Clear Sky Science · en
Prospective multicentre study of upper respiratory mucosal transcriptomics reveals two major endotypes of critically ill COVID-19 patients
Why this matters for patients and families
Even after years of living with COVID-19, doctors still struggle to explain why some people with the Omicron variant become critically ill while others do not. This study zooms in on the lining of the nose and upper airways in patients sick enough to need intensive care, asking a simple but important question: are there different “types” of immune reactions in the airways, and could those invisible patterns one day guide more personalized treatments?
Looking inside the nose, not just the blood
Most hospital tests for severe COVID-19 focus on blood samples, scans, and oxygen levels. But the virus first settles and multiplies in the nose and throat, where the immune system launches its earliest defense. In this multicenter French study, researchers collected standard nose swabs from 94 adults admitted to intensive care units with life-threatening Omicron-related pneumonia between spring 2022 and summer 2023. From these swabs, they extracted and sequenced human genetic messages, known as RNA, to see which immune-related genes were turned on or off in the airway lining. By using a mathematical clustering method focused on cell–signaling molecules called cytokines, they grouped patients purely on the basis of these airway immune signatures.
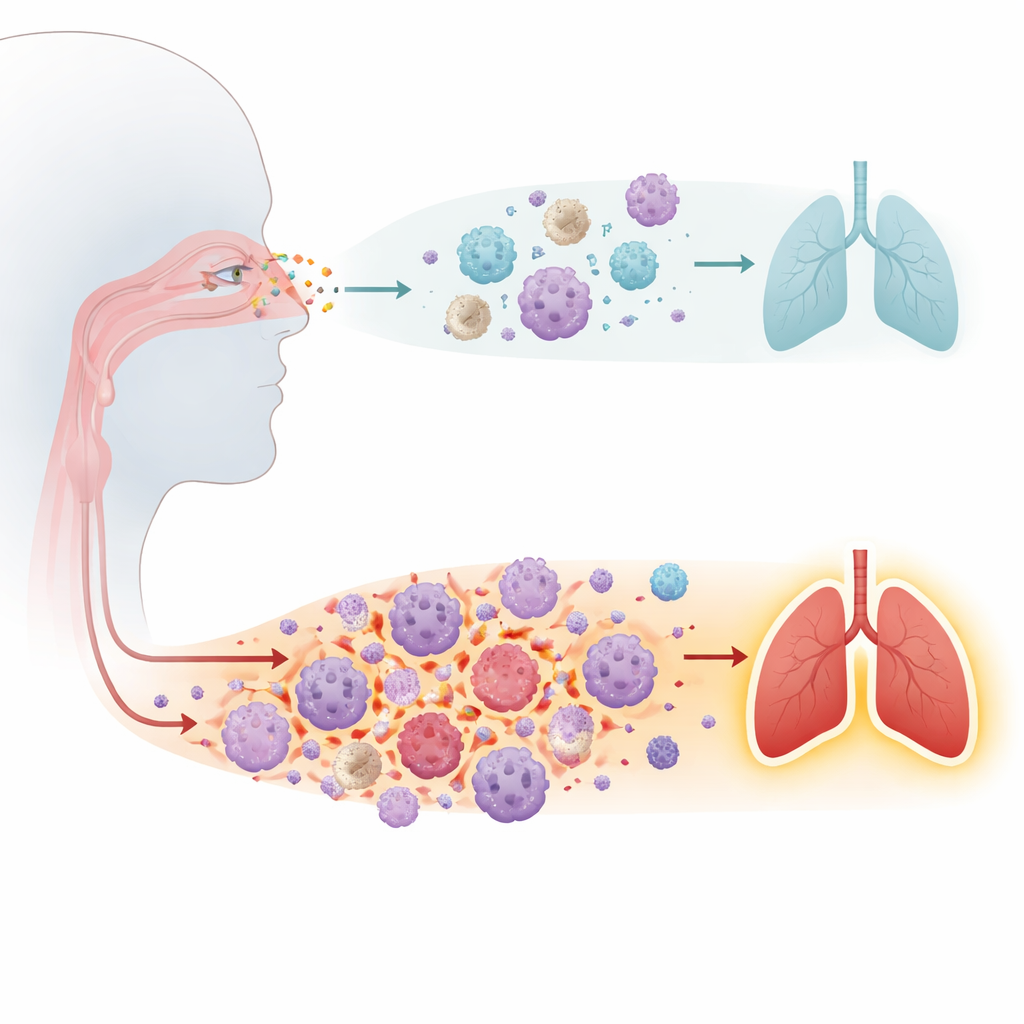
Two hidden immune “types” in the airways
Among 56 patients with high-quality data, the analysis revealed two clear immune patterns in the upper airways, which the team called COVID-19 Immune Transcriptomic Respiratory Profiles, or CITRP-1 and CITRP-2. Both groups were similar in age, underlying illnesses, vaccination status, virus levels, and overall severity when they arrived in the ICU. Yet their airway immune activity looked very different. Patients in the CITRP-2 group showed a much stronger inflammatory response, with higher activity of genes linked to early, fast-acting defenses and to certain helper T cells that drive allergic- and asthma-like inflammation. In contrast, patients in the CITRP-1 group had a more muted, less inflamed airway profile, despite being just as sick clinically.
When defenders become damaging
The overactive pattern in CITRP-2 centered on a type of white blood cell called the neutrophil, which rushes to infection sites and releases toxic granules and web-like traps to kill microbes. Gene pathways tied to neutrophil degranulation, engulfing of germs, and sensors that detect viral components were all more active in this group. So were signals associated with so-called Th2 responses, involving molecules similar to those seen in allergies and asthma. One standout signal, produced at especially high levels, was a chemokine that powerfully attracts neutrophils to the site of infection. Using computational tools, the authors estimated that CITRP-2 patients had more neutrophils and fewer virus-killing T cells in their nasal tissue than CITRP-1 patients, suggesting a shift toward a hot, neutrophil-heavy front line that may injure delicate airway structures while trying to clear the virus.
Same bedside picture, different biology
Surprisingly, these striking immune differences did not translate into obvious differences at the bedside. Rates of mechanical ventilation, organ support, complications such as ventilator-associated pneumonia, length of stay in intensive care, and death at 28 days were similar between the two endotypes. The only routine lab value that clearly differed was a lower blood lymphocyte count in the more inflamed CITRP-2 group. The authors note that because all patients in this study were already critically ill, it may be harder to see how immune profiles relate to outcomes than if patients with milder disease were included. Still, the fact that two biologically distinct groups look almost identical using standard clinical tools underscores how much important information is currently hidden from routine practice.
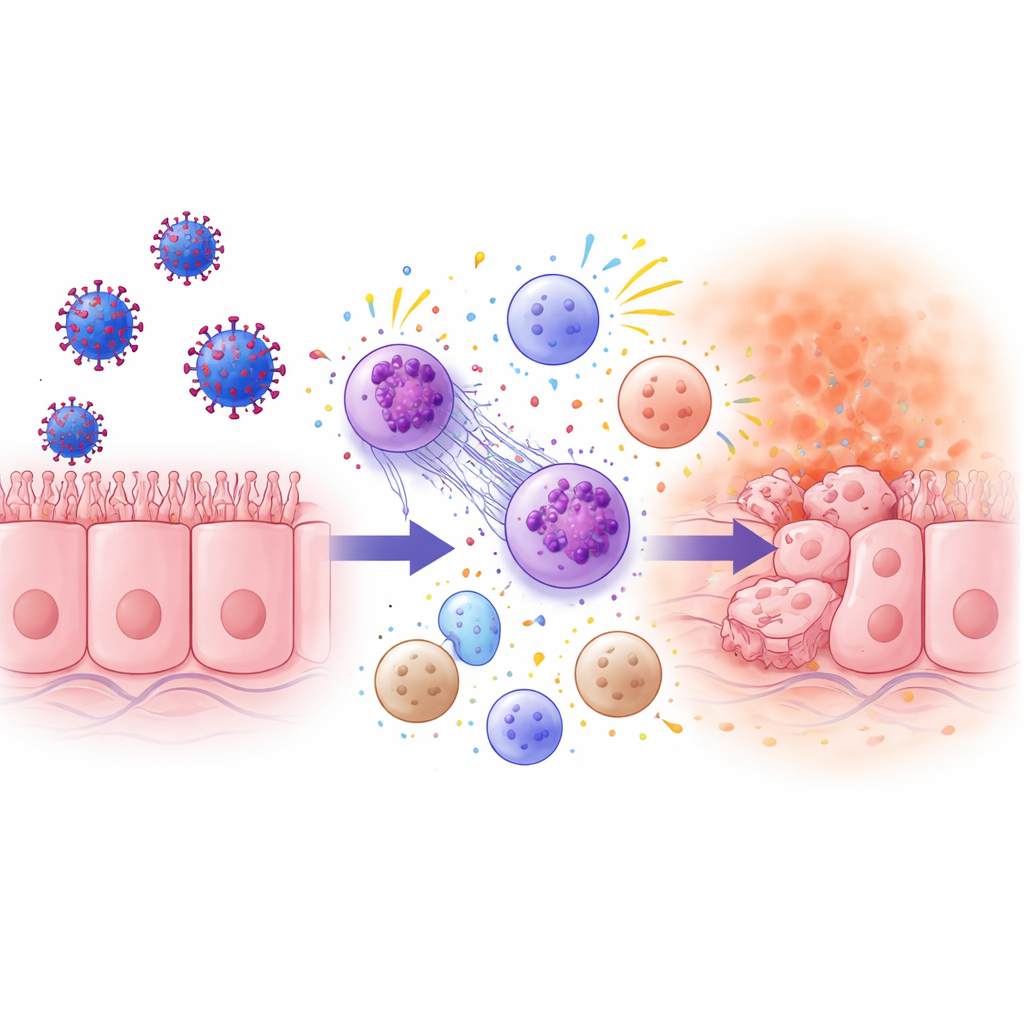
Toward more tailored treatment for severe COVID-19
The study’s main message is that severe Omicron COVID-19 in the ICU is not a single disease, even when patients look very similar. Instead, at least two distinct airway immune patterns exist: one comparatively restrained, and one dominated by intense neutrophil- and Th2-driven inflammation that may itself contribute to lung damage. Today, most critically ill patients receive the same set of anti-inflammatory drugs. The authors argue that future care should move toward matching therapies to these hidden immune endotypes—such as selectively dampening neutrophil activity or specific cytokines in the most inflamed subgroup. Although more and larger studies are needed, especially including people with less severe illness, this work shows that a simple nose swab could one day help doctors choose the right immune-targeting treatment for the right patient at the right time.
Citation: Bay, P., Boizeau, L., Préau, S. et al. Prospective multicentre study of upper respiratory mucosal transcriptomics reveals two major endotypes of critically ill COVID-19 patients. Commun Med 6, 238 (2026). https://doi.org/10.1038/s43856-026-01474-0
Keywords: COVID-19, Omicron variant, immune response, neutrophils, personalized medicine