Clear Sky Science · en
Targeted long-read sequencing for high-resolution repeat profiling in myotonic dystrophy type 1
Why this matters for families and doctors
Some inherited diseases are driven by stuttering stretches in our DNA—short sequences repeated hundreds or even thousands of times. When these repeats grow too long, they can cause serious conditions such as myotonic dystrophy, a muscle-wasting disease. Yet exactly measuring how long these repeats are, and how they are chemically marked, is surprisingly hard with today’s hospital tests. This paper introduces a streamlined way to read these difficult DNA regions in detail within about a day, potentially giving doctors clearer answers and patients better guidance.
A closer look at a tricky muscle disease
Myotonic dystrophy type 1 (DM1) is caused by an overgrown run of three DNA letters in a gene important for muscle and other tissues. People with more repeats tend to have more severe disease, so knowing the exact repeat length is vital for diagnosis and prognosis. Standard tools in clinical labs—such as special PCR tests and Southern blots—work well when the repeat is modest in size, but they often fail or become very imprecise once expansions exceed a few hundred copies. That leaves a gray zone where families may know that a mutation is present but lack a clear picture of how extensive it is or how it might evolve over time.
Combining molecular scissors and long-read DNA reading
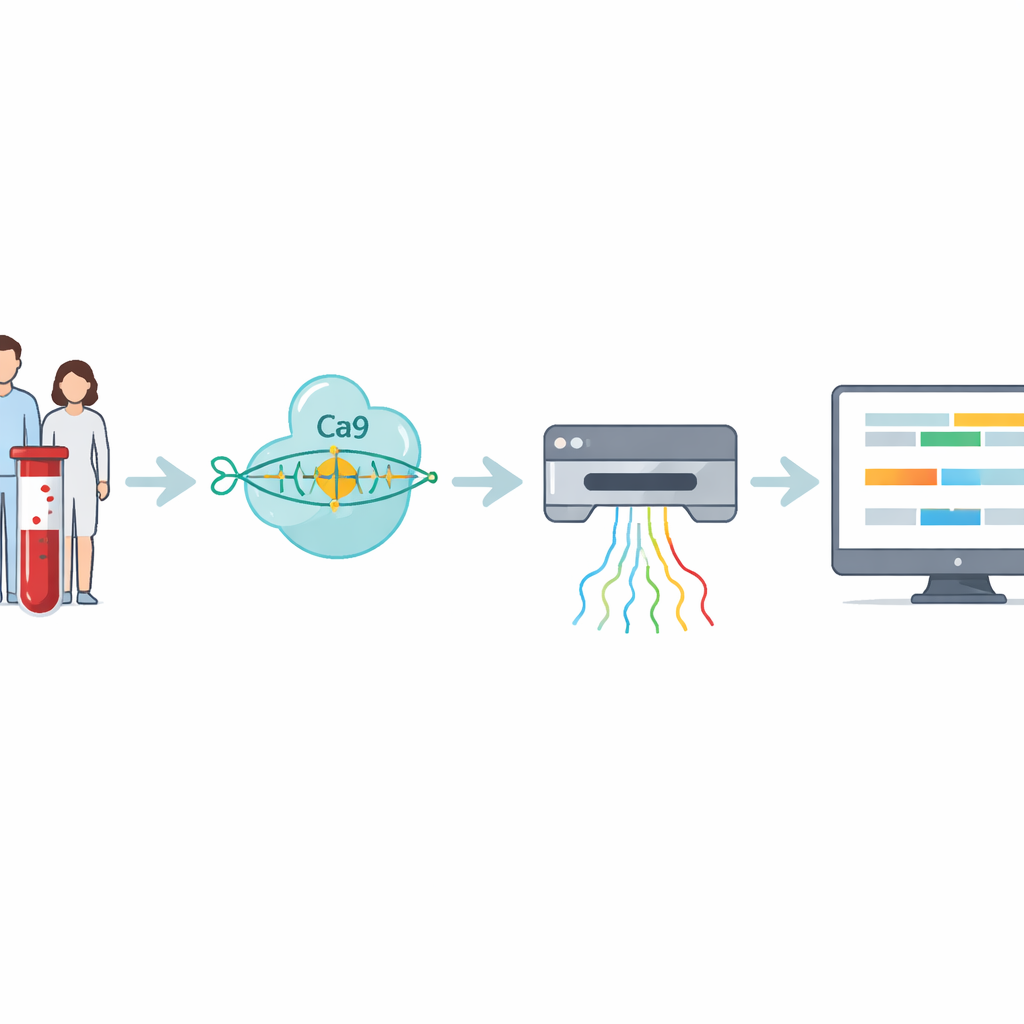
The researchers built a four-step workflow that tackles these hard-to-read repeats directly. First, DNA is extracted from blood or cell samples. Next, the team uses CRISPR–Cas9—molecular scissors guided to a specific address in the genome—to cut around the stretch of DNA containing the repeat. Because only those cut pieces are prepared for sequencing, the method strongly enriches for the region of interest while avoiding copying the DNA by PCR, which tends to miss very large expansions. The prepared DNA is then fed into a nanopore sequencer, a handheld device that threads DNA strands through tiny pores and senses their composition in real time as electrical signals.
Smart software to read length and chemical marks
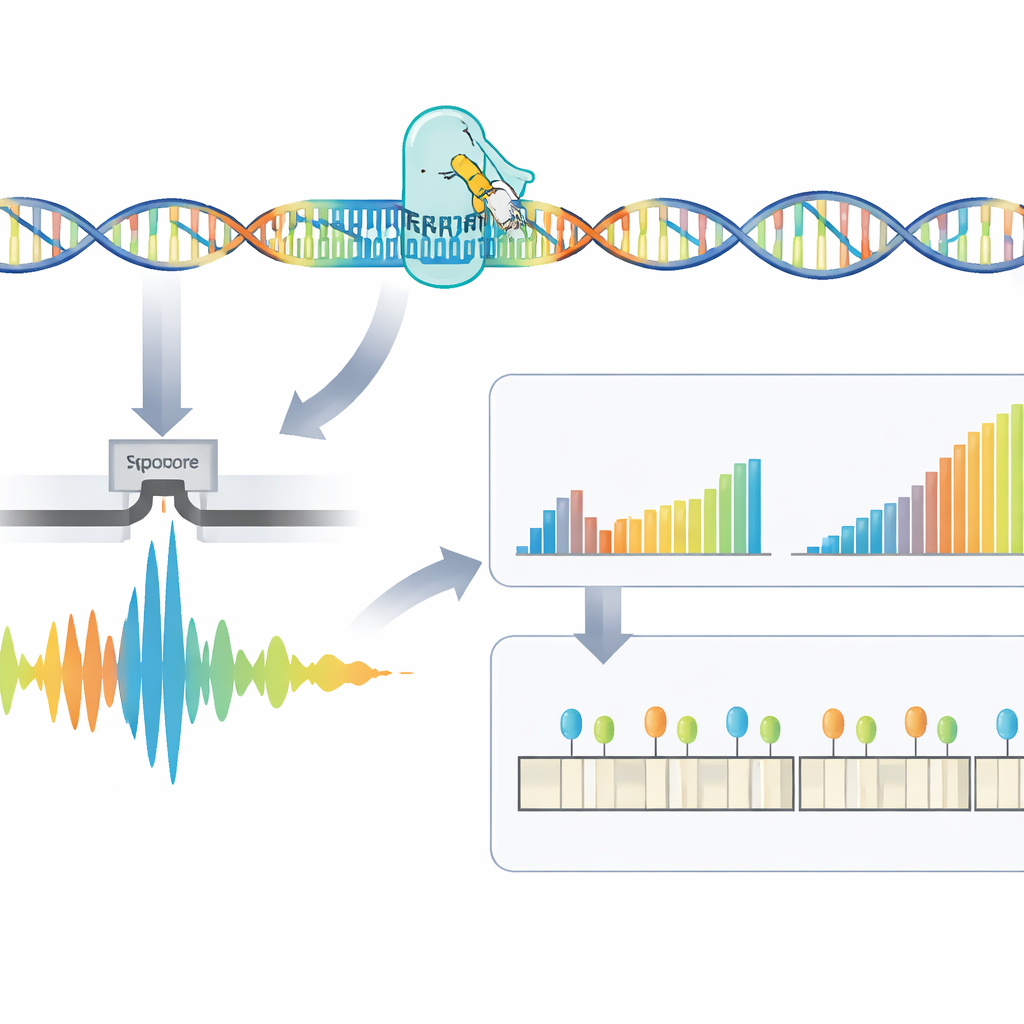
Raw signals from the sequencer must be translated into DNA code and then into meaningful measurements. Here the authors introduce RepeatLab, an automated analysis pipeline designed to be used even on an ordinary cloud notebook. RepeatLab first performs a quick pass to find reads that cover the targeted gene, then reprocesses those reads with a more demanding, high-accuracy setting tailored for long, repetitive stretches. It uses a modified statistical strategy to cluster single-molecule measurements into two groups—typically the normal and expanded copies of the gene in each person—achieving reliable length estimates with as few as about a dozen reads. The same framework can analyze many samples or multiple genes in one run, keeping costs comparable to existing clinical assays.
Peering into interruptions and epigenetic signals
Beyond simply counting repeats, the method examines the fine structure of the repeated region and how it is chemically decorated. Some patients carry interruptions—small patches of different sequence within the repeat—that may be linked to milder disease. The team found that many apparent interruptions were actually artifacts of how the raw signal was processed, and they showed that using larger analysis windows during decoding greatly reduced these false positives. RepeatLab also uses a specialized mode to read DNA methylation, a chemical tag that can switch genes on or off. Around the DM1 gene, the authors mapped methylation patterns at single-allele resolution and identified several key zones where heavier methylation tracks with longer repeat expansions, supporting the idea that these chemical marks help shape how the disease unfolds.
A faster, richer test for hard-to-measure repeats
Taken together, this work shows that CRISPR-guided enrichment, nanopore long-read sequencing, and user-friendly software can deliver a detailed profile of disease-causing DNA repeats in under 24 hours. For DM1 and related conditions, the approach not only measures how long the repeats are but also captures subtle sequence changes and epigenetic patterns that current hospital tests largely miss. While further validation and adaptation to newer sequencing chemistries are still needed, this integrated platform points toward future genetic tests that are faster, more informative, and easier to deploy in routine care.
Citation: Han, Y., Jang, JH. & Chang, H. Targeted long-read sequencing for high-resolution repeat profiling in myotonic dystrophy type 1. Exp Mol Med 58, 1203–1215 (2026). https://doi.org/10.1038/s12276-026-01683-6
Keywords: myotonic dystrophy, tandem repeat expansion, nanopore sequencing, CRISPR Cas9 enrichment, DNA methylation