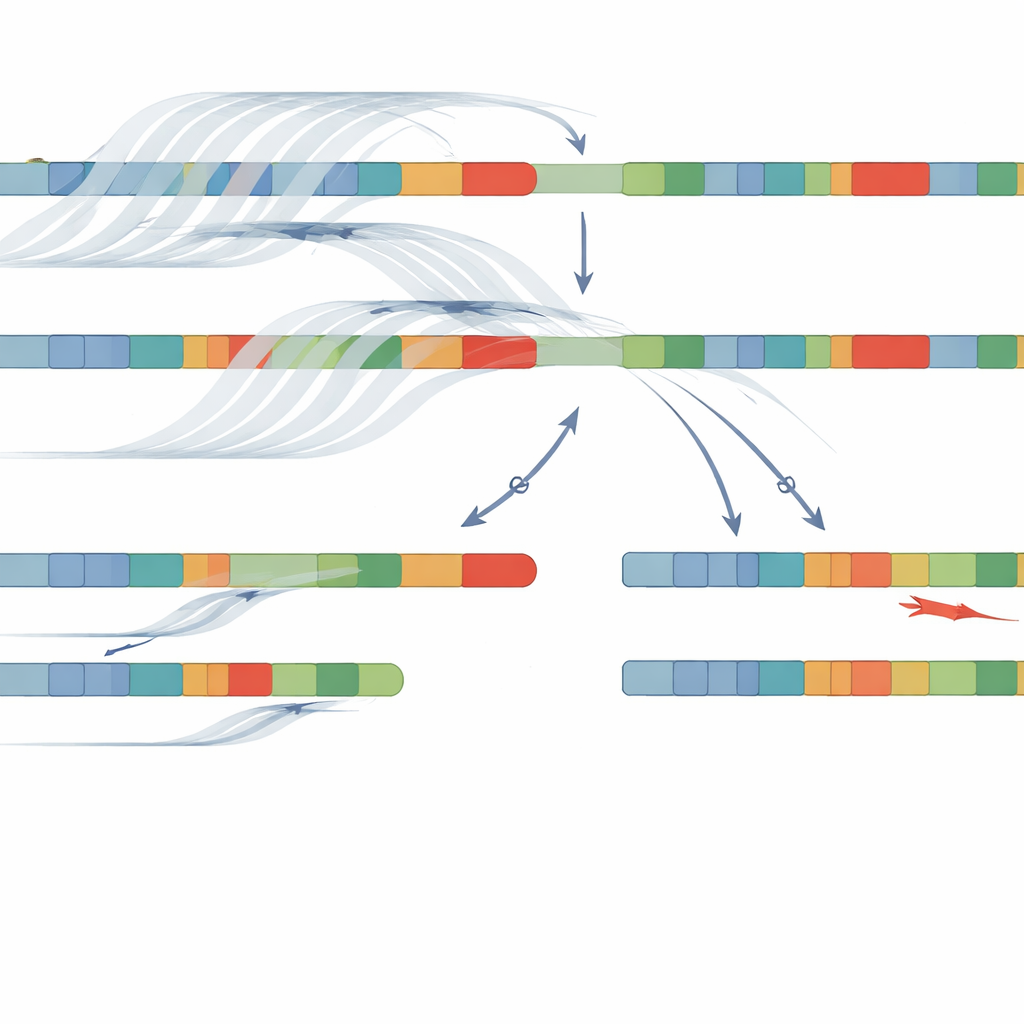
Using this combination of adaptive sampling long‑read sequencing and NanoCAH, the team pinpointed disease‑causing DNA changes in 32 of the 34 patients, a 94 percent success rate. They not only confirmed nearly all findings from older methods but also uncovered important details that had been missed or misclassified. In some patients, earlier tests had called a change “homozygous,” as if both chromosomes carried the same error; the new method revealed that one chromosome actually carried a deletion or a complex fusion between the gene and pseudogene. In others, previously unseen misspellings in the gene were identified because the new approach reads the entire region rather than a fixed set of common hotspots. Crucially, the long reads allowed the team to “phase” variants—showing which changes sit together on the same chromosome—without needing DNA from parents.
What this means for future testing
The study concludes that adaptive sampling long‑read sequencing, paired with specialized analysis software, can deliver faster, more scalable, and more accurate genetic diagnoses for CAH than the current standard tests. By giving a clearer picture of complex gene rearrangements and reliably separating the real gene from its near‑twin, this approach can improve diagnosis, guide treatment decisions, and support more precise carrier and prenatal testing in affected families. Because many other disease‑related genes also lie in tangled, repetitive stretches of our DNA, the same strategy could help untangle difficult genetic questions far beyond CAH.
Citation: Lildballe, D.L., Huno, M.R., Ridder, L.O.R. et al. Genetic diagnosis of CYP21A2-related CAH: adaptive sampling long-read sequencing is an accurate and scalable solution. Eur J Hum Genet 34, 535–542 (2026). https://doi.org/10.1038/s41431-026-02019-8
Keywords: congenital adrenal hyperplasia, CYP21A2, long-read sequencing, genetic diagnosis, adaptive sampling