Clear Sky Science · en
Smoothie: efficient inference and integration of spatial co-expression networks from denoised spatial transcriptomics data
Seeing Cellular Neighborhoods in New Detail
Every tissue in the body is a bustling city of cells, each turning genes on and off in precise locations. New "spatial transcriptomics" technologies can now read which genes are active at almost every point in a tissue slice, but the raw data are extremely noisy and incomplete. This paper introduces Smoothie, a computational method that cleans up those blurry measurements and reveals groups of genes that act together in space, helping scientists decode how tissues are built, how they function, and how they change over time or disease.
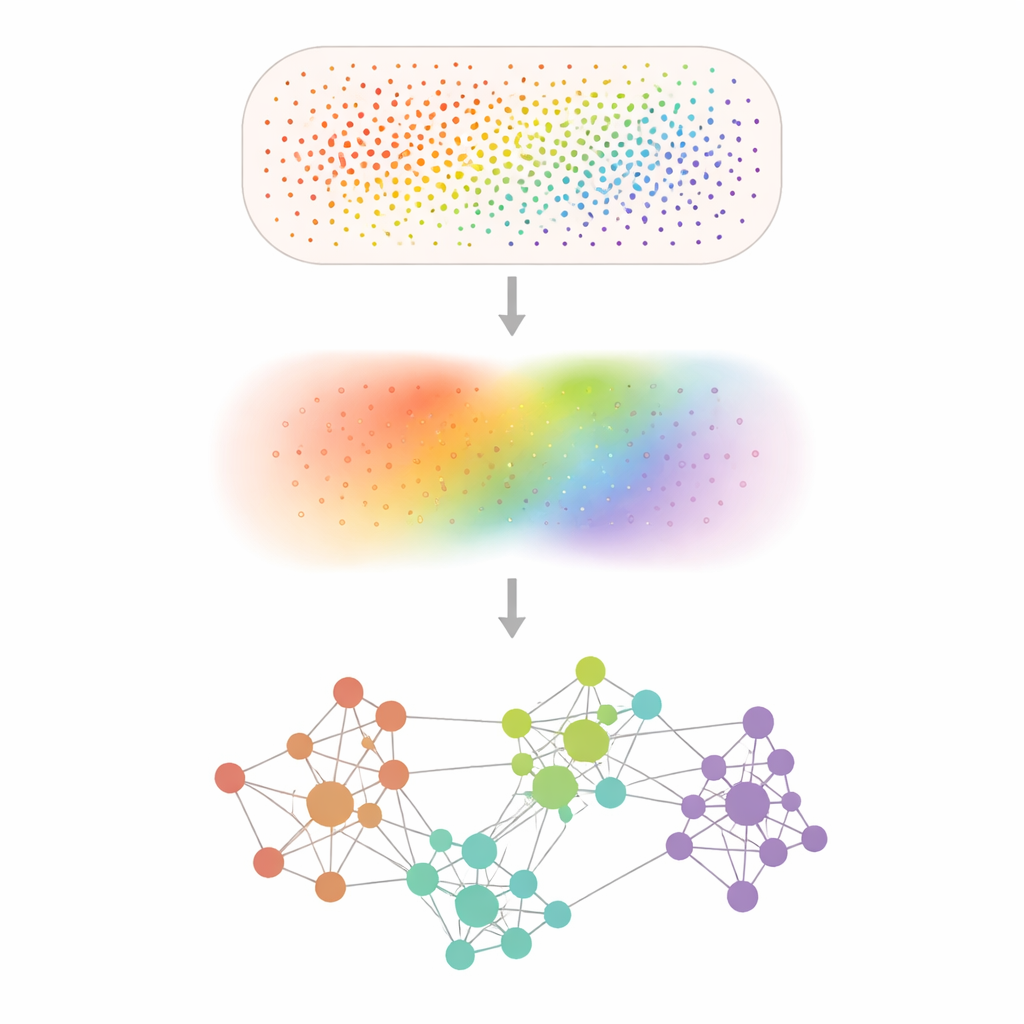
Turning Noisy Dots into Clear Patterns
Spatial transcriptomics experiments record gene activity at tens of thousands to hundreds of millions of tiny positions, but many genes are barely detected and measurements are speckled by random noise. Smoothie tackles this by gently blurring the data with a mathematical "smoothing" step that averages each measurement with its neighbors, much like applying a soft-focus filter to a pixelated photograph. After this denoising, Smoothie compares every pair of genes to see where their smoothed patterns rise and fall together across the tissue. Pairs that are strongly in sync are connected in a network, and clusters within this network reveal groups of genes that share a common spatial pattern and likely a common biological role.
Reading the Gene Map of the Brain and Embryo
The authors first tested Smoothie on a high‑resolution map of gene activity in the adult mouse cerebellum, a part of the brain involved in balance and coordination. Smoothie uncovered distinct gene groups that neatly matched known cell types, including Purkinje neurons, granule cells, oligodendrocytes, and support cells. By comparing results with three leading methods, the team showed that Smoothie finds more gene groups, includes more genes overall, and forms tighter, better‑separated clusters, all while running much faster. Crucially, the smoothing step made real biological relationships easier to detect by boosting correlations between known marker genes that share a cell type.
Discovering Hidden Cell Types and Unknown Genes
Next, the researchers pushed Smoothie to a demanding test: a massive spatial dataset from a mid‑gestation mouse embryo measured at sub‑micron resolution. Without resorting to heavy preprocessing, Smoothie analyzed over 175 million spatial points and more than 20,000 genes in about an hour. It revealed hundreds of gene modules that corresponded to organs, sub‑regions within organs, and specialized cell types in the liver, kidney, brain, and skin. Because so many genes could be included, Smoothie also attached likely functions to dozens of poorly characterized genes by placing them into modules with known markers. Intriguingly, some modules consisted of genes that sit next to each other along the genome, hinting that local DNA organization helps shape which genes are active together in space.
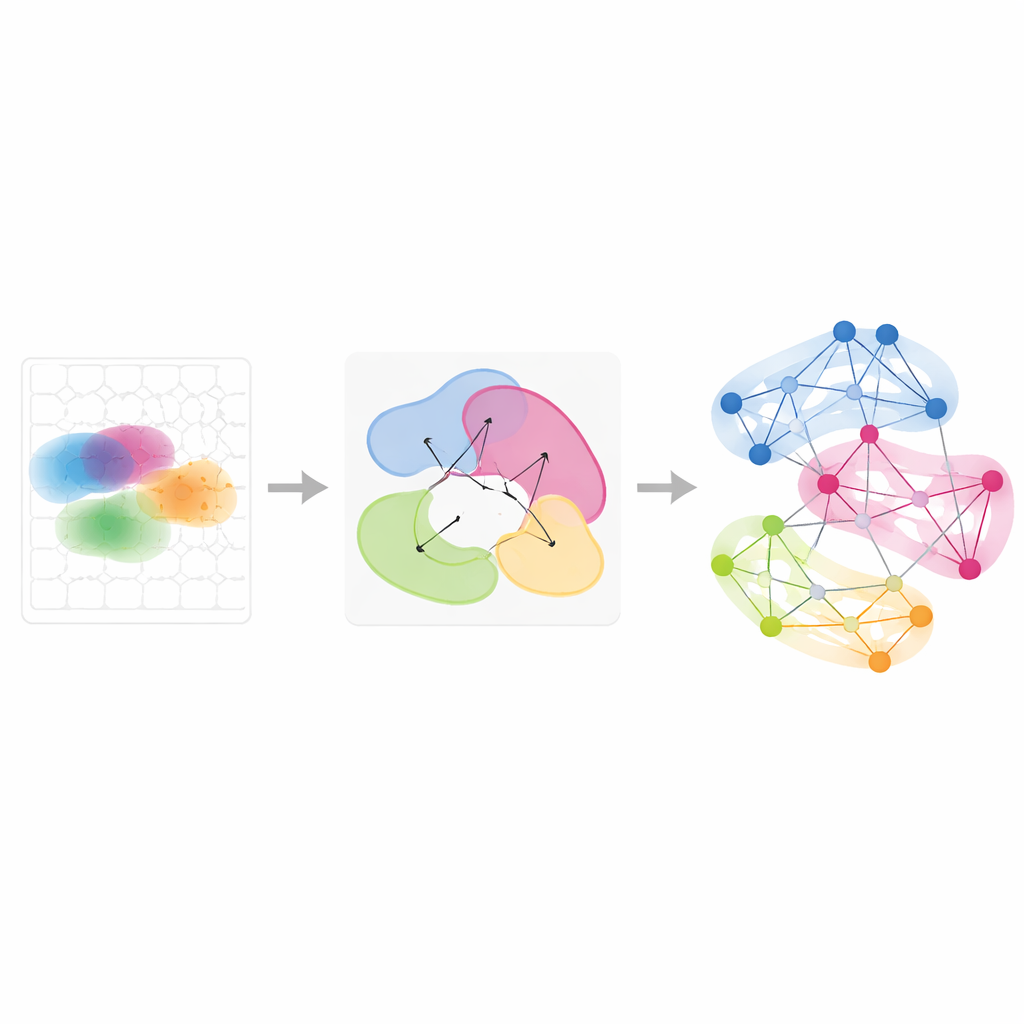
Following Gene Programs Across Time and Conditions
Beyond single tissues, Smoothie can integrate multiple spatial experiments to see how gene patterns are shared or change. The authors applied this to a series of mouse embryos from slightly different developmental stages and to ovaries sampled at eight time points during hormone‑induced ovulation. By comparing how a gene’s partners in the network shift between samples, Smoothie ranks genes as stable or dynamic. This revealed, for example, gene groups that move from liver to blood vessels as red blood cells migrate during development, and wave‑like gene programs in ovarian support cells that switch on in sequence to drive ovulation. These analyses show that gene networks, rather than raw images, provide a powerful common language for aligning mismatched tissues and time points.
A New Lens for Spatial Biology
In practical terms, Smoothie turns vast, noisy spatial gene maps into clear, interpretable networks that highlight where and how genes work together in real tissues. For non‑experts, the key message is that this method makes it far easier to spot meaningful patterns, connect unknown genes to known cell types, and track how tissues change over time or treatment. As spatial genomics technologies continue to sharpen, tools like Smoothie will help transform millions of tiny molecular measurements into coherent stories about development, health, and disease.
Citation: Holdener, C., De Vlaminck, I. Smoothie: efficient inference and integration of spatial co-expression networks from denoised spatial transcriptomics data. Commun Biol 9, 459 (2026). https://doi.org/10.1038/s42003-026-09898-z
Keywords: spatial transcriptomics, gene co-expression networks, tissue architecture, developmental biology, computational genomics