Clear Sky Science · en
Dual-branch attention network with deep split convolution and multi-dimensional transformers for medical image segmentation
Sharper views for doctors
Modern scans can reveal tumors, clogged vessels, and damaged organs in stunning detail, but turning those gray-and-white images into clear outlines a computer can understand is still surprisingly hard. Doctors need precise boundaries around organs and diseased tissue to plan surgery, track treatment, and avoid mistakes. This study introduces a new artificial intelligence system, called D3T-Net, that draws those boundaries more accurately and more reliably than many leading methods, potentially easing the workload on radiologists and improving diagnostic confidence.
Why drawing lines on medical scans is so difficult
When a radiologist looks at a CT or X‑ray image, they mentally separate overlapping structures, ignore noise, and infer missing edges. Traditional computer programs struggle with this, especially when organ shapes vary from person to person, or when a tumor’s border is blurred. Earlier systems based on convolutional neural networks excel at picking up local textures and edges, but they tend to see only a small neighborhood at a time. That makes it easy for them to miss the broader context needed to distinguish, for example, a faint tumor edge from normal tissue. On the other hand, newer “Transformer” models are good at capturing long‑range relationships across an entire image, but they often gloss over fine details such as tiny lesions or thin boundaries.
Two complementary ways of seeing
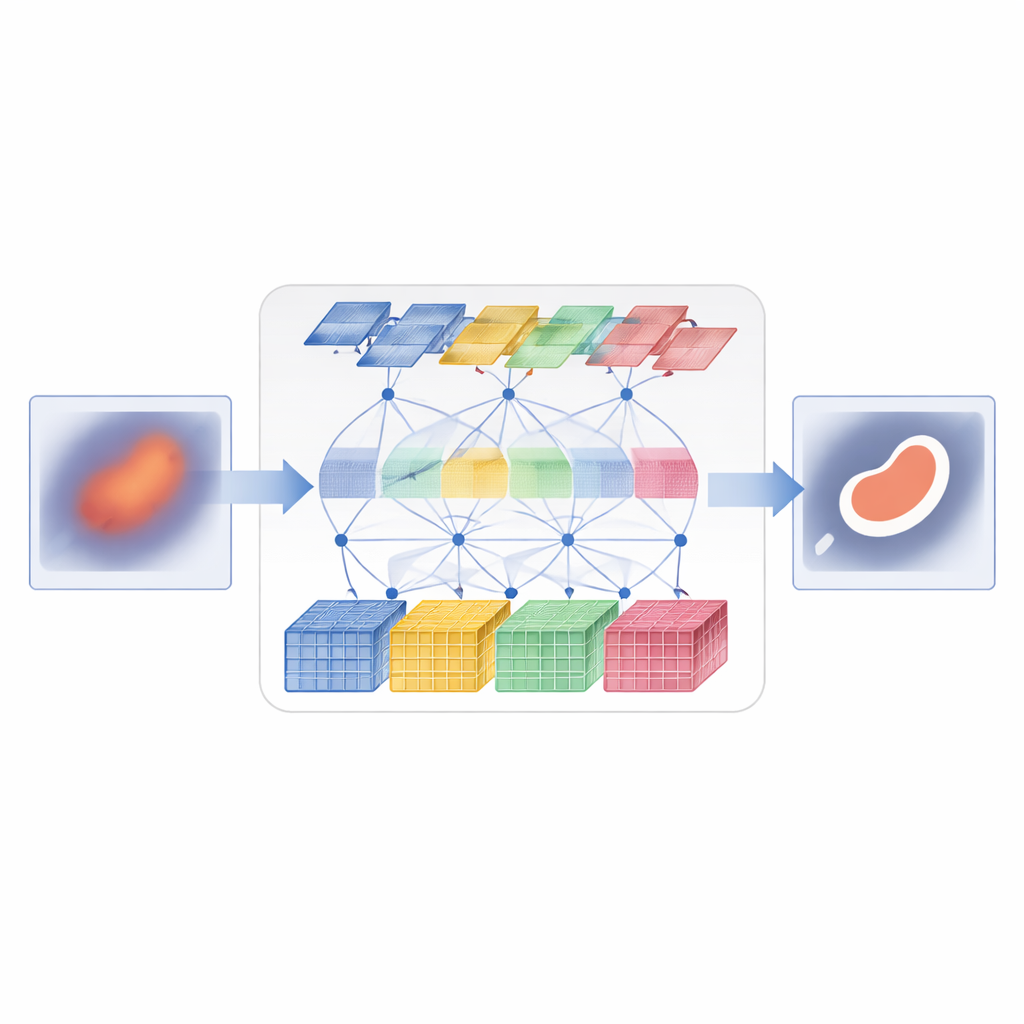
D3T-Net tackles this challenge by combining both ways of seeing into a single, tightly coordinated network. One branch behaves like a traditional image analyzer, focusing on small patches to capture fine textures and crisp edges. It uses a “deep splitting” strategy: the incoming image features are divided into multiple parallel streams, processed separately, and then fused with an attention mechanism that decides which streams carry the most useful structural information. The other branch behaves more like a global observer, using Transformer-style attention to compare distant parts of the image and understand how regions relate to one another. It looks not only across the image plane but also across feature channels, allowing it to capture both where things are and how their appearance patterns hang together.
Getting both branches to cooperate
Simply running two branches in parallel is not enough; they must exchange information in a smart way. In the encoder part of D3T-Net, a special interaction module examines patterns from multiple directions across the image, using pooling and attention to highlight the most informative structures—such as organ outlines or lesion cores—and to share that emphasis between the local and global branches. In the decoder part, where the final segmentation map is assembled, a cross-attention mechanism learns how to combine what each branch has learned, reorganizing features so that global context sharpens local edges and local detail refines the broad global picture. Multi-scale skip connections carry information from early, high‑resolution stages of processing straight to later stages, helping the system keep track of small objects and delicate boundaries that might otherwise be lost.
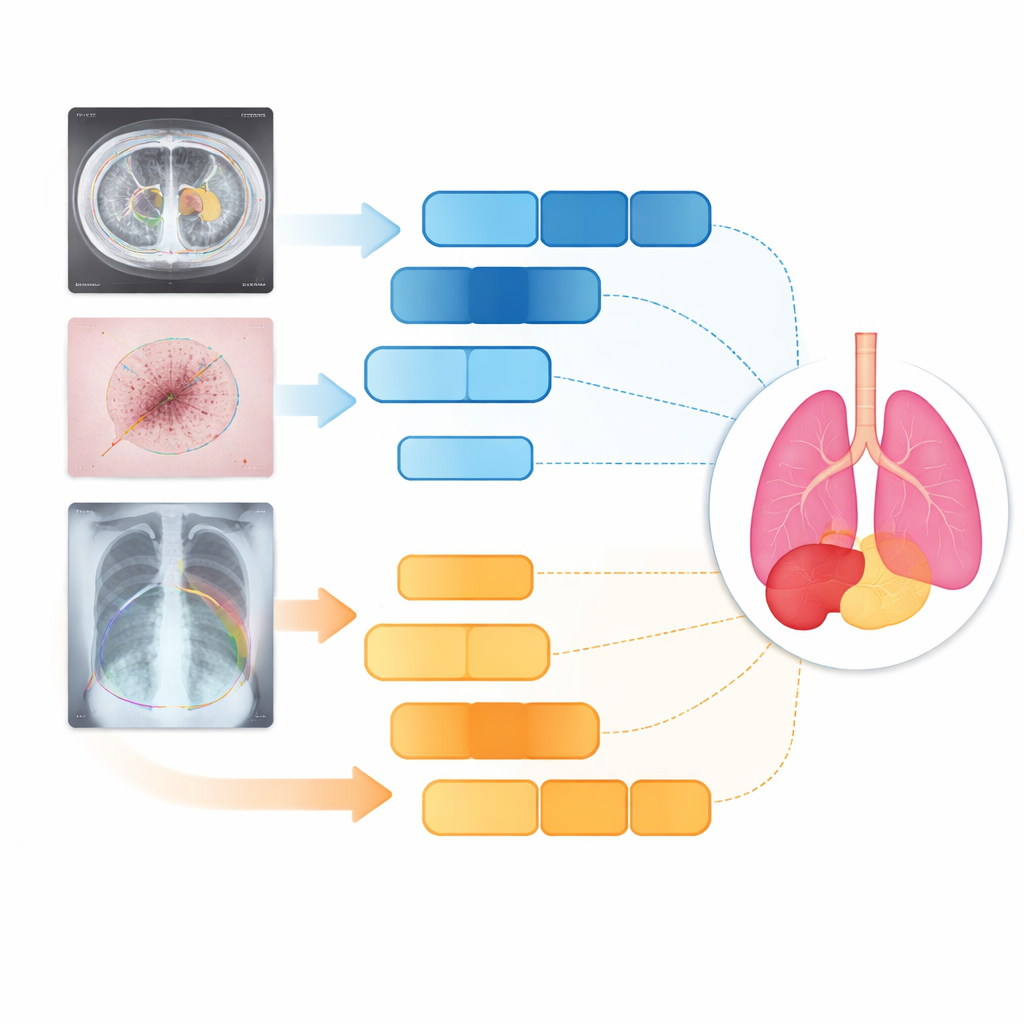
Testing on organs, skin, and lungs
The researchers tested D3T-Net on three very different medical tasks: outlining abdominal organs on CT scans, tracing skin lesions in clinical photographs, and segmenting lungs on chest X‑rays. Across standard accuracy and boundary-sharpness measures, D3T-Net consistently outperformed a broad lineup of state-of-the-art systems, including well-known U-Net variants and Transformer-based hybrids. It was particularly strong at keeping organ contours continuous, correctly separating neighboring structures, and capturing small or low-contrast targets such as the gallbladder or irregular skin lesions. Importantly, these gains came without an extreme increase in computation time: the model’s processing cost remained comparable to many widely used networks, making it plausible for clinical deployment.
What this means for patients and clinicians
In plain terms, the study shows that letting an algorithm "think" both locally and globally at the same time leads to cleaner outlines of organs and disease on medical images. By carefully coordinating a detail‑oriented branch with a context‑aware branch, D3T-Net can separate healthy and unhealthy tissue more accurately than many existing tools. While it will not replace radiologists, it can serve as a powerful assistant—automatically pre‑segmenting scans, flagging subtle lesions, and providing more reliable masks for downstream tasks such as 3D planning or treatment monitoring. As similar dual‑view designs are applied to other imaging problems, patients could benefit from faster, more consistent, and more personalized care.
Citation: Li, D., Yuan, C., Yao, Y. et al. Dual-branch attention network with deep split convolution and multi-dimensional transformers for medical image segmentation. Sci Rep 16, 14238 (2026). https://doi.org/10.1038/s41598-026-44413-8
Keywords: medical image segmentation, deep learning, transformer networks, liver and organ analysis, computer-aided diagnosis