Clear Sky Science · en
Genome sequence assembly of the 5S rDNA loci informs haplotype specificity and evolution in the greater duckweed Spirodela polyrhiza
Why tiny floating plants matter
Greater duckweed is a small aquatic plant that spreads quickly across ponds and canals, but inside its cells lies a surprisingly streamlined genome. This study dives into one of the most mysterious parts of that genome: the stretches of DNA that make the core machinery for building proteins, the cell’s ribosomes. By completely decoding these hard-to-read regions in duckweed, the authors show how essential genes are organized and fine-tuned, offering clues to how plant genomes evolve and how cells control one of their most basic life-support systems.
Building blocks of the cell’s protein factories
Ribosomes, the molecular machines that assemble proteins, are built from both proteins and special RNA molecules. Some of these RNAs are encoded by the so‑called 5S ribosomal DNA (5S rDNA), which normally appears in hundreds or thousands of repeated copies in plant genomes. Because these repeats are long and nearly identical, they are notoriously difficult to assemble using standard DNA sequencing. In greater duckweed (Spirodela polyrhiza), however, earlier work showed that the number of 5S copies is unusually low, making this plant an ideal model to finally map an entire 5S rDNA region from end to end and see how its copies are arranged on chromosomes.
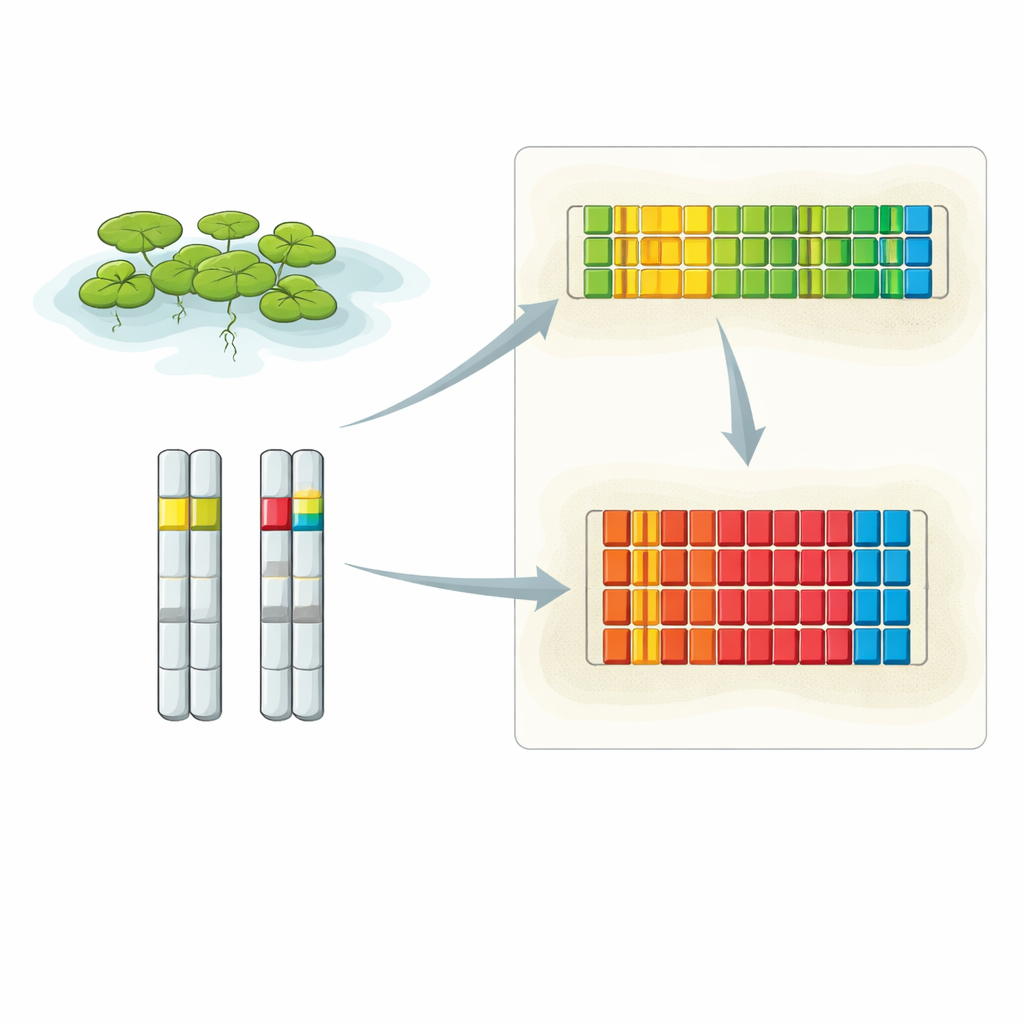
Zooming in on two key DNA neighborhoods
The researchers combined several advanced techniques to uncover the full layout of the duckweed 5S rDNA. They used extra-long DNA reads from Oxford Nanopore sequencing, conventional high-accuracy sequencing of cloned fragments, and high-resolution fluorescence in situ hybridization (FISH), which lights up specific DNA stretches inside intact nuclei. Their analyses revealed that 5S rDNA is concentrated in two main locations, each on a different chromosome. One locus sits on chromosome 6 and contains a series of short repeating units, whereas the other lies on chromosome 13 and is built from longer repeat units. FISH images showed that the signal strength from these loci differs between the pair of chromosome copies in each cell, hinting that one homolog can carry noticeably more 5S repeats than the other.
Unequal repeats and mirrored clusters
By carefully stitching together long sequencing reads, the team assembled nearly complete DNA sequences for both loci. On one version of chromosome 6, they reconstructed a continuous stretch of 40 5S repeat units; the partner chromosome carries more than 65 units of the same short type. On chromosome 13, they traced a more complex landscape: a major cluster of at least 64 long repeats and a smaller cluster of two repeats positioned in the opposite orientation, separated by over 12,000 base pairs of ordinary chromosomal DNA. Within the long-repeat locus, some units carry a tiny 13-base insertion in the spacer region between genes, and these slightly different units tend to group together into sub-clusters. Overall, the two loci together account for roughly 320–390 5S gene copies per diploid genome, consistent with independent estimates of copy number.
DNA chemistry and control switches
When the authors examined the chemical makeup of these regions, a striking pattern emerged. The 5S repeat arrays themselves are unusually rich in the DNA bases G and C, while the flanking chromosome segments around them are strongly enriched in A and T. These AT-rich bordering zones resemble DNA elements associated with replication origins and open, active chromatin in other organisms, and similar motifs appear scattered across all 20 duckweed chromosomes. At the finer scale of individual repeat units, the team identified small but consistent differences in short control sequences upstream and downstream of the 5S gene, including variations in TATA-like motifs and stretches of GA repeats. These are known in other species to interact with general transcription factors and GAGA-binding proteins, suggesting that each locus may be tuned differently even though the 5S RNA they produce is identical.
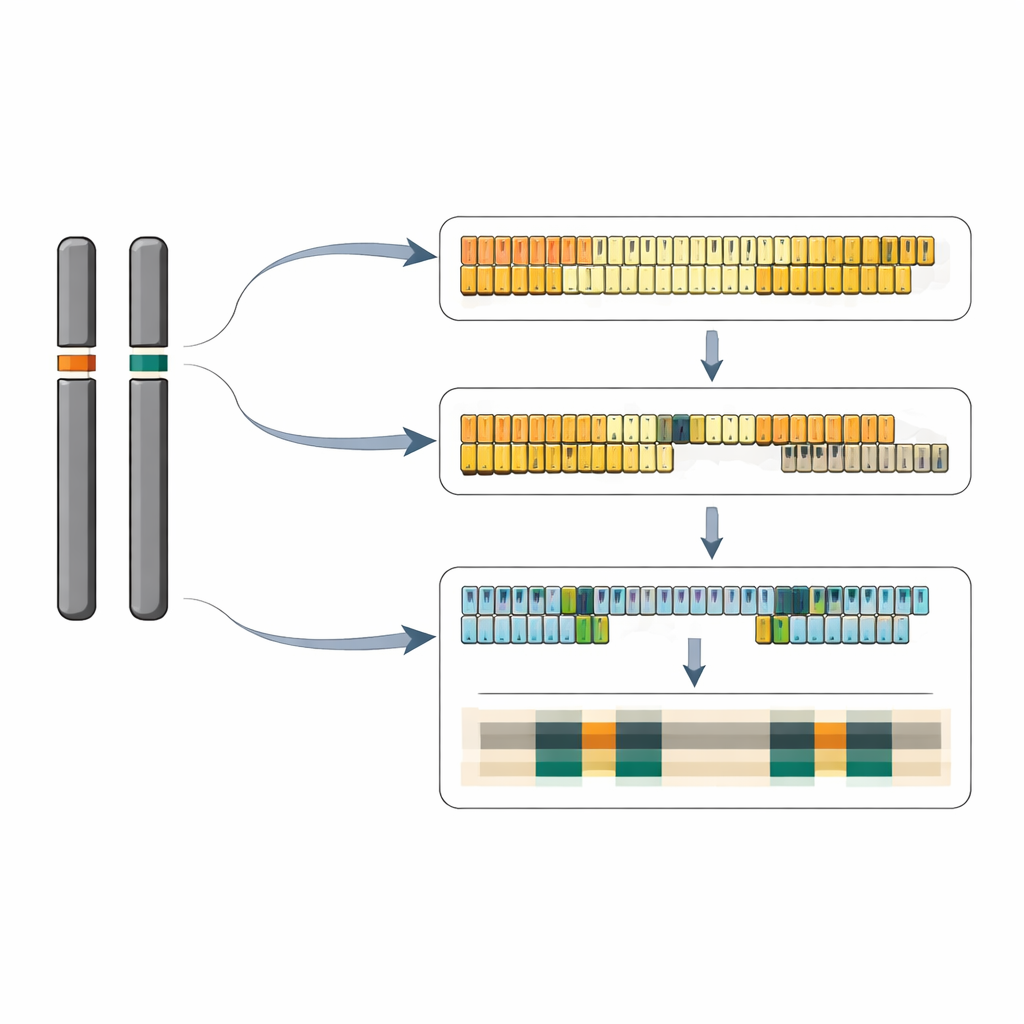
What this means for plant genomes
Taken together, the work delivers the first complete, nucleotide-level view of a plant 5S rDNA system, revealing how repeat number, sequence variants, and surrounding chromosome context come together in a compact genome. Greater duckweed appears to have shed many surplus ribosomal gene copies compared with other flowering plants, yet it maintains two distinct, likely active 5S loci with carefully organized clusters of gene variants. The authors propose that both loci can contribute to 5S RNA production, depending on conditions, and that subtle differences in neighboring DNA and spacer sequences help regulate when and how strongly each cluster is used. Beyond duckweed, the study offers a blueprint for decoding similar repetitive regions in other species and deepens our understanding of how essential gene families are shaped over evolution.
Citation: Stepanenko, A., Schubert, V., Chen, G. et al. Genome sequence assembly of the 5S rDNA loci informs haplotype specificity and evolution in the greater duckweed Spirodela polyrhiza. Commun Biol 9, 516 (2026). https://doi.org/10.1038/s42003-026-09598-8
Keywords: ribosomal DNA, duckweed genome, plant genetics, repetitive DNA, genome evolution