Clear Sky Science · en
Estimating major pathological response in non-small cell lung cancer patients with post-neoadjuvant therapy using MMT-net
Helping Doctors See Treatment Success
When people with lung cancer receive powerful treatments before surgery, doctors need to know how well those therapies worked inside the tumor. Today, that judgment relies on pathologists who painstakingly examine huge digital microscope images by eye. This study introduces a new artificial intelligence tool, called MMT-Net, that can rapidly and precisely estimate how much of a lung tumor has been destroyed by treatment, potentially making care faster, more consistent, and more personalized.
Why Measuring What Is Left Matters
For patients with non-small cell lung cancer, doctors often use drugs and radiation before surgery to shrink or weaken the tumor. After surgery, pathologists estimate the “major pathological response” by checking what fraction of the tumor bed still contains living cancer cells. If only a small amount—about one‑tenth or less—remains, patients tend to live longer and do better overall. However, modern digital slides can contain billions of pixels, and therapy can leave behind a confusing mixture of dead tissue, scar-like areas, and scattered surviving cells. Manually reviewing these enormous images is slow, exhausting, and prone to variation between experts.
Teaching Computers to Read Cancer Slides
The researchers built MMT-Net to assist pathologists in this demanding task. The system analyzes whole slide images of lung tumors that have already been treated. Instead of relying on standard computer vision models trained on everyday photos, they first let the network teach itself the visual “language” of pathology using thousands of unlabeled tissue snippets. This self-directed learning step helps the model recognize subtle patterns specific to stained microscope images, rather than cats, cars, or landscapes.
Looking Close and Far at the Same Time
MMT-Net also mimics how human experts zoom in and out while reading a slide. The system takes small image patches at three magnifications: low power to capture overall architecture, medium to see patterns of damage, and high power to inspect individual cells. A specialized attention mechanism then fuses these views, allowing the model to weigh fine details and broad context together. A transformer module—originally developed for language translation—uses these fused features to sort each patch into three categories: remaining live tumor, treatment‑related scar tissue, or dead and keratinized material. By summing over the entire slide, MMT-Net estimates the fraction of living tumor cells, the key quantity doctors use to judge response.
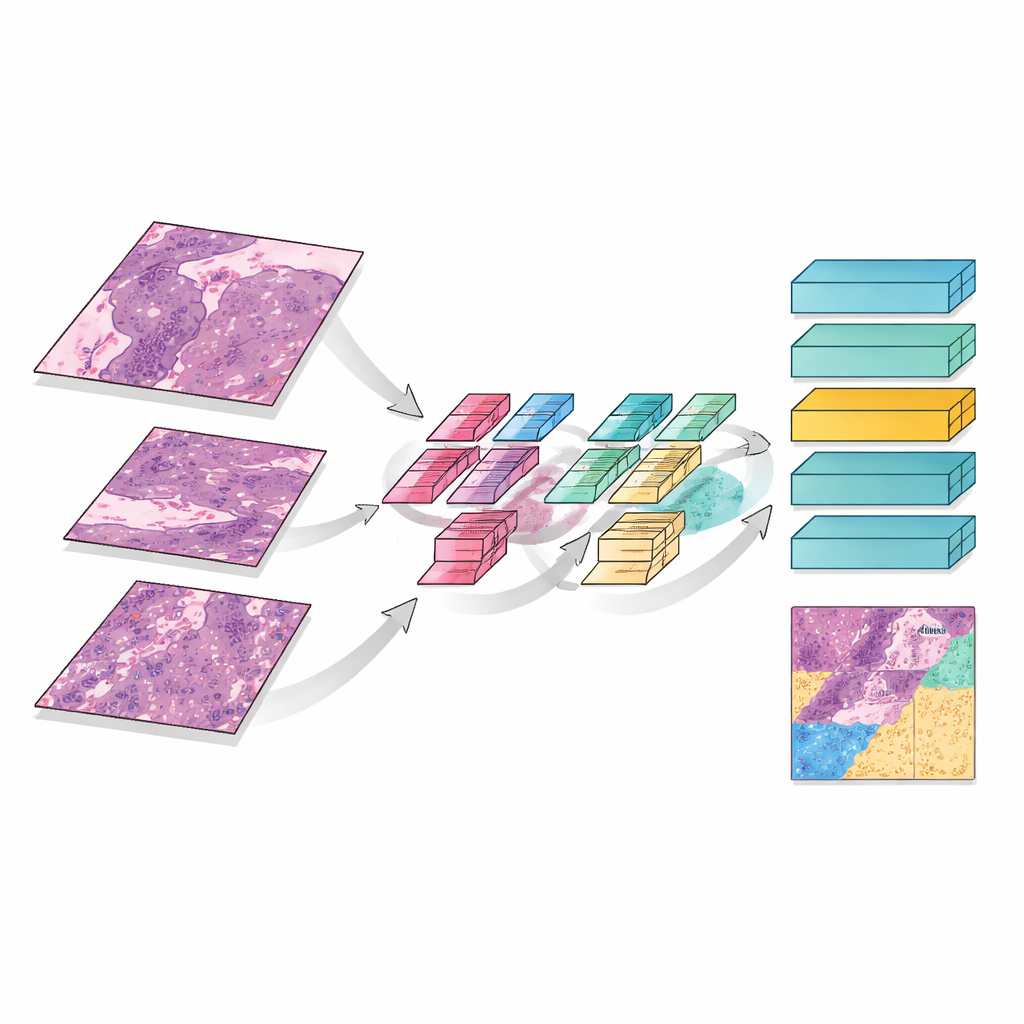
How Well the System Performs
To test the model, the team used slides from 23 patients treated at a single cancer center. They divided the data into training and testing groups, with expert pathologists carefully marking regions of viable tumor, scar-like stroma, and dead tissue according to international guidelines. Compared with several leading deep learning methods, MMT-Net achieved the highest accuracy, correctly classifying tissue types in about 99 percent of test patches when pre-trained on pathology images. The areas highlighted by the model’s internal attention maps closely matched the regions human specialists focus on, suggesting that the system is “looking” at the right structures rather than relying on spurious cues.
Checking Against Molecular Markers
Beyond visual comparison, the researchers asked whether MMT-Net’s predictions align with biological measures of tumor activity. They examined additional slides stained for two common laboratory markers: cytokeratins, which highlight tumor cells of epithelial origin, and Ki67, which marks dividing cells. The proportion of regions labeled as viable tumor by the model showed a strong statistical correlation with both markers, meaning that where the computer saw active tumor, the laboratory stains agreed. This close match supports the idea that MMT-Net is capturing real aspects of disease biology, not just superficial color differences.
What This Could Mean for Patients
In simple terms, this study shows that a carefully designed AI system can estimate how much lung tumor remains after pre-surgery treatment almost as well as expert pathologists, and in a way that is explainable and biologically grounded. While the work was done on a relatively small group of patients from a single center, and larger multi-hospital studies are still needed, the approach is flexible and could be adapted to other cancers and tasks. In the future, tools like MMT-Net may help standardize how response to therapy is measured, support more reliable decisions about follow‑up treatment, and bring precision medicine closer to everyday clinical practice.
Citation: Yang, C., Liu, L., Xiao, W. et al. Estimating major pathological response in non-small cell lung cancer patients with post-neoadjuvant therapy using MMT-net. Sci Rep 16, 14144 (2026). https://doi.org/10.1038/s41598-026-44633-y
Keywords: lung cancer, digital pathology, artificial intelligence, treatment response, deep learning