Clear Sky Science · en
Machine and Deep Learning Reveal Sequence Determinants Encoding Bivalent Histone Modifications
How DNA’s Punctuation Marks Shape a Cell’s Future
Every cell in your body carries essentially the same DNA, yet brain cells and muscle cells behave worlds apart. One reason is that chemical tags on DNA-packaging proteins can flip genes on or off without changing the genetic code itself. This study asks a surprisingly simple question with big implications: are there hidden patterns in the DNA sequence that tell the cell where to place a special kind of “mixed” tag that keeps crucial genes poised between silence and activity?
A Tale of Two Opposing Tags
Inside the nucleus, DNA is wrapped around protein spools called histones. These histones can carry signals that either encourage gene activity (“go”) or suppress it (“stop”). Sometimes, both types of signals sit together in the same spot, creating what scientists call a “bivalent” state—genes are held in a ready-but-waiting mode. Using mouse embryonic stem cells, which can become almost any tissue, the researchers mapped three key histone tags across the genome. They found that regions carrying mixed tags differed from single-tag regions: they were slightly narrower, richer in the DNA letters G and C, and more strongly conserved across evolution, hinting that these poised stretches of DNA are especially important and carefully protected.
Poised Switches for Development and Disease
When the team linked these tagged regions to nearby genes, a pattern emerged. Genes marked by mixed histone signals tended to be turned on only modestly and were heavily involved in early development and in the decision of stem cells to remain flexible or specialize. Pathways such as Hippo, MAPK, Wnt, and TGF-beta—core communication circuits for growth and tissue formation—were strongly represented. Some bivalently marked genes have also been tied to cancers, suggesting that the same poised control system that guides healthy development can be hijacked in disease. Overall, mixed marks appear to work like finely tuned dimmer switches, giving genes a subtle baseline of activity while keeping them ready to ramp up or shut down when signals arrive.
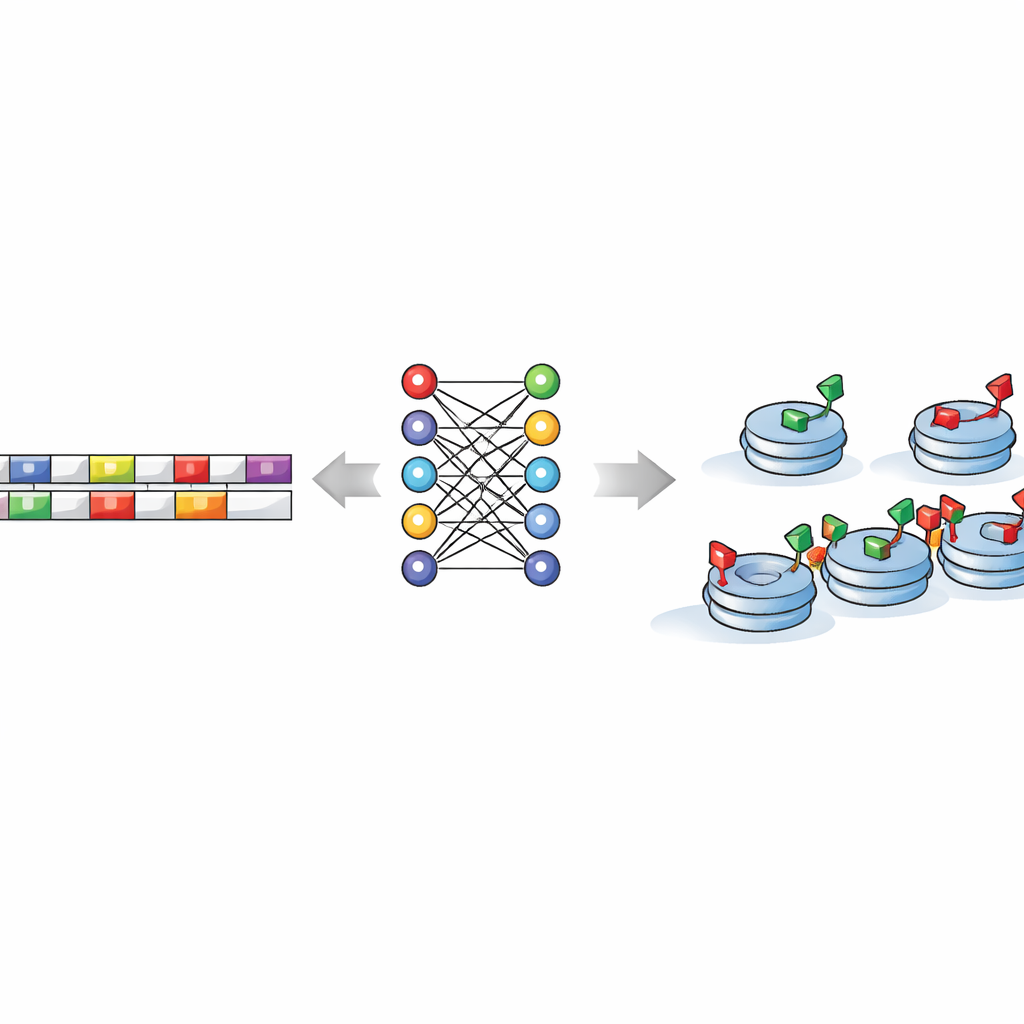
Teaching Machines to Read Hidden DNA Patterns
The heart of the study asks whether the DNA sequence itself encodes instructions for where these poised states should form. To test this, the researchers fed short stretches of DNA—broken down into all possible tiny “words” of a few letters—into a set of machine learning and deep learning models. These algorithms learned to distinguish regions with mixed tags from those with only activating or only repressive tags, often with high accuracy. Crucially, when the DNA letters were shuffled at random, the models failed, showing that the real genome carries authentic predictive signals rather than accidental noise. This means that without looking at any experimental measurements, a computer can use the DNA text alone to guess where the cell is likely to place these mixed histone marks.
Sequence Motifs as Molecular Road Signs
By peering inside the models, the authors uncovered a handful of short DNA motifs—recurring letter patterns—that were especially informative. Some, like sequences resembling TCTGAA and TCACAG, matched known binding sites of master stem cell regulators such as OCT4, SOX2, ESRRB, and a factor called TCFCP2l1. Others tended to cluster near the edges of bivalently marked regions, hinting that certain motifs may help set the boundaries of these poised chromatin zones. Different combinations and placements of motifs distinguished one type of mixed marking from another, implying that each class of bivalency follows its own “grammar” of sequence rules even while sharing many of the same regulatory proteins.
What This Means for Stem Cells and Beyond
Put simply, the study shows that DNA is not just a list of genes; it also carries embedded instructions about how tightly those genes should be packaged and how ready they are to respond. In embryonic stem cells, specific short DNA patterns help recruit protein factors and shape regions where opposing histone tags coexist, keeping developmental genes balanced on a knife edge between on and off. By harnessing machine learning and deep learning to read this hidden code, the authors provide both a practical tool for predicting epigenetic states from sequence and a clearer picture of how cells program flexibility into their genomes during early life—and how that programming might go awry in disease.
Citation: Zhao, X., Wu, J., Che, Y. et al. Machine and Deep Learning Reveal Sequence Determinants Encoding Bivalent Histone Modifications. Commun Biol 9, 491 (2026). https://doi.org/10.1038/s42003-026-09962-8
Keywords: bivalent chromatin, histone modifications, embryonic stem cells, DNA sequence motifs, machine learning in genomics